ABSTRACT
In this study, DNA sequence data were used to characterize 290 Fusarium strains isolated during a survey of root-colonizing endophytic fungi of agricultural and nonagricultural plants in northern Kazakhstan. The Fusarium collection was screened for species identity using partial translation elongation factor 1-α (TEF1) gene sequences. Altogether, 16 different Fusarium species were identified, including eight known and four novel species, as well as the discovery of the phylogenetically divergent F. steppicola lineage. Isolates of the four putatively novel fusaria were further analyzed phylogenetically with a multilocus data set comprising partial sequences of TEF1, RNA polymerase II largest (RPB1) and second-largest (RPB2) subunits, and calmodulin (CaM) to assess their genealogical exclusivity. Based on the molecular phylogenetic and comprehensive morphological analyses, four new species are formally described herein: F. campestre, F. kazakhstanicum, F. rhizicola, and F. steppicola.
INTRODUCTION
Fungal endophytes that colonize living plant tissues without causing visible symptoms on their hosts can be found in many plant species and play crucial roles in ecosystems (Porras-Alfaro and Bayman Citation2011; Rodriguez et al. Citation2009). Roots harbor a diverse community of fungal endophytes (Rodriguez et al. Citation2009; Vandenkoornhuyse et al. Citation2002), but the nature of the relationship is poorly understood (Newsham Citation2011; Porras-Alfaro and Bayman Citation2011). Knowledge of root endophytic fungal diversity is increasing (Kia et al. Citation2017; Knapp et al. Citation2018; Sieber and Grünig Citation2013), but the focus has mostly been on root-colonizing communities of grassland ecosystems (Jumpponen et al. Citation2017; Knapp et al. Citation2019, Citation2012; Li et al. Citation2018; Mandyam et al. Citation2010; Porras-Alfaro et al. Citation2008). Studies of root endophytes in some regions of the Eurasian steppe belt (Khidir et al. Citation2010; Knapp et al. Citation2019, Citation2012) have revealed that those in grass roots form communities similar to other grassland regions of the world. However, no detailed study of endophyte diversity has been conducted in the extensive grasslands of the semiarid steppe ecosystem in the Republic of Kazakhstan, which occupies one-third of the country (Klein et al. Citation2012). This ecosystem is characterized by black, fertile, humus-rich soil (i.e., chernozem) that is excellent for cultivating cereals and other crops (Mermut Citation1999).
Kazakhstan is one of the main producers and exporters of cereal crops to Europe, Russia, Ukraine, northern Africa, and Central Asia (FAOSTAT Citation2021). During our studies focusing on isolation and characterization of root-associated endophytic fungi of agronomically important crops and native plants in northern Kazakhstan, 550 fungal isolates were collected. The core root endophytic taxa detected were similar to those found in other grassland areas (Akhmetova et al. Citation2021). Interestingly, Fusarium was the dominant genus, constituting two-thirds of the isolates cultured from healthy roots (Akhmetova et al. Citation2021). Although many Fusarium species are important plant pathogens and produce diverse mycotoxins that pose a significant threat to global food safety, plant health, and agricultural biosecurity (Audenaert et al. Citation2009), they frequently inhabit healthy roots asymptomically (Kuldau and Yates Citation2000; Maciá-Vicente et al. Citation2008), as reported in previous studies (Knapp et al. Citation2012, Citation2019). Endophytic Fusarium strains have been shown to improve plant performance under various stress conditions (Kavroulakis et al. Citation2018). Moreover, comparative genomic analyses revealed that root-colonizing F. oxysporum possess similar carbohydrate-activated enzymes (CAZymes) to other root endophytes (Knapp et al. Citation2018).
