ABSTRACT
Pseudognaphalium Kirp. (Compositae: Gnaphalieae) comprises approximately 90 species and it has a worldwide but fragmented distribution. This study provides a total of 20 chromosome counts, nine of which are the first report for a species, and the rest are confirmation of previous reports. With the addition of these data to the previously published reports, the chromosome numbers for a total of 32 species of the genus Pseudognaphalium are now available. Considering all chromosome data available at present, c.22% of the species are diploids and c.78% are polyploids, mainly tetraploids. African and Asian species are predominantly diploid whereas the vast majority of the American species are tetraploid, suggesting that polyploidy played a key role in the colonization of the American continent by the genus Pseudognaphalium.
Introduction
The genus Pseudognaphalium Kirp. (Compositae: Gnaphalieae) is one of the most species-rich genera of the tribe Gnaphalieae and it is constituted by annual, biennial or perennial herbs with clusters of capitula usually arranged in corymbiform or paniculiform synflorescences and with monochromous involucral bracts (Anderberg Citation1991; Hilliard and Burtt Citation1981; Hilliard Citation1983). It comprises approximately 90 species and has a worldwide but fragmented distribution. It includes African, Asian and American taxa, including some species endemic to the Caribbean and the Hawaiian archipelagos, and the subcosmopolitan Pseudognaphalium luteoalbum (L.) Hilliard & B.L. Burtt. However, the vast majority of the species are distributed in America (Anderberg Citation1991; Hilliard and Burtt Citation1981; Freire et al. Citation2014).
Existing chromosome data show that base chromosome number of Pseudognaphalium is x = 7, and the species mainly present 2n = 14 chromosomes (diploids) or 2n = 28 chromosomes (tetraploids). Additionally, the numbers 2n = 16, 2n = 20 and 2n = 40 have occasionally been reported (). Smissen et al. (Citation2011) and Galbany-Casals et al. (Citation2014), based on molecular phylogenies, hypothesized that Pseudognaphalium could have one or multiple allopolyploid origins. However, these authors highlighted the need for obtaining more chromosome counts and also more complete and well-resolved phylogenies to corroborate this hypothesis. Allopolyploidy is considered a significant mechanism in the evolutionary success of many species, in particular by enabling the colonization of new niches and the adaptation of polyploids to new habitats (Ramsey Citation2011; Mota et al. Citation2016). Also, a pattern of higher probability of long distance dispersal events and colonization of new areas has been observed in polyploid groups (Linder and Barker Citation2014), and several adaptive radiations in islands have been preceded by allopolyploidization (Barrier et al. Citation1999; Mandáková et al. Citation2010; Joly et al. Citation2009). The recurrent presence of polyploids in habitats or geographic areas different from those of their diploid progenitors suggests the ability of polyploids to colonize new environmental niches (Hegarty and Hiscock Citation2008). However, it may show also their difficulty in succeeding in areas in which the parental species occur (Thompson Citation2005; Ramsey Citation2011).
Table 1. Available chromosome counts for the genus Pseudognaphalium, with indication of the relevant publication references and of the geographic area of the species. North America here refers to the American part of the Boreal Kingdom. Central America here refers to the area from Guatemala and Belize to Panama.
Given the reported variation in chromosome numbers in Pseudognaphalium and their probable significance in its origin and biogeographical history, it is essential to have a more complete picture of chromosome data for the genus, as chromosome number is only available for 22 Pseudognaphalium species, which represent c.24% of the species of the genus. In this context, the aims of this work are to provide new chromosome counts for Pseudognaphalium, to compile all chromosome number reports available, and to examine the distribution patterns of ploidy levels in the genus. These data will be essential for the interpretation of future phylogenetic works focused on the origin and biogeographic history of the genus.
Material and methods
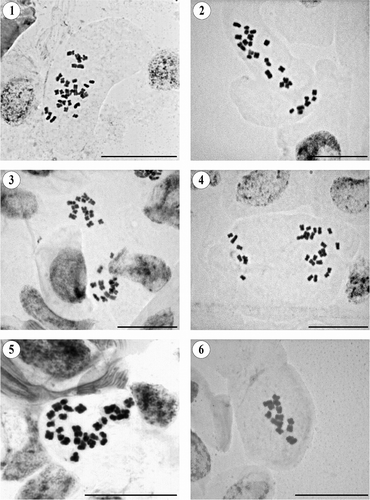
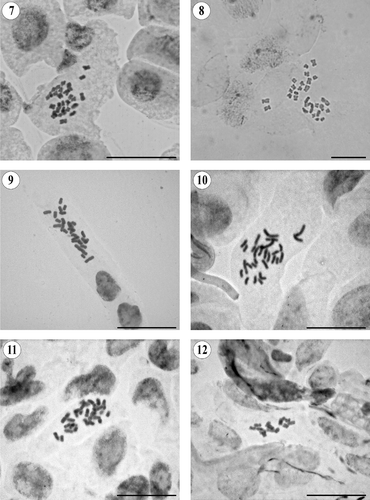
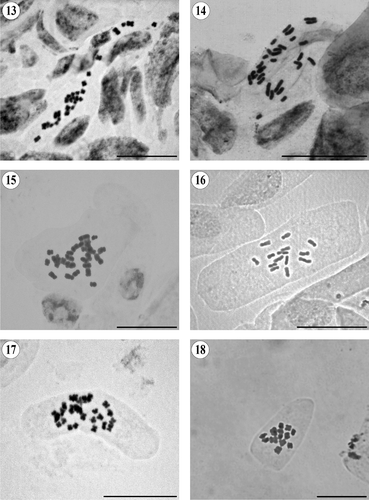
Chromosome counts were made on mitotic metaphases using standard squash techniques. Seeds collected in the wild were used, and voucher specimens were deposited in the herbaria of the Botanical Institute of Barcelona (BC) and in the National Herbarium of Mexico (MEXU). For one species, seeds were taken from a specimen deposited in the herbarium of the University of Salamanca (SALA). Root-tip meristems were obtained by germinating seeds on wet filter paper in Petri dishes at approximately 20°C. Samples were pretreated with 0.05% colchicine for 2 h 15 min at room temperature. The material was fixed in 3: 1 v/v absolute ethanol: glacial acetic acid at 4°C. Meristems were hydrolysed in 5 M hydrogen chloride (HCl) for 50 min to 1 h at room temperature. They were then stained in 2% acetic orcein for a minimum of 3 h at room temperature. Squashes were made in 45% acetic acid. Photographs were taken through a Zeiss Axioscope (Oberkochen, Germany) compound microscope with a Jenoptik ProGres C3 (Jena, Germany) digital camera using the program ProGres Capture 7 (Jenoptik, Jena, Germany).
