Figures & data
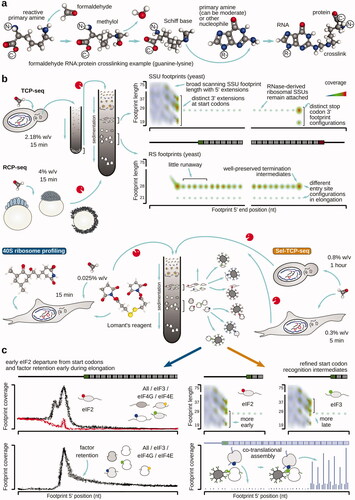
Table 1. Summary of the major approaches employed to arrest or stabilize translational complexes on mRNA for downstream footprinting and footprint profiling.
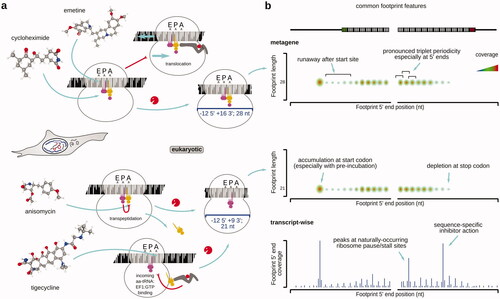
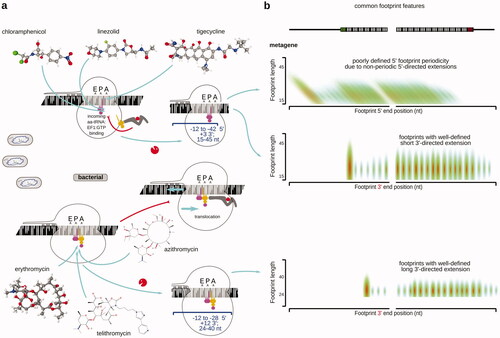
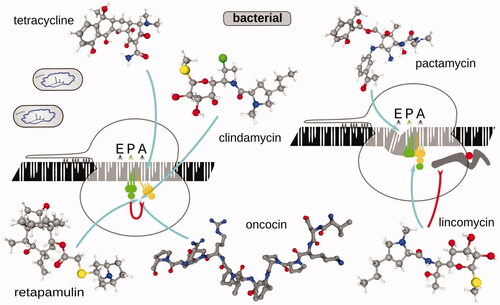
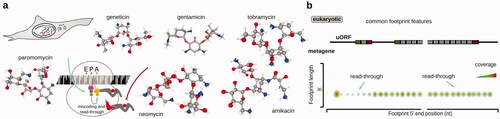
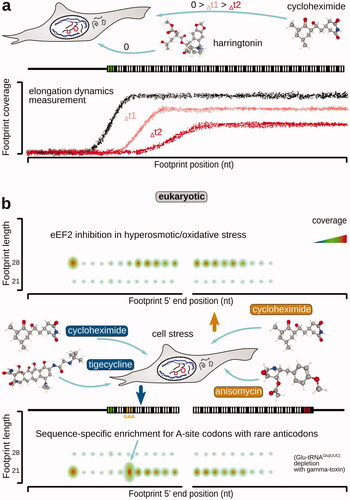
Table 2. Examples of antibiotics and inhibitors used in ribosomal footprinting research to study peptide elongation phase of protein biosynthesis.
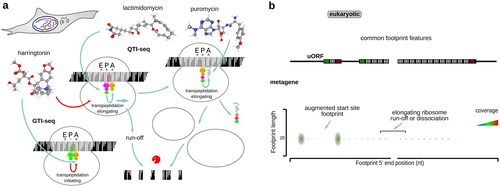
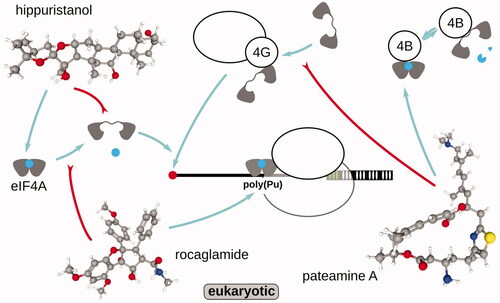
Table 3. Examples of antibiotics and inhibitors used in ribosomal footprinting research to study peptide initiation phase of protein biosynthesis.
Table 4. Examples of antibiotics and inhibitors used in ribosomal footprinting research to study peptide termination and ribosomal recycling phases of protein biosynthesis.