Figures & data
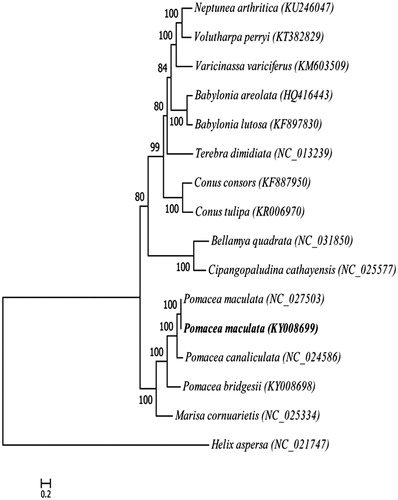
Figure 1. Phylogenetic tree generated by the maximum-likelihood method based on the complete mitochondrial genomes of 15 species of Caenogastropoda, using Helix aspersa as outgroup. The published sequences in GenBank adopted are Neptunea arthritica (KU246047), Volutharpa perryi (KT382829), Varicinassa variciferus (KM603509), Babylonia areolata (HQ416443), Babylonia lutosa (KF897830), Terebra dimidiate (NC_013239), Conus consors (KF887950), Conus tulipa (KR006970), Bellamya quadrata (NC_031850), Cipangopaludina cathayensis (NC_025577), Pomacea maculate (NC_027503), Pomacea maculate (KY008699), Pomacea canaliculata (NC_024586), Pomacea bridgesii (KY008698), Marisa cornuarietis (NC_025334), Helix aspersa (NC_021747).