Published chromosome data for the genus Pseudognaphalium were retrieved from the “Index to chromosome numbers in Compositae” website (http://www.lib.kobe-u.ac.jp/infolib/meta_pub/G0000003asteraceae_e), “Index to plant Chromosome Numbers” website (http://www.tropicos.org/Project/IPCN) and Nesom (Citation2006). Most of the original references cited in these online sources were later searched and consulted to cover one or more references for each of the chromosome numbers reported for every species.
Results
The newly obtained chromosome numbers for each species are given below along with some relevant notes. The species are listed in alphabetical order. The data on species habitat and geographic distribution were taken from Hilliard (Citation1983), McVaugh (Citation1984), Anderberg (Citation1991), Espinosa-García (Citation2005), Nesom (Citation2006), Chen and Bayer (Citation2011), Freire et al. (Citation2014) and Hinojosa-Espinosa and Villaseñor (Citation2014), and from our own field observations. The new chromosome counts obtained in this work are also included in , where all gathered information of previously published chromosome counts of the genus is compiled. We only cite the references that have been directly checked by us, which are representative of all the chromosome numbers documented for each species up to the present.
Pseudognaphalium arizonicum (A. Gray) Anderb.
This taxon is an annual herb distributed from southern USA to central Mexico. It grows in open woodlands, chaparral and in pine-oak forest clearings (Espinosa-Garcia Citation2005; Nesom Citation2006).
Mexico: Veracruz, road 140, between Las Vigas de Ramírez and La Joya, 19°37ʹ35.2ʺ N, 97°4ʹ20.8ʺ W, 2412 m, 1 September 2014, Galbany GC-2470–2 & Arrabal (BC).
2n = 28 (). This is the first chromosome count for this species, which is shown to be tetraploid.
Pseudognaphalium canescens (DC.) Anderb.
This species is an annual herb, distributed from the south of the USA to central Mexico. It occurs in lava beds, rocky sites, grasslands, scrublands, oak, pine-oak, and pine woodlands, often on slopes (McVaugh Citation1984; Espinosa-García Citation2005; Nesom Citation2006).
Mexico: Mexico City, Tlalpan, Ajusco Medio, ca. Km 5 of the Picacho-Ajusco road, about 150 meters south of the park Ecoguardas and the Tlalpan Police Subdelegation, 19°16ʹ16ʺ N, 99°12ʹ21ʺ W, 2610 m, 13 December 2014, Hinojosa-Espinosa 590 (MEXU).
2n = 28 (). Our result agrees with previous gametophytic counts for this tetraploid species (Keil and Pinkava Citation1976; Keil et al. Citation1988; Carr et al. Citation1999), but this is the first mitotic count.
Pseudognaphalium chartaceum (Greenm.) Anderb.
This species is an annual or biennial herb endemic to Mexico (Espinosa-García Citation2005; Villaseñor Citation2016). It occurs on forested slopes, hills, ravines, forest clearings, and roadsides, mostly in tropical deciduous forests and oak or oak-pine forests (McVaugh Citation1984).
Mexico: Mexico City, “Bosque de Tlalpan”, Las Antenas road, at 500 m to the north of the entrance located in De las Torres Avenue, Colonia Miguel Hidalgo (2nd Section), 19°17ʹ21ʺ N, 99°11ʹ44ʺ W, 2410 m, 14 December 2014, Hinojosa-Espinosa 592 (BC, MEXU).
2n = 28 (). Our mitotic chromosome count of this tetraploid species agrees with that of Turner and Zhao (Citation1992) and with the gametophytic count of Keil and Stuessy (Citation1977).
Pseudognaphalium cheiranthifolium (Lam.) Hilliard & B.L. Burtt
This taxon is a perennial herb distributed in southern South America, where it grows on rocky and sandy soils (Freire et al. Citation2014).
