Figures & data
Figure 1. Evolutionary timeline. Note large intervals for evolution of microbial to eukaryotic life, and for single-celled eukaryotes to metazoans. (see colour version of this figure online at www.informahealthcare.com/bmgwww.informahealthcare.com/bmg).
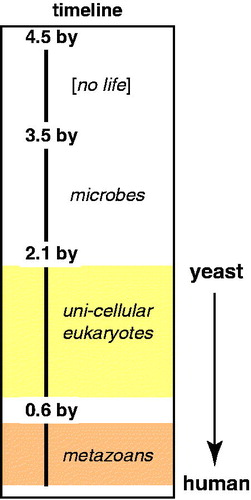
Table 1. Basic comparison of Mediator subunits in humans (Hs), yeast (Sc), fly (Dm), and mouse (Mm) by percent identity, percent similarity, and size.
Table 2. List of human Mediator subunits, along with their approximate molecular weight.
Figure 2. EM structure of human Mediator compared with human Mediator lacking the MED1 and MED26 subunits. Both complexes are bound to the activation domain of VP16, and each is rendered at their predicted molecular weight (1.2 MDa or 0.9 MDa, respectively). The circled region indicates one area of missing protein density in the complex lacking MED1 and MED26. Note, however, that a pol II interaction surface (dashed yellow line; see text) is maintained in both structures, consistent with a general ability of each complex to activate transcription by VP16 (Taatjes & Tjian, Citation2004). (see colour version of this figure online at www.informahealthcare.com/bmgwww.informahealthcare.com/bmg).
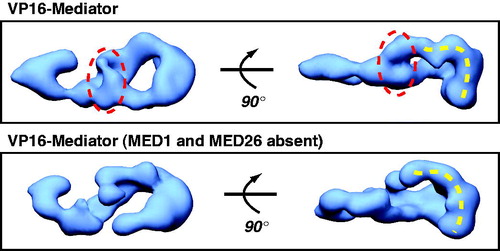
Table 3. DNA-binding TFs and their identified Mediator subunit target(s)*.
Figure 3. Cryo-EM structure of yeast Mediator. The EM data reveal structural flexibility (Cai et al., Citation2009) that can even be inferred from the 3D reconstruction, with its large domains connected by narrow linkers. Note also the extensive surface area, due to channels and cavities in the structure. (see colour version of this figure online at www.informahealthcare.com/bmgwww.informahealthcare.com/bmg).
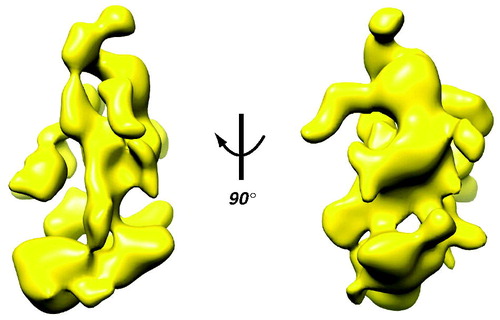
Figure 4. Human Mediator undergoes a structural shift upon binding the pol II CTD. EM structures of unliganded Mediator (left) and CTD-bound Mediator (right) are shown. Note the CTD-Mediator sample is bound to native, full-length (52 YSPTSPS heptad repeat) mammalian CTD (Naar et al., Citation2002). (see colour version of this figure online at www.informahealthcare.com/bmgwww.informahealthcare.com/bmg).
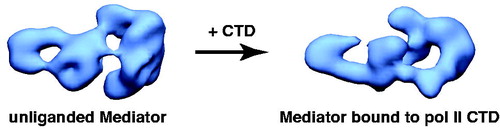
Figure 5. Schematic outlining human Mediator structural changes induced by pol II-TFIIF binding. Two different views (front and side 1) are shown. Three Mediator domains (labeled 1, 2 and 3) are highlighted in the side 1 view and their putative locations are indicated following pol II-TFIIF binding. Note that structural re-organization occurs throughout the Mediator complex upon pol II-TFIIF binding, including the distal “leg/tail” domain (boxed), which represents a site for CDK8 module binding (Bernecky et al., Citation2011). The “Mediator only” and the “Mediator–pol II–TFIIF” structures each are bound to the activation domain of VP16. Pol II is shown in red (PDB 1Y1V). (see colour version of this figure online at www.informahealthcare.com/bmgwww.informahealthcare.com/bmg).