Initially, our 550-isolate collection was typed with DNA sequences of the nuclear ribosomal internal transcribed spacer (ITS) region (Schoch et al. Citation2012). All 290 Fusarium isolates were typed using translation elongation factor 1-alpha (TEF1; O’Donnell et al. Citation1998b; Villani et al. Citation2016), the most informative marker locus for species identification within this genus (Geiser et al. Citation2004). Because several of the Fusarium isolates appeared to represent novel species based on phylogenetic analyses of the partial TEF1 sequence data, the present study was conducted to (i) characterize the putatively novel species using multilocus DNA sequence data and phylogenetic species recognition based on genealogical exclusivity (GCPSR sensu Taylor et al. Citation2000) and (ii) formally describe the new Fusarium species using GCPSR and detailed morphological analyses. Herein, four novel Fusarium species are formally described, including one that represents a novel, phylogenetically divergent lineage that is a distant sister to F. nurragi + the F. tricinctum and F. heterosporum species complexes (Geiser et al. Citation2021; Laraba et al. Citation2022)
MATERIALS AND METHODS
Isolation and identification of fungal root endophytes.—
Root samples were collected in the arid Shortandy District of the Akmola Region in northern Kazakhstan where the climate is sharply continental, with cold winters and hot summers. The growing season is short, and drought represents a major stress factor (Pavlova et al. Citation2014). Root samples were collected from natural habitats of the region and from agricultural fields of the A.I. Barayev Scientific Production Center for Grain Farming at the laboratories of Crop Rotation and Precision Agriculture (SUPPLEMENTARY TABLE 1). During the first sampling (10–14 Sep 2018), roots of five gramineous plants were sampled: wheat (Triticum aestivum), barley (Hordeum vulgare), oat (Avena sativa), barley grass (Hordeum leporinum), and the native steppe feather or needle grass (Stipa capillata). During the second sampling (11–13 Aug 2019), roots were collected from the plants listed above together with lentil (Lens culinaris, Fabaceae) and flax (Linum usitatissimum, Linaceae). Roots were collected in labeled paper bags with moist soil in triplicate from 10 plants 1 m apart from each field. Samples were kept in cool conditions when they were transported to the laboratory where the roots were cleaned with tap water until all soil particles were removed. Once cleaned, roots were wrapped in moist paper towels, placed in a refrigerator (4 C), and processed within 1 wk. Roots were surface sterilized for isolation of endophytes following the technique of Knapp et al. (Citation2012). Each root sample was cut into 2–3-cm segments and placed in 30% H2O2 for 2 min, then in 70% alcohol for 60s, and washed twice in sterile tap water for 2 min. Each root segment was cut into four pieces and placed on potato dextrose agar (PDA). After 4–6 d incubation at room temperature in the dark, hyphae growing from samples were subcultured on PDA. Fusarium isolates (N = 290) were identified based on cultural characteristics (Leslie and Summerell Citation2006) and DNA sequence data from the ITS rDNA region (see below; SUPPLEMENTARY TABLE 1).
Nine isolates of the newly described species were deposited in the CBS Collection of the Westerdijk Fungal Biodiversity Institute, Utrecht, the Netherlands (SUPPLEMENTARY TABLE 2).
DNA isolation and sequencing.—
Total DNA was isolated from the 290 isolates using a modified cetyltrimethylammonium bromide (CTAB) method (Kovács et al. Citation2001; Murray and Thompson Citation1980), a NucleoSpin Plant II DNA isolation kit (Macherey-Nagel, Düren, Germany), or a Quick-DNA Fungal/Bacterial Miniprep Kit (Zymo Research, Orange, California) from mycelium grown and harvested as described by Gargouri et al. (Citation2020).
Published protocols were used to PCR (polymerase chain reaction) amplify and Sanger sequence the ITS rDNA region (White et al. Citation1990) and a portion of translation elongation factor 1-alpha (TEF1; O’Donnell et al. Citation1998b) for the 290 Fusarium isolates. Based on BLASTn queries of the National Center for Biotechnology Information (NCBI) GenBank (https://www.ncbi.nlm.nih.gov/nucleotide/), 16 isolates representing four putative novel Fusarium species from Kazakhstan were analyzed morphologically and subjected to a multilocus molecular phylogenetic analysis that included portions of RNA polymerase II largest (RPB1) and second-largest (RPB2) subunits and calmodulin (CaM) ().
Table 1. Primers used for PCR amplification and sequencing portions of four genes comprising six marker loci.