Chile: Valparaíso Region, Valparaiso, Quintero, Ritoque beach, rocky spur at the north end of the beach, 32°49ʹ36.7ʺ S, 71°31ʹ45.5ʺ W, 10 m, 18 November 2015, Galbany GC-2513 & Arrabal (BC).
2n = 28 (). Our mitotic count for this tetraploid species is the first one from Chilean material and is consistent with previously published gametophytic counts based on material from Argentina (Hunziker et al. Citation1990; Carr et al. Citation1999).
Pseudognaphalium conoideum Anderb.
This taxon is an annual herb distributed in Mexico, where it appears in disturbed areas, meadows and crops (Espinosa-García Citation2005).
Mexico: State of Mexico, around Lagunas de Zempoala, 19°2ʹ52.7ʺ N, 99°19ʹ25.5ʺ W, 2887 m, 17 August 2014, Galbany GC-2443, Magallón & Arrabal (BC).
2n = 28 (). This is the first chromosome count for this tetraploid species.
Pseudognaphalium cymatoides (Kuntze ex DC.) Anderb.
This species is an annual or biennial herb distributed in South America, particularly in Bolivia, Argentina and Chile (Freire et al. Citation2014).
Chile: Región IV, Coquimbo, arriving to Sotaquí from Monte Patria or Combarbalá, Quebrada Seca, 30°39ʹ29.1ʺ S, 71°05ʹ15.8ʺ W, 98 m, 20 November 2015, Galbany GC-2526 & Arrabal (BC).
2n = 14 (). Our chromosome count is the first one for the species. It is diploid, which seems to be rather exceptional in the American species of the genus Pseudognaphalium.
Pseudognaphalium elegans (Kunth) Kartesz.
This taxon is distributed in Mexico, Central America and South America. It grows in volcanic beds, disturbed areas and forest clearings.
Mexico: Veracruz, road 140, between Las Vigas de Ramírez and La Joya, 19°37ʹ35.2ʺ N, 97°4ʹ20.8ʺ W, 2412 m, 1 September 2014, Galbany GC-2471–2 & Arrabal (BC).
2n = 28 (). Our report is the first mitotic count for this tetraploid species, and the first count from Mexican material. It is consistent with previous gametophytic counts based on material from Colombia (Powell and King Citation1969; Jansen et al. Citation1984) and Venezuela (Carr et al. Citation1999).
Pseudognaphalium greenmanii (S.F. Blake) Anderb.
This annual species is endemic to Mexico and occurs in oak-pine forests in steep grassy or rocky hills (McVaugh Citation1984).
Mexico: Jalisco, Zapopan, way to La Gotera, Pinar de la Venta, 20°42ʹ56.4ʺ N, 103°31ʹ29.0ʺ W, 1725 m, 20 August 2014, Galbany GC-2449–1, Arrabal & Juárez (BC).
2n = 28 (). This is the first chromosome count for this tetraploid species.
Pseudognaphalium heterotrichum (Phil.) Anderb.
This taxon is a perennial herb endemic to Chile (Reiche Citation1903), where it grows in open shrublands with Cactaceae, in herbaceous communities of disturbed areas or in road margins.
Chile: IV Coquimbo Region, Coquimbo, Canela Baja, next to a chapel, 31°24ʹ29.3ʺ S, 71°28ʹ14.8ʺ W, 286 m, 19 November 2015, Galbany GC-2522 & Arrabal (BC).
2n = 28 (). This is the first report for this tetraploid species. Freire et al. (Citation2014) consider this species a synonym of Pseudognaphalium gayanum (J. Rémy) Anderb., for which there are no available chromosome counts.
Pseudognaphalium jaliscense (Greenm.) Anderb.
This annual plant is endemic to western and north-western Mexico, where it grows in meadows or grasslands, openings in oak-pine forests, often on roadsides or in disturbed places (McVaugh Citation1984).
Mexico: Jalisco, Zapopan, surroundings of the “Centro Universitario de Ciencias Biológicas y Agropecuarias”, 20°44ʹ22.5ʺ N, 103°30ʹ51.1ʺ W, 1647 m, 20 August 2014, Galbany GC-2446–1, Arrabal & Juárez (BC).
2n = 28 (). This is the first chromosome count for this tetraploid species.
Pseudognaphalium liebmannii (Sch. Bip. ex Klatt) Anderb. var. liebmannii
This perennial herb is distributed in Mexico and Central America. It grows mostly at the timberline and above, occupying openings in pine forests, on grassy, rocky or gravelly slopes, in alpine grasslands and disturbed sites such as in volcanic debris (McVaugh Citation1984; Espinosa-García Citation2005).
Mexico: State of Mexico, Iztaccihuatl volcano, between Paso de Cortés and the parking area called “La Joya”, 19°5ʹ53.6ʺ N, 98°39ʹ7.2ʺ W, 3753 m, 6 September 2014, Galbany GC-2479 & Arrabal (BC) ().
2n = 28 (). Beaman et al. (Citation1962) indicated Gnaphalium vulcanicum I.M. Johnst., which is a synonym of P. liebmannii var. liebmannii (McVaugh Citation1984), presents 2n = 28. Our chromosome count of P. liebmannii var. liebmannii is consistent with this previous count and confirms this species is tetraploid.