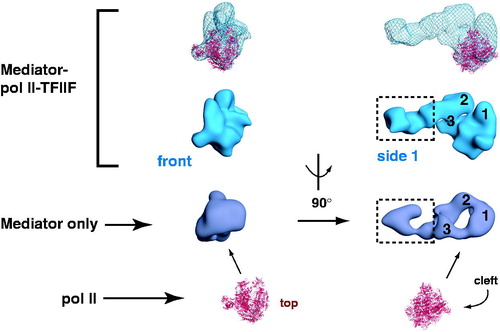
Figure 6. CDK8 module–Mediator binding appears to occlude pol II–Mediator binding by an allosteric mechanism. EM structures of Mediator and CDK8-Mediator (both bound to the activation domain of VP16) are shown (Taatjes et al., Citation2002). The lower panel shows “bottom” views of each complex, with the dashed line on Mediator representing the surface that appears to make direct contacts with pol II (Bernecky et al., Citation2011). The bracket shows the general region occupied by pol II upon binding human Mediator, and the corresponding position in the CDK8-Mediator complex. The structural difference in this bracketed region may reflect a structural change important to prevent pol II (and pol II CTD) binding to CDK8-Mediator. (see colour version of this figure online at www.informahealthcare.com/bmgwww.informahealthcare.com/bmg).
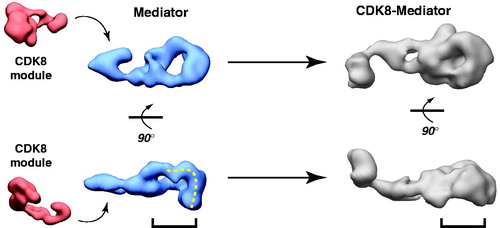
Figure 7. Distinct modes of CDK8 module (CKM) binding to yeast Mediator. EM structure at left shows a single CKM-Mediator interaction via Med13, whereas the structure on the right shows a more extensive interface that also involves Cdk8 (Tsai et al., Citation2013). Scale bar: 100 Å. (see colour version of this figure online at www.informahealthcare.com/bmgwww.informahealthcare.com/bmg).
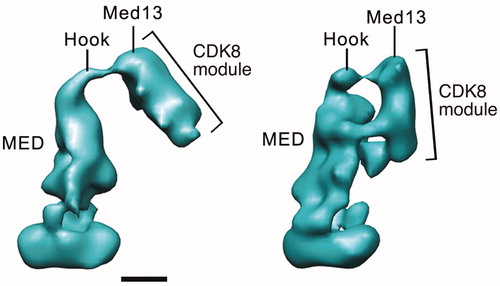
Figure 8. TF binding induces structural shifts throughout the human Mediator complex. Similar views of EM structures of Mediator without a TF bound (top), or bound to the activation domain of SREBP or VP16 are shown (Taatjes et al., Citation2002). Note that structural changes appear to propagate throughout the complex, and that structural changes are distinct for each TF. Localization of the VP16 (X) and SREBP (+) binding sites are shown. Scale bar: 100 Å. (see colour version of this figure online at www.informahealthcare.com/bmgwww.informahealthcare.com/bmg).
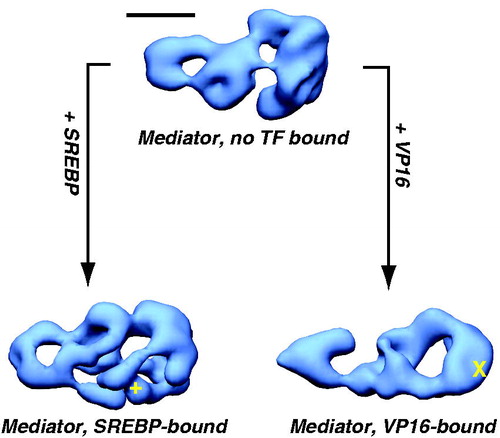
Figure 9. A working model for Mediator and CDK8-Mediator regulation of transcription initiation and elongation. This model depicts four functionally distinct structural states (I–IV) for Mediator. We hypothesize that different Mediator surfaces will be exposed in each state, which may help coordinate timing of factor recruitment to the promoter, in accordance with requirements for various stages of transcription. According to this model, state I and state II are compatible with pre-initiation events, state III represents transcription initiation (possibly including paused pol II), and state IV represents an elongation-competent structure. In state I, Mediator is not bound to a TF; Mediator is capable of binding pol II in this structural state, but pol II will be inactive or minimally active (i.e. basal transcription). TF binding (e.g. VP16) causes a structural shift to state II. Mediator is also capable of binding pol II in this conformational state, with the potential to direct high levels of “activated” transcription. This structural state might also coordinate timing of other Mediator-cofactor interactions at the promoter that could regulate subsequent stages of transcription (Ebmeier & Taatjes, Citation2010). If pol II binds the TF-Mediator complex, this leads to structural state III. This structural state may be compatible with activated transcription, perhaps by promoting synergy among PIC factors (e.g. TFIIH, TFIID and TFIIB) that assemble around the Mediator–pol II complex. Note that in this structural state, the CDK8 module is incapable of binding Mediator. Upon transcription initiation and pol II transition to productive elongation, pol II breaks