All PCRs were performed in a 20-μL final volume that included 1 μL of 10 μM forward and reverse primers (Sigma-Aldrich, Sternheim, Germany), 1 µL of 50 ng/µL DNA template, and one of two different commercially available PCR master mixes. With DreamTaq Green PCR Master Mix (Thermo Fisher Scientific, Waltham, Massachusetts) the PCR program included an initial denaturation step of 5 min at 94 C, followed by 35 cycles of 45s at 94 C, 45s at 55 C for ITS and at 57 C for RPB2 and TEF1, and 60s at 72 C, with a final extension step of 10 min at 72 C. For Phusion Green Hot Start II High-Fidelity PCR Master Mix (Thermo Fisher Scientific), PCR cycling times and temperatures were as follows: 98 C for 2 min, followed by 38 cycles of 5 s at 98 C, 5 s at 60 C, and 30s at 72 C, and a final extension step of 72 C for 5 min for the four marker loci.
Isolates chosen for multilocus analysis were screened for mating type idiomorph using the diagnostic primer pairs Fmat1f × Fmat1r for MAT1-1 and Fmat2f × Fmat2r for MAT1-2 (Kerényi et al. Citation2004). PCRs were performed in a 20-µL final volume. Reaction mixes included 1 µL of a 10 μM stock of the forward and reverse primers (Sigma-Aldrich), 10 ng of DNA template, and the recommended amount of DreamTaq Green PCR Master Mix. Amplification conditions included an initial denaturation step for 2 min at 95 C, followed by 30 cycles of 30s at 95 C, 30s at 60 C, and 30s at 72 C, followed by a final extension step of 5 min at 72 C. PCR products were sent to LGC Genomics (Berlin, Germany) for sequencing. Staden Package with pregap4 and gap4 tools was used to edit the electrophoregrams and combine the sequences for phylogenetic analysis (Staden et al. Citation2000). Sequences were deposited in NCBI GenBank (ON951728–ON951743, ON960618–ON960734; see SUPPLEMENTARY TABLE 2).
Phylogenetic analyses.—
ITS rDNA sequences of the collection from Kazakhstan were used to place the 290 isolates in Fusarium species complexes, based on BLASTn queries of NCBI GenBank. The ITS rDNA sequences were not analyzed phylogenetically because the spacer regions are too divergent to align across Fusarium and because paralogous or xenologous ITS2 sequences have been reported in the F. fujikuroi species complex and related lineages (O’Donnell and Cigelnik Citation1997; O’Donnell et al. Citation1998a). Three data sets were analyzed phylogenetically. The first data set included partial TEF1 sequences of the 290 Fusarium isolates from Kazakhstan and 124 reference sequences downloaded from NCBI GenBank (SUPPLEMENTARY FIG. 1). The second data set was represented by 67 partial TEF1, RPB1 (two regions), RPB2 (two regions), and CaM sequences, with sequences of 16 of these representing seven species from Kazakhstan generated in the present study. Sequences for 51 strains that included representatives of 13 species complexes were downloaded from NCBI GenBank. Reference sequences of 36 strains were mined from whole-genome sequences deposited in NCBI GenBank (SUPPLEMENTARY TABLE 2). In addition to the 4-gene combined analysis, the six partitions were analyzed separately. The third data set analyzed phylogenetically included partial TEF1, RPB1, RPB2, and CaM sequences of seven isolates within the F. burgessii species complex from two different regions of Kazakhstan.
DNA sequences were aligned with MAFFT 7 (http://mafft.cbrc.jp/alignment/server/index.html; Katoh and Standley Citation2013), using the E-INS-i method. Alignments were visually inspected and edited in MEGA7 (Kumar et al. Citation2016). Informative sites in the alignments were identified using the DIVEIN Web server (Deng et al. Citation2010). Bayesian inference (BI) analyses were performed with MrBayes 3.1.2 (Ronquist and Huelsenbeck Citation2003) using the GTR+G model of molecular evolution. Four Markov chains were run for 10 million generations, sampling every 1000 generations with a burn-in value set at 4000 sampled trees. Maximum likelihood (ML) phylogenetic analyses were carried out with raxmlGUI 1.3 (Silvestro and Michalak Citation2012; Stamatakis Citation2014). A GTR+G nucleotide substitution model was used for nucleotide partitions with ML estimation of base frequencies. ML bootstrap (BS) analysis with 1000 replicates was used to assess clade support. Phylogenetic trees were visualized and edited in MEGA7 (Kumar et al. Citation2016).