Pseudognaphalium liebmannii (Sch. Bip. ex Klatt) Anderb. var. monticola (McVaugh) Hinojosa & Villaseñor
This taxon is an annual or perennial herb and it is distributed in Mexico and Central America. It grows in clearings of fir forests, humid pine forests, and pine-oak forests, on rocky slopes and summits, and on roadsides and in disturbed areas (McVaugh Citation1984; Espinosa-García Citation2005; Hinojosa-Espinosa and Villaseñor Citation2014).
Mexico: State of Mexico, Iztaccihuatl volcano, between Paso de Cortés and the parking area called “La Joya”, 19°5ʹ38.3ʺ N, 98°39ʹ4.1ʺ W, 3710 m, 6 September 2014, Galbany GC-2476–3 & Arrabal (BC).
2n = 28. As in the previous case, our chromosome count agrees with that of Beaman et al. (Citation1962); however, this is the first report for this variety.
Pseudognaphalium luteoalbum (L.) Hilliard & B.L.Burtt
This is an annual herb with a subcosmopolitan distribution, known from Europe, Asia, America, Australia, New Zealand and other Pacific Islands (Nesom Citation2006; Chen and Bayer Citation2011). It is considered introduced in most part of these areas and its original distribution is not known with certainty. It grows along roadsides, in fields and pastures, humid grasslands near streams and marshes, in gardens, river beds and disturbed areas (Hilliard Citation1983; Nesom Citation2006).
Mexico: Mexico City, “Ciudad Universitaria”, “Jardín Botánico”, 19°19ʹ5ʺ N, 99°11ʹ39.1ʺ W, 2327 m, 5 August 2014, Galbany GC-2411 & Arrabal (BC).
2n = 14 (). Many studies have reported chromosome counts of this subcosmopolitan species (see ). The most commonly found chromosome number is 2n = 14, which indicates it is diploid. Vogt and Aparicio (Citation1999) reported the number 2n = 28, but to our knowledge this is the only report of a tetraploid level. This may indicate that the species can experience intraspecific polyploidy, or that the chromosome count might be an error. Unfortunately, the authors did not include an image of the chromosome count. Additional and exceptional chromosome numbers have been reported for P. luteoalbum: Gill and Omoigui (Citation1987) reported n = 8, and Subramanian (Citation1987) 2n = 20. None of these works contained an image of the chromosome count cited. These odd numbers have been found only once each. Thus, they could be considered as anomalous counts within the species, or perhaps errors. Finally, in the “Index to chromosome numbers in Compositae” database is a report of 2n = 18 (Fernandes and Queiros Citation1971). Actually, Fernandes and Queiros (Citation1971) reported several counts of 2n = 14 for P. luteoalbum, but none of 2n = 18.
Pseudognaphalium oxyphyllum (DC.) Kirp. var. oxyphyllum
This annual herb is distributed in Central America and Mexico (Espinosa-García Citation2005). This species grows in clearings of oak or pine forests or in disturbed areas (Espinosa-García Citation2005).
Mexico: Mexico City, “Ciudad Universitaria”, close to “Ciudad Universitaria” metrobus station, between the station and the Pumabus stop, 19°19ʹ25.6ʺ N, 99°11ʹ19.8ʺ W, 2304 m, 8 September 2014, Galbany GC-2480 & Arrabal (BC).
2n = 28 (). Our chromosome count is the first one for this tetraploid species.
Pseudognaphalium semilanatum (DC.) Anderb.
This taxon is an annual plant endemic to Mexico (Villaseñor Citation2016). It grows in rocky or steep mountainsides and summits, grasslands, open oak woodlands, pine or pine-oak forests, often in disturbed ground (McVaugh Citation1984).
Mexico: Veracruz, road 140, between Perote and Las Vigas de Ramírez, a little after passing Cruz Blanca, 19°38ʹ10.6ʺ N, 97°9ʹ9.1ʺ W, 2440 m, 1 September 2014, Galbany GC-2469 & Arrabal (BC).
2n = 28 (). This is the first chromosome count for this tetraploid species. García-Espinosa (Citation2005) stated that this species may be synonym of Pseudognaphalium semiamplexicaule (DC.) Anderb. or that both could be part of a complex of closely related species. McVaugh (Citation1984) recognized it as a separate species closely related to P. oxyphyllum, P. semiamplexicaule, P. canescens and Pseudognaphalium roseum (Kunth) Anderb.
Pseudognaphalium stramineum (Kunth) Anderb.
This species is an annual or biennial plant, widely distributed in North America, Mexico and Central America. It occurs in a wide variety of habitats, such as humid fields and stream banks, dunes, chaparral slopes, roadsides or disturbed places (Espinosa-García Citation2005; Nesom Citation2006).
Mexico: State of Mexico, around Xalatlaco, 19°9ʹ29.2ʺ N, 99°24ʹ6.8ʺ W, 2850 m, 9 August 2014, Galbany GC-2430 & Arrabal (BC).
2n = 28 (). Our report is the first mitotic count for this tetraploid species, which is consistent with the previous gametophytic counts (Keil and Stuessy Citation1977; Sundberg and Dillon Citation1986).