contacts with Mediator; Mediator structure transitions back to state II (TF bound, but no pol II). The CDK8 module is able to bind Mediator in this structural state. If the CDK8 module binds Mediator, Mediator adopts structural state IV. This structural state (i.e. CDK8-Mediator) does not allow pol II binding. Thus, the CDK8-Mediator complex prevents a second pol II enzyme from immediately re-engaging the promoter, which might otherwise cause defects in mRNA processing or defects during initiation by this second pol II. Furthermore, the CDK8-Mediator complex could help assemble and/or regulate elongation factors, thereby influencing ongoing elongation events. The ability of CDK8-Mediator or core Mediator (i.e. Mediator containing MED26) to positively influence pol II elongation has been documented by several groups (Donner et al., Citation2010; Galbraith et al., Citation2013; Takahashi et al., Citation2011). Yet Mediator and other PIC components remain at the promoter following pol II promoter escape, leaving a “scaffold” complex (Yudkovsky et al., Citation2000). These apparently contradictory findings are reconciled by growing evidence that elongating pol II complexes are likely stationary, and that rather than moving directionally along DNA, pol II instead “reels in” the DNA template (Papantonis et al., Citation2010). This has already been demonstrated for bacterial polymerases (Kapanidis et al., Citation2006; Revyakin et al., Citation2006), and DNA polymerases work in much the same way (Anachkova et al., Citation2005). Stationary, elongating pol II complexes could be juxtaposed with promoter-bound factors, facilitating Mediator- or CDK8-Mediator-dependent regulation of pol II elongation. We emphasize that this is a model, and that many aspects remain to be rigorously tested. (see colour version of this figure online at www.informahealthcare.com/bmgwww.informahealthcare.com/bmg).
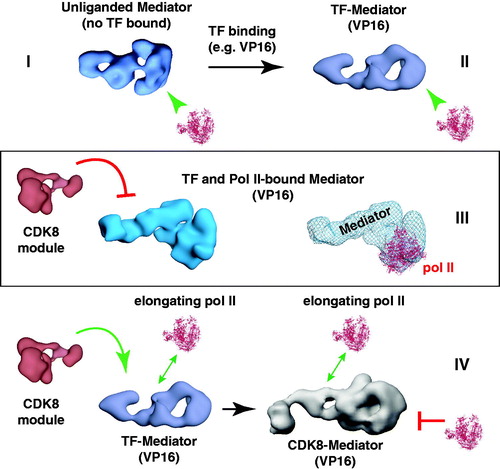
Figure 10. A structural model of the human PIC. The cryo-EM structure of human pol II, TBP, TFIIA, TFIIB, TFIIE and TFIIF bound to promoter DNA (closed complex (He et al., Citation2013)) was docked into the cryo-EM map of human Mediator–pol II–TFIIF (Bernecky et al., Citation2011). In the docked structures, the Mediator-pol II-TFIIF cryo-EM map is shown in blue mesh, whereas the color-coding for the other PIC factors is indicated. For reference, the same orientation of the Mediator–pol II–TFIIF structure alone is shown in solid blue. Addition of TFIIH (pink) to the model blocks details of the structure, therefore, we show the model with and without TFIIH (below). The view without TFIIH also indicates an open region for its assembly into the PIC. Note that some structural reorganization occurs within the PIC upon TFIIH binding (He et al., Citation2013). To generate the model, the docked pol II crystal structure was used as a reference to align both cryo-EM maps in Chimera. (see colour version of this figure online at www.informahealthcare.com/bmgwww.informahealthcare.com/bmg).
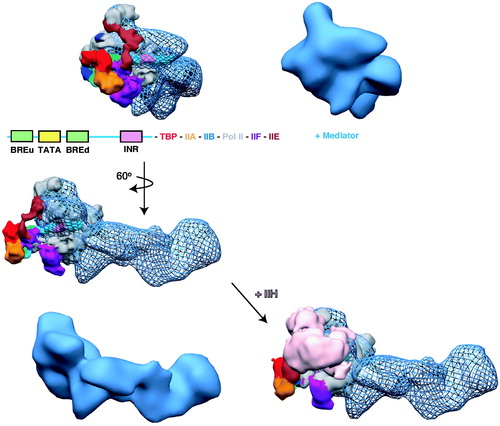
Figure 11. A common structural feature among TF-bound human Mediator complexes. EM structures are shown for Mediator bound to different TF activation domains and compared with Mediator that is not TF-bound (inset). A shared structural feature is a “pocket” (green arrow) and the surfaces corresponding to probable sites of pol II interaction are highlighted with the dashed yellow line. Note these features are absent from the unliganded Mediator structure. (see colour version of this figure online at www.informahealthcare.com/bmgwww.informahealthcare.com/bmg).