Morphology.—
Colony morphology, color, and odor were characterized on PDA after 8 d incubation under an alternating 12 h dark/12 h near-UV (near-ultraviolet) black and white fluorescent light cycle at 20 C. The optimal temperature for mycelial growth was determined by incubating strains in the dark on PDA at four different temperatures (20, 25, 30, and 35 C). Radial growth was measured in mm daily from the second day post inoculation until the 90-mm Petri dish was fully covered. Micromorphological characters, including shape and size of conidia produced in sporodochia or on the aerial mycelium, shape, size, and mode of formation of microconidia, and chlamydospores, were examined by cultivating strains on synthetic low-nutrient agar (SNA; Nirenberg and O’Donnell Citation1998) overlaid with pieces of sterile filter paper and on carnation leaf agar (CLA; Fisher et al. Citation1982) for 4 wk at 20 C using the same photoperiods described above. Images of sporodochia were obtained using a VHX-5000 (Keyence, Osaka, Japan) digital microscope. Differential interference contrast (DIC) images were taken with a Zeiss Axioskop 2 Plus microscope (Carl Zeiss Microscopy, Göttingen, Germany) equipped with a Zeiss AxioCam ICc5 camera and Zeiss ZEN 2011 software using water as the mounting medium.
RESULTS
A total of 290 Fusarium isolates from asymptomatic agricultural and nonagricultural plant roots in Kazakhstan were placed a species complex using the ITS rDNA sequence data, and species-level identifications were made using partial TEF1 sequences and morphological data. Phylogenetic analysis of the TEF1 sequences suggested that 16 Fusarium species from the following eight species complexes were represented in the collection: F. tricinctum (FTSC; 5 species, N = 95), F. incarnatum-equiseti (FIESC; 3 species, N = 77), F. sambucinum (FSAMSC; 2 species, N = 72), F. redolens (FRSC; 2 species, N = 32), F. burgessii (FBURSC; 1 species, N = 8, plus 3 from Karaganda and Almaty, Kazakhstan, collected by Göksel Özer in a separate pathogen survey, see below), F. oxysporum (FOSC; 1 species, N = 2), F. solani (FSSC; 1 species, N = 1), and F. steppicola lineage (FSTEPSC; 1 species, N = 3), which represents a novel monotypic sister lineage of F. nurragi + the F. heterosporum and F. tricinctum species complexes (SUPPLEMENTARY FIG. 1; SUPPLEMENTARY TABLE 1).
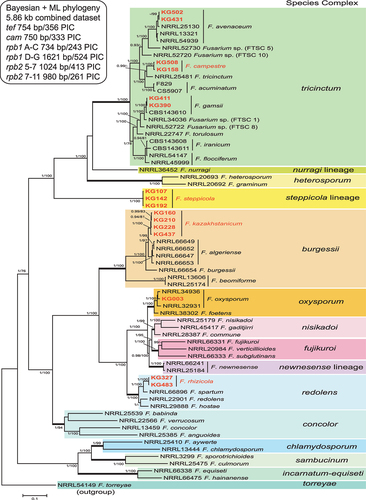
To further assess their species diversity, using phylogenetic species recognition based on genealogical exclusivity (Taylor et al. Citation2000), multilocus DNA sequence data were collected from 16 putatively novel Kazakhstan strains and aligned with reference sequences of 51 phylogenetically diverse fusaria constituting 13 species complexes downloaded from NCBI GenBank (). The aligned 5.86 kb multilocus data set (TreeBASE S29061) included a total of 1774 parsimony-informative characters (PIC) from portions of the following four genes: TEF1 (356 PIC/754 bp alignment), CaM (333 PIC/750 bp alignment), RPB1 A–C (243 PIC/734 bp alignment), RPB1 D–G (524 PIC/1621 bp alignment), RPB2 5–7 (413 PIC/1024 bp alignment), and RPB2 7–11 (261 PIC/980 bp alignment). Bayesian and maximum likelihood phylogenetic analyses of the partial TEF1 sequence data and the 4-gene data sets revealed that the 290 isolates from Kazakhstan were genetically diverse and included representatives of the following eight species complexes, ordered by most to fewest isolates per complex.
Figure 1. Phylogenetic tree of 16 representative Fusarium isolates from Kazakhstan (in bold red) and 51 related reference strains of various Fusarium species. The 50% majority rule consensus phylogram was inferred from Bayesian analysis of the combined data set of six loci (TEF1, CaM, RPB1 A–C, RPB1 D–G, RPB2 5–7, and RPB2 7–11). Bayesian posterior probabilities (≥0.90) and ML bootstrap support (≥70) are shown before and after slashes, respectively. Scale bar indicates 0.1 expected changes per site.