Pseudognaphalium undulatum (L.) Hilliard & B.L. Burtt
This species is an annual herb, naturally distributed in southern Africa and Madagascar, and naturalized in several parts of Europe and North Africa. It grows on damp places, particularly around rock outcrops, on stream- and river-banks, or near forest margins or in open fynbos (Hilliard Citation1983).
South Africa: Matroosberg Private Reserve, Matroosberg, Ski Hut., 33°20ʹ46.7ʺ S, 19°37ʹ26.5ʺ E, 1219 m, 21 January 2014, Andrés-Sánchez SA827 & Smuts (SALA 156,613).
2n = 14 (). Our report is the first one for this African diploid species.
Pseudognaphalium viravira (Molina) Anderb.
This species is a perennial herb distributed in Chile and Argentina (Freire et al. Citation2014), where it grows on dry soils in open shrublands, along disturbed roadsides with other herbaceous species, or in forest clearings.
Chile: IV Coquimbo Region, from Quilimarí (Los Vilos) inland by D-875 road, 32°07ʹ17.4ʺ S, 71°27ʹ47.4ʺ W, 40 m, 19 November 2015, Galbany GC-2517 & Arrabal (BC).
2n = 28 (). Our result is consistent with the gametophytic count of Spooner et al. (Citation1987), but this is the first mitotic chromosome count for this species.
Pseudognaphalium viscosum (Kunth) Anderb.
This annual herb is distributed from the southern USA to Honduras (Espinosa-García Citation2005; Nesom Citation2006). It grows in pine or oak forest clearings, grasslands, fields, shrublands, often in rocky sites, disturbed areas or roadsides (Espinosa-García Citation2005; Nesom Citation2006).
Mexico: Mexico City, “Ciudad Universitaria”, “Jardín Botánico”, 19°19ʹ5ʺ N, 99°11ʹ39.1ʺ W, 2327 m, 5 August 2014, Galbany GC-2412–1 & Arrabal (BC); “Ciudad Universitaria”, next to the metrobus station called “Ciudad Universitaria”, 19°19ʹ25.6ʺ N, 99°11ʹ19.8ʺ W, 2304 m, 31 July 2014, Galbany GC-2408–1 & Arrabal (BC).
2n = 14 (). We obtained the same chromosome diploid number for two different specimens. Our results are consistent with the gametophytic count of Turner et al. (Citation1962) and are the first mitotic counts of 2n = 14. However, two counts of 2n = 28 also exist. The first one is from Canada (Chinnappa and Chmielewski Citation1987) and is clearly an error, given that P. viscosum is not present in that country. Nesom (Citation2006) noted that reports of P. viscosum in states other than Texas are based on plants of Pseudognaphalium macounii (Greene) Kartesz, a very similar species that is widely distributed in the USA and Canada. In particular, P. macounii is present in British Columbia, where the material for that count was taken (Chinnappa and Chmielewski Citation1987). The other published tetraploid report is a gametophytic count based on material from Chihuahua, Mexico (Carr et al. Citation1999). We examined the voucher corresponding to this count [Chihuahua, 33.7 miles west of La Junta, 30 September 1989, R. M. King 9863 & P. M. Peterson (US 3,140,891)], and concluded it does not belong to P. viscosum, but to P. stramineum. This is consistent with the chromosome number obtained by us for P. stramineum in the present work, 2n = 28. Therefore, we note that the two reports of 2n = 28 for P. viscosum are errors and that this species is diploid with 2n = 14.
Discussion
This study includes a total of 20 chromosome counts, nine of which are the first report for a species, and the rest are confirmation of previous reports. With these data, the chromosome number is known for a total of 32 species of the genus Pseudognaphalium, c.35% of the species of the genus.
It is already known that Pseudognaphalium includes diploids and tetraploids (Smissen et al. Citation2011; Galbany-Casals et al. Citation2014). With chromosome data available at present, only seven species are diploids with 2n = 14 (c.22% of the species with available data) and 24 species are tetraploids with 2n = 28 (c.78% of the species with available data). Two of the diploid species, P. luteoalbum and P. hypoleucum (DC.) Hilliard & B.L. Burtt, are actually reported to present both ploidy levels, 2n = 14 and 2n = 28. Indeed, intraspecific polyploidy has been reported for many plant species (e.g. Gültepe et al. Citation2015; Meng et al. Citation2016). However, for the two mentioned species, the chromosome number 2n = 28 has only been reported once for each. Thus, additional counts of 2n = 28 would be desirable to confirm the tetraploid level in these two species. Finally, Carr et al. (Citation1999) reported Pseudognaphalium purpurascens (DC.) Anderb. as 2n = 40. These authors interpreted this odd number as a dysploid derivate of the hexaploid based on x = 7 or alternatively a tetraploid based on x = 10. Hexaploids with base x = 7 (2n = 42) occur within Helichrysum (Febles Citation1989; Galbany-Casals and Romo Citation2008), a genus that is closely related to Pseudognaphalium. These data, and the fact that all New World Pseudognaphalium species with available chromosome data present the base number of x = 7, makes the hypothesis of a dysploid derivate of an hexaploid based on x = 7 more probable.