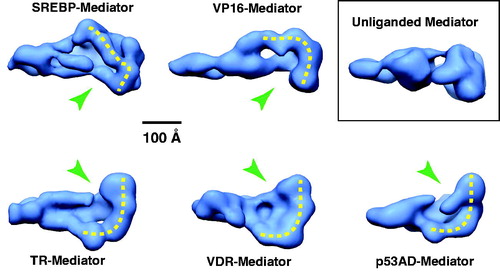
Figure 12. A model for TF-dependent “post-recruitment” activation of a fully assembled but latent PIC. In the absence of a key TF, a PIC may occupy the promoter, but pol II remains largely inactive or paused. Upon TF binding to the promoter, it interacts with Mediator and triggers a structural shift in the complex, which activates the PIC and allows pol II to escape the promoter region and transition to a productively elongating state. Part of this process could involve functional synergy between Mediator and pausing/elongation factors such as DSIF, Gdown1/POLR2M or the SEC. (see colour version of this figure online at www.informahealthcare.com/bmgwww.informahealthcare.com/bmg).
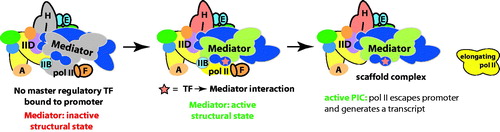
Figure 13. Gdown1/POLR2M, TFIIF and pol II each converge on the same structural interface of Mediator. At left is shown a “bottom” view of the VP16-Mediator complex (Taatjes et al., Citation2002). A pol II interaction surface is highlighted by the yellow dashed line. This distinctive interaction surface forms upon TF binding (see ). At center is a “front” view of the Mediator complex, with pol II (red ribbon; PDB 1Y1V) oriented consistent with its bound state orientation in the VP16-Mediator-pol II-TFIIF assembly (Bernecky et al., Citation2011), shown at right. Highlighted at right is a general location for Gdown1 binding to pol II, based upon cryo-EM data (Wu et al., Citation2012), as well as the approximate location of TFIIF, based upon crosslinking-MS data and cryo-EM data (Chen et al., Citation2010b; Eichner et al., Citation2010; He et al., Citation2013). The cryo-EM map for the VP16-Mediator-pol II-TFIIF assembly is shown in blue mesh, with pol II docked as described (Bernecky et al., Citation2011). (see colour version of this figure online at www.informahealthcare.com/bmgwww.informahealthcare.com/bmg).
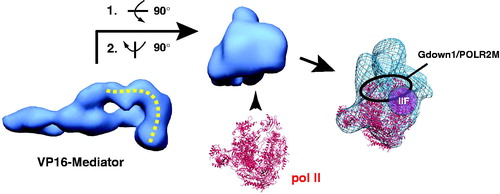
Figure 14. A simple schematic illustrating enhancer-promoter communication via Mediator. Mediator can bind simultaneously to enhancer-bound TFs and the PIC, including pol II. (see colour version of this figure online at www.informahealthcare.com/bmgwww.informahealthcare.com/bmg).
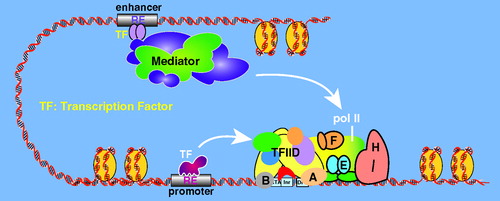
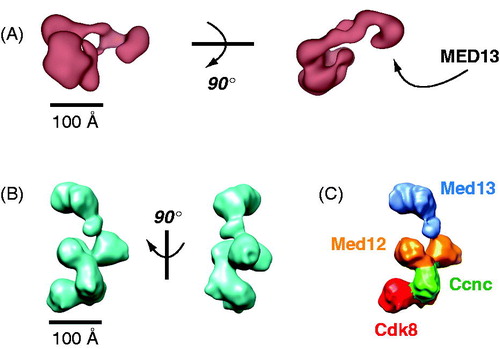
Figure 15. EM structures of the human (A) and yeast (B) CDK8 module. The structures are distinct and not shown in similar orientations. Structural distinctions may derive from sequence differences, in particular, the much larger sizes for human MED12 and MED13. MED13 forms an extended hook-like structure in each, and this subunit has been shown to contact core Mediator (Knuesel et al., Citation2009a; Tsai et al., Citation2013). Whereas the general location of MED13 was determined for the human CDK8 module, localization of each subunit was determined for the yeast structure with antibody labeling (C). (see colour version of this figure online at www.informahealthcare.com/bmgwww.informahealthcare.com/bmg).