FTSC (N = 95).
The five species included 80 isolates of F. acuminatum recovered from all of the plants sampled in both years, 9 F. iranicum from feather grass and wheat in both years, 2 F. avenaceum from barley grass and feather grass from separate sampling sites in 2019, 2 F. gamsii from oat in 2019, and 2 F. campestre, sp. nov. (KG158 and KG508), from feather grass and wheat in different years, which was previously reported as Fusarium sp. FTSC 12 (; Laraba et al. Citation2022; Senatore et al. Citation2021). Monophyly of the FTSC was well supported by the multilocus analysis and single-locus phylogenies (ML-BS/Bayesian posterior probability [PP] values: all six loci = 100/1, TEF1 = 100/1, CaM = 100/1, RPB1 A–C = 89/0.95, RPB1 D–G = 100/1, RPB2 5–7 = 100/1, and RPB2 7–11 = 100/1; , SUPPLEMENTARY FIG. 2).
FIESC (N = 77).
The three species included 61 isolates of F. clavum (FIESC 5) from the Poaceae sampled, 15 F. equiseti (FIESC 14) from gramineous hosts, and one F. gracilipes (FIESC 13) from barley grass.
FSAMSC (N = 72).
The two species included 63 isolates of F. culmorum from all plants sampled except lentil and nine isolates of F. sporotrichioides from native steppe barley grass (Hordeum leporinum) and feather grass (Stipa capillata) (SUPPLEMENTARY FIG. 1; SUPPLEMENTARY TABLE 1).
FRSC (N = 32).
The two species included 30 isolates of F. redolens from wheat, barley, and lentil together with two isolates of F. rhizicola, sp. nov. (KG327 and KG483), collected in 2019 from lentil and wheat. The latter species was strongly supported as sister to F. spartum (). Monophyly of the FRSC was strongly supported by the multilocus analysis and single-locus phylogenies (ML-BS/Bayesian PP values: all six loci = 100/1, TEF1 = 100/1, CaM = 99/1, RPB1 A–C = 100/1, RPB1 D–G = 100/1, RPB2 5–7 = 100/1, and RPB2 7–11 = 100/1; , SUPPLEMENTARY FIG. 2).
FBURSC (N = 8).
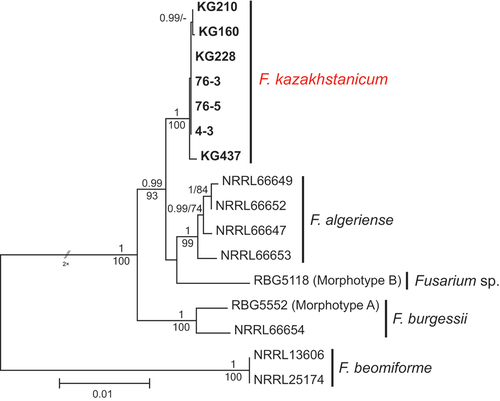
The eight isolates of Fusarium kazakhstanicum, sp. nov. (KG159, KG160, KG210, KG211, KG228, KG231, KG232, and KG437), recovered in the present survey were from feather grass at three different sampling sites in 2018 and from wheat in 2019 (SUPPLEMENTARY TABLE 1). In addition, three isolates of this novel species (4-3, 76-3, 76-5) were collected by GÖ in a separate pathogen survey from the roots of wheat plants showing crown and root rot symptoms in the provinces of Karaganda (4-3) and Almaty (76-3 and 76-5) in 2019. Monophyly of the FBURSC was strongly supported by the multilocus and single-locus phylogenies (ML-BS/Bayesian PP values: all six loci = 100/1, TEF1 = 100/1, CaM = 91/0.98, RPB1 A–C = 99/1, RPB1 D–G = 78/0.95, RPB2 5–7 = 97/0.85, and RPB2 7–11 = 97/0.99; , SUPPLEMENTARY FIG. 2).