The chromosome data available in this study show that the African and Asian species are predominantly diploid, whereas the vast majority of the New World species are tetraploid. Considering Galbany-Casals et al. (Citation2014) placed the origin of Pseudognaphalium in Africa, chromosome data available may indicate that the colonization of the American continent by this genus could have been stimulated or facilitated by one or more polyploidization events, the colonization routes being still unknown. Chromosome data presented here will help to disentangle the role of allopolyploidy in the colonization of new territories when a more complete phylogeny of Pseudognaphalium is provided.
Acknowledgements
M. G.-C. greatly appreciates Sebastián Arrabal’s help, enthusiastic support, dedication and joy in all the shared field expeditions. David Juárez and Sebastián Teillier provided very valuable help and advice for field trips and plant collections, and Santiago Andrés provided material from his field collections; all of them are gratefully acknowledged. We thank Susana Magallón for organizing the stay of M. G.-C. at the “Universidad Nacional Autónoma de México”. We thank the staff of the herbaria of “Institut Botànic de Barcelona” (BC, Neus Nualart), “Universidad de Concepción” (CONC, Alicia Marticorena), “Universidad Nacional Autónoma de México” (MEXU, David Gernandt and José Luis Villaseñor), Missouri Botanical Garden (MO, John Pruski), “Universidad de Salamanca” (SALA, Francisco Javier Hernández), “Museo Nacional de Historia Natural de Santiago de Chile” (SGO, Gloria Rojas), University of Texas at Austin (TEX, George Yatskievych), and Smithsonian Institution (US, Ingrid P. Lin) for kindly attending M. G.-C. and for providing specimens and/or institutional support for plant collections. Laia Guàrdia provided the microscope and camera, and the technical support necessary for using them. Joan Prunera provided technical support for some of the chromosome counts. Samuel Pyke is acknowledged for the English correction. Two anonymous reviewers contributed to improve the paper.
Disclosure statement
No potential conflict of interest was reported by the authors.
Additional information
Funding
References
- Anderberg AA. 1991. Taxonomy and phylogeny of the tribe Gnaphalieae (Asteraceae). Opera Bot. 104:1–195.
- Arano H. 1956. Karyotaxonomic studies in tribe Tubuliflorae of Compositae. I. The karyotype analysis and its phylogenic consideration in subtribe Gnaphaliineae. Jap J Genet. 31(5):137–143.
- Arano H. 1963. Cytological studies in subfamily Carduoideae (Compositae) of Japan. XIII. The karyotype analysis on subtribe Gnaphaliineae. Bot Mag (Tokyo). 76(905):419–427.
- Bala S, Gupta RC. 2013. Male meiosis and chromosome number in Asteraceae family from district Kangra of H. P. (Western Himalayas). Int J Bot Res. 3(1):43–58.
- Barrier M, Baldwin BG, Robichaux RH, Purugganan MD. 1999. Interspecific hybrid ancestry of a plant adaptive radiation: allopolyploidy of the Hawaiian Silversword Alliance (Asteraceae) inferred from floral homeotic gene duplications. Molec Biol Evol. 16:1105–1113.
- Beaman JH, De Jong DCD, Stoutamire WP. 1962. Chromosome studies in the alpine and subalpine floras of Mexico and Guatemala. Amer J Bot. 49(19):41–50.
- Carr GD, King RM, Powell AM, Robinson H. 1999. Chromosome numbers in Compositae. XVIII. Amer J Bot. 86(7):1003–1013.
- Chen Y, Bayer RJ. 2011. Pseudognaphalium Kirp. In: Wu CY, Raven PH, Hong D, editors. Flora of China. Vol. 20-21. Beijing and St. Louis: Science Press and Missouri Botanical Garden Press; [accessed 2017 Jun]. http://www.efloras.org/florataxon.aspx?flora_id=2&taxon_id=127088.
- Chinnappa CC, Chmielewski JG. 1987. Documented plant chromosome numbers 1987: 1. Miscellaneous counts from western North America. Sida. 12:409–417.
- Dawson MI, Beuzenberg EJ. 2000. Contributions to a chromosome atlas of the New Zealand flora — 36. Miscellaneous families. New Zealand J Bot. 38:1–23.
- De Lange PJ, Murray BG, Datson PM. 2004. Contributions to a chromosome atlas of the New Zealand flora—38. Counts for 50 families. New Zealand J Bot. 42:873–904.
- Dempsey RE, Gornall RJ, Bailey JP. 1994. Contributions to a cytological catalogue of the British and Irish flora, 4. Watsonia. 20:63–66.
- Espinosa-García FJ. 2005. Gnaphalium L. In: Calderón G, Rzedowski J, editors. Flora fanerogámica del Valle de México. Michoacán (Pátzcuaro): Instituto de Ecología, A.C. y Comisión Nacional para el Conocimiento y Uso de la Biodiversidad; p. 840–856.
- Febles R. 1989. Estudios en la flora Macaronésica: algunos números de cromosomas VI. Bot Macaronés. 17:57–71.
- Fernandes A, Queiros M. 1971. Contribution a la connaissance cytotaxinomique des Spermatophyta du Portugal. II. Compositae. Bol Soc Broteriana. 45:5–122.