Figure 2. Maximum likelihood phylogenetic tree of representative isolates of Fusarium burgessii species complex. Isolates of the novel species from Kazakhstan, F. kazakhstanicum, are shown in bold. RAxML analysis was conducted on combined data set of six loci (TEF1, CaM, RPB1 A–C, RPB1 D–G, RPB2 5–7, and RPB2 7–11). Bayesian posterior probabilities (≥0.90) and ML bootstrap support (≥70) are shown before and after slashes, respectively. Scale bar indicates 0.01 expected changes per site.
FSTEPSC (N = 3).
Three isolates of F. steppicola, sp. nov., were isolated from wheat and feather grass roots (KG107, KG142, and KG192) collected in 2018. This novel monotypic lineage was strongly supported as sister to F. nurragi + the F. tricinctum and F. heterosporum species complexes by the multilocus analysis and single-locus phylogenies (ML-BS/Bayesian PP values: all six loci = 100/1, TEF1 = 100/1, CaM = 100/1, RPB1 A–C = 100/1, RPB1 D–G = 100/1, RPB2 5–7 = 100/1, and RPB2 7–11 = 100/1; , SUPPLEMENTARY FIG. 2).
FOSC (N = 2).
The two F. oxysporum isolates (KG003 and KG214) were recovered from the roots of wheat and feather grass in 2018 and in 2019.
FSSC (N = 1).
Fusarium vanettenii (KG458) was isolated from lentil in 2019.
The 16 isolates constituting seven species from Kazakhstan included in the multilocus phylogenetic analyses were screened for mating type idiomorph using a published PCR assay (Kerényi et al. Citation2004). The two isolates of F. campestre (KG158 and KG508) were MAT1-1, whereas the 14 isolates of the other six species were MAT1-2. Based on the genealogical exclusivity of the novel species within the FTSC, FRSC, FBURSC, and FSTEPSC, the following four novel species are formally described below.
TAXONOMY
Fusarium kazakhstanicum Akhmetova, Özer, D.G. Knapp, Kovács & Molnár, sp. nov.
MycoBank MB842148
Typification: REPUBLIC OF KAZAKHSTAN. AKMOLA REGION: Shortandy District, 51°34ʹ28.8″N, 71°15ʹ53.8″E, originally isolated from the roots of feather or needle grass (Stipa capillata), 13 Sep 2018, G.K. Akhmetova KG160 (holotype 111751BP, a dried culture of KG160, Mycological Collection of the Hungarian Natural History Museum, HNHM-MYC-020593). Ex-holotype culture: KG160 = CBS 148870.
Etymology: kazakhstanicum, for the country of origin, Kazakhstan.
Diagnosis: Distinguished from F. algeriense by slower radial growth on PDA at 25 C in the dark, rare production of chlamydospores, and genealogical exclusivity based on multilocus molecular phylogenetic analyses.
Colony color on PDA white (KG228), beige (KG211), or salmon (KG231, KG437). Odor moldy. Aerial mycelium abundant or sparse, loosely or densely floccose, colony margin entire to undulate. Fastest average radial growth rate on PDA 4–7.3 mm/d at 25 C in dark; 4 mm/d for KG232 and KG437, 4.8 mm/d for KG231, 5.8 mm/d for KG210 and KG211, 6 mm/d for KG160 and KG228, and 7.3 mm/d for KG159. Sporulation on SNA under alternating dark/fluorescent white and black light cycle starting after 5 d, pale orange sporodochia formed after 3 wk by KG159, KG160, and KG210. Pale orange to white-cream sporodochia produced by all observed isolates on carnation leaves under alternating dark/fluorescent white and black light cycle after 3 wk.