- Fernández-Casas FJ, Fernández-Piqueras J. 1981. Estudio cariológico de algunas plantas bolivianas. Anales Jard Bot Madrid. 38:149–152.
- Freire SE, Bayón ND, Baeza CM, Giuliano DA, Monti C. 2014. Revision of the genus Pseudognaphalium (Asteraceae, Gnaphalieae) in Chile. Gayana Bot. 71:68–107.
- Gadella WJ, Kliphuis E. 1966a. Chromosome numbers of flowering plants in the Netherlands. III. Proc Kon Neerl Acad Wetenschaf Amsterdam Ser C. 70:7–20.
- Galbany-Casals M, Romo A. 2008. Polyploidy and new chromosome counts in Helichrysum Mill. (Asteraceae, Gnaphalieae). Bot J Linn Soc. 158:511–521.
- Galbany-Casals M, Unwin M, Garcia-Jacas N, Smissen RD, Susanna A, Bayer RJ. 2014. Phylogenetic relationships in Helichrysum (Compositae: Gnaphalieae) and related genera: incongruence between nuclear and plastid phylogenies, biogeographic and morphological patterns, and implications for generic delimitation. Taxon. 63:608–624.
- Gill LS, Omoigui ID. 1987. The incidence of polyploidy in family Asteraceae in southern Nigeria. Rev Cytol Biolo Vég Botaniste. 10:177–184.
- Gosavi KVC, Kambale SS, Yadav SR. 2012. Gnaphalium luteoalbum L. In: Marhold K, editor. IATP/IOPB chromosome data 14. Vol. 61 (6). Taxon. 61(6):1338.
- Groves BE. 1977. Contributions to a chromosome atlas of the New Zealand flora—19. Gnaphalium (Compositae). New Zealand J Bot. 15:17–18.
- Gültepe M, Coşkunçelebi K, Makbul S, Vladimirov V. 2015. Chromosome counts of Tragopogon L. (Asteraceae) from Turkey. Caryologia. 68(3):193–199.
- Gupta H, Gupta RC, Kumar R, Singhal VK. 2017. A profile of chromosome counts, male meiosis and pollen fertility in 45 species of Asteraceae from Parvati Valley in Kullu district, Himachal Pradesh. Caryologia. 70:128–140.
- Hegarty MJ, Hiscock SJ. 2008. Genomic clues to the evolutionary success of polyploid plants. Curr Biol. 18:R435–444.
- Hilliard OM. 1983. Pseudognaphalium Kirp. In: Leistner OA, editor. Flora of southern Africa. Vol. 33 (7, 2). Pretoria: Department of Agriculture; p. 53–56.
- Hilliard OM, Burtt BL. 1981. Some generic concepts in Compositae-Gnaphaliinae. Bot J Linn Soc. 82:181–232.
- Hinojosa-Espinosa O, Villaseñor JL. 2014. New combinations in Pseudognaphalium (Gnaphalieae-Asteraceae) of Mexico. Bot. Sciences. 92(4):489–491.
- Hsu CC. 1970. Preliminary chromosome studies on the vascular plants of Taiwan. (III). The aster family. Compositae. Taiwania. 15(1):17–29.
- Hunziker JH, Escobar A, Xifred CC, Gamerro JC. 1990. Estudios cariológicos en Compositae. VI. Darwiniana. 30:115–121.
- Jansen RK, Stuessy TF, Díaz-Piedrahíta S, Funk VA. 1984. Recuentos cromosómicos de Compositae de Colombia. Caldasia. 14(66):7–20.
- Joly S, Heenan PB, Lockhart PJ. 2009. A Pleistocene intertribal allopolyploidization event precedes the species radiation of Pachycladon (Brassicaceae) in New Zealand. Molec Phylogen Evol. 51:365–372.
- Karna Mallick P, Manandhar Laxmi M, Laxmi VB. 2011. Chromosome numbers of some taxa of the nepalese Asteraceae. Bull Pure Appl Sci – Bot. 30b(1 and 2):55–68.
- Keil DJ, Luckow MA, Pinkava DJ. 1988. Chromosome studies in Asteraceae from the United States, Mexico, the West Indies, and South America. Amer J Bot. 75:652–668.
- Keil DJ, Pinkava DJ. 1976. Chromosome counts and taxonomic notes for Compositae from the United States and Mexico. Amer J Bot. 63(10):1393–1403.
- Keil DJ, Pinkava DJ. 1981. IOPB Chromosome number reports LXXII. Taxon. 30:705–706.
- Keil DJ, Stuessy TF. 1977. Chromosome counts of Compositae from Mexico and the United States. Amer J Bot. 64(6):791–798.
- King RM. 1965. Chromosome numbers of Thailand Compositae. Phytologia. 11(4):217–218.
- Kockx-Van Roon M, Wieffering JH. 1982. IOPB chromosome number reports LXXV. Taxon. 31:367.
- Lane MA, Li J. 1993. Documented chromosome numbers 1993: 1. Chromosome number reports in Compositae with emphasis on tribe Astereae of the southwestern United States and Mexico. Sida. 15:539–546.
- Linder HP, Barker NP. 2014. Does polyploidy facilitate long-distance dispersal? Ann. Bot. (Oxford). 113:1175–1183.
- Love Á, Kjellqvist E. 1974. Cytotaxonomy of Spanish plants. IV. Dicotyledon: Caesalpinaceae-Asteraceae. Lagascalia. 4(2):153–211.
- Mandáková T, Heenan PB, Lysak MA. 2010. Island species radiation and karyotypic stasis in Pachycladon allopolyploids. BMC Evol Biol. 10:367.
- McVaugh R. 1984. Gnaphalium L. In: Flora Novo-Galiciana. vol. 12, Compositae. Ann Arbor (Michigan): The University Press; p. 446–468.
- Mehra PN, Gill BS, Mehta JK, Sidhu SS. 1965. Cytological investigations on the Indian Compositae. I. North — indian Taxa. Caryologia. 18(1):35–68.
- Meng Y, Yang Y-P, Deng T, Nie Z-L. 2016. New chromosome counts, polyploidy, and karyotype evolution in Aster L. (Asteraceae: astereae) from the Qinghai-Tibet Plateau. Caryologia. 69(4):370–378.
- Morton JK. 1981. Chromosome numbers in Compositae from Canada and the U.S.A. Bot J Linn Soc. 82:357–368.
- Morton JK. 1993. Chromosome numbers and polyploidy in the flora of Cameroon Mountain. Oper Bot. 121:159–172.
- Mota L, Torices R, Loureiro J. 2016. The evolution of haploid chromosome numbers in the sunflower family. Genome Biol Evol. 8(11):3516–3528.
- Nesom GL. 2006. Pseudognaphalium Kirp. In: Flora of North America Editorial Committee, editors. Flora of North America North of Mexico. Vol. 19. New York: Oxford: Oxford University Press; [ accessed 2016 Jun]. http://www.efloras.org/florataxon.aspx?flora_id=1&taxon_id=127088.
- Powell AM, Cuatrecasas J. 1970. Chromosome numbers in Compositae: Colombian and Venezuelan species. Ann Missouri Bot Gard. 57(3):374–379.
- Powell AM, King RM. 1969. Chromosome numbers in the Compositae: Colombian species. Amer J Bot. 56(1):116–121.
- Powell AM, Kyhos DW, Raven PH. 1974. Chromosome numbers in Compositae. X. Amer J Bot. 61(8):909–913.
- Raj B. 1965. Chromosome numbers in some Indian angiosperms. II. Proc Indian Acad Sci. Section B. 61(5):253–261.
- Ramsey J. 2011. Polyploidy and ecological adaptation in wild yarrow. Proc Natl Acad Sci USA. 108:7096–7101.
- Razaq ZA, Vahidy AA, Ali SI. 1994. Chromosome numbers in Compositae from Pakistan. Ann Missouri Bot Gard. 81:800–808.
- Reiche C. 1903. Gnaphalium L. In: Flora de Chile. Vol. 4 (I). Santiago de Chile: Imprenta Barcelona; p. 47–73.
- Ruíz De Clavijo E. 1990. Números cromosomáticos de plantas occidentales, 619–630. Anales Jard Bot Madrid. 47(2):431–437.
- Shetty BV. 1967. IOPB chromosome numbers reports XIV. Taxon. 16(6):552–571.
- Smissen RD, Galbany-Casals M, Breitwieser I. 2011. Ancient allopolyploidy in the everlasting daisies (Asteraceae: Gnaphalieae): complex relationships among extant clades. Taxon. 60:649–662.
- Smith EB. 1969. IOPB chromosome number reports XXII. Taxon. 18:433–442.
- Spooner DM, Stuessy TF, Crawford DJ, Silva M. 1987. Chromosome numbers from the flora of the Juan Fernandez Islands. II. Rhodora. 89:351–356.
- Subramanian D. 1987. Cytotaxonomical studies in the South Indian species of Inuleae (Asteraceae). J Cytol Genet. 22:77–82.
- Sundberg SD, Dillon MO. 1986. IOPB Chromosome number reports XCI 91. Taxon. 35:409–410.
- Thompson JD. 2005. Plant evolution in the Mediterranean. Oxford: Oxford University Press.
- Turner BL. 1970. Chromosome numbers in the Compositae. XII. Australian species. Amer J Bot. 57(4):382–389.
- Turner BL, King RM. 1964. Chromosome numbers in the Compositae. VIII. Mexican and Central American species. Southwest Nat. 9(1):27–39.
- Turner BL, Powell AM, King RM. 1962. Chromosome numbers in the Compositae. IV. Additional Mexican and Guatemalan species. Rhodora. 64(759):251–271.
- Turner BL, Zhao Z. 1992. Documented chromosome numbers 1992: 2. Miscellaneous U.S.A. and Mexican species, mostly Asteraceae. Sida. 15:147–150.
- Villaseñor JL. 2016. Checklist of the native vascular plants of Mexico. Revista Mexicana De Biodiversidad. 87:559–902.
- Vogt R, Aparicio A. 1999. Chromosome numbers of plants collected during Iter Mediterraneum IV in Cyprus. Bocconea. 11:117–169.
- Vogt R, Oberprieler C. 1993. Chromosome numbers of North African phanerogams. I. Fl Medit. 3:187–210.
- Vogt R, Oberprieler C. 1994. Chromosome numbers of North African phanerogams. IV. Candollea. 49(2):549–570.