Conidiophores borne on aerial mycelium unbranched or branched, bearing terminal or lateral monophialides. Aerial monophialides ellipsoidal, clavate to cylindrical, arise laterally from hyphae. Aerial conidia hyaline, falcate, thin-walled, slightly curved, 1–5-septate, mostly 3-septate, measuring: 1-septate (n = 50) (18–)23–32(–38.5) × (3.5–)5–7(–8.5) μm total range, 27.8 ± 5 × 5.7 ± 0.9 μm average ± SD; 2-septate (n = 20) (27–)35–46(–49) × (4–)5–7(–8.5) μm total range, 40.6 ± 5.2 × 6.2 ± 1 μm average ± SD; 3-septate (n = 150) (35–)44–58(–72) × (5–)5.7–7(–9) μm total range, 41.8 ± 6.9 × 6.5 ± 0.8 μm average ± SD; 4-septate (n = 50) (46–)52–68(–77) × (6–)6.4–7.8(–8.5) μm total range, 59.9 ± 7.8 × 7.2 ± 0.7 μm average ± SD; 5-septate (n = 10) (56–)57–80(–89.5) × (6–)7–8(–8.5) μm total range, 68.7 ± 11.7 × 7.5 ± 0.6 μm average ± SD. Sporodochial conidia falcate, slender, straight to slightly curved, with a papillate apical cell and distinct basal foot cell. Sporodochial conidia typically 3- or 4-septate, measuring: 1-septate (n = 24) (24.5–)28–37(–43) × (4.6)5–6.2(–6.5) μm total range, 32.2 ± 4.9 × 5.7 ± 0.6 μm average ± SD; 2-septate (n = 16) (29)33–48(–59.5) × (4.5–)6.7–8(–8.5) μm total range, 40.7 ± 7.2 × 6.9 ± 1.2 μm average ± SD; 3-septate (n = 147) (34.5–)42–56(–66.5) × (5–)6.3–8(–10) μm total range, 48.9 ± 7 × 7.1 ± 0.9 μm average ± SD; 4-septate (n = 76) (52–)56–67.6(–75) × (5–)6.6–8.4(–10) μm total range, 61.8 ± 5.9 × 7.5 ± 0.9 μm average ± SD; 5-septate (n = 4) (58–)68–73(–90.5) × (6.5–)7–8(–9) μm total range, 72.1 × 7.6 μm average. Microconidia 0- or occasionally 1-septate, ellipsoidal or fusiform in false heads on monophialides, measuring: 0-septate (n = 210) (7–)10–15(–20) × (2.5–)3.8–5.5(–7) μm total range, 12.6 ± 2.8 × 4.6 ± 0.8 μm average ± SD; 1-septate (n = 14) (14.5–)15.3–17.5(–21) × (4–)5–6(–6.5) μm total range, 16.9 ± 1.7 × 5.4 ± 0.8 μm average ± SD. Globose terminal chlamydospores formed sporadically on SNA and CLA after 1 mo, measuring (n = 10) (13.5–)13.5–18(–19.5) × (12–)12.5–15.5(–17) μm total range, 15.3 ± 2.5 × 14.1 ± 1.4 μm average ± SD.
Additional isolates examined: Isolates 4-3, 76-3, 76-5 (NCBI GenBank [TEF1]: ON960732–ON960734) collected from diseased wheat roots from Karaganda and Almaty, Kazakhstan, in 2019 ().
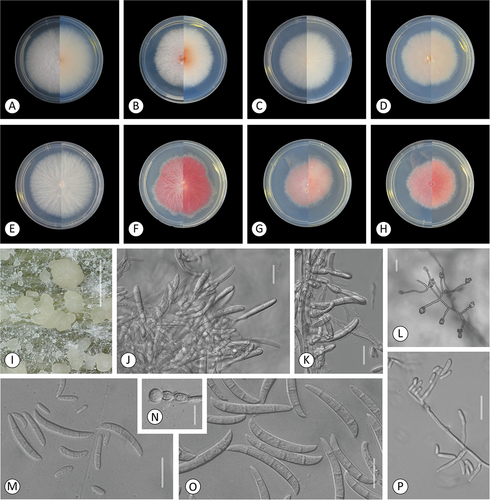
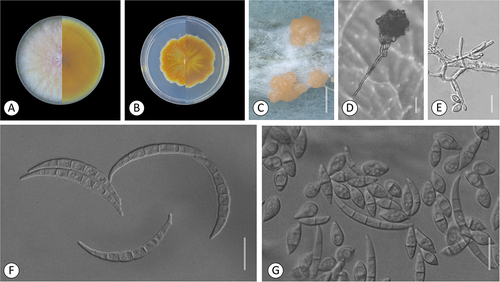
Figure 3. Fusarium kazakhstanicum. A–H. Colony morphology after 8 d growth on PDA using a 12/12 h photoperiod. Colony surface is shown on left half of each plate and colony undersurface on right half. A. KG159. B. KG160. C. KG210. D. KG211. E. KG228. F. KG231. G. KG232. H. KG437. I–P. KG160. I–J. Sporodochia on carnation leaf. K. Aerial conidia formed on monophialides. L, P. Microconidia in false heads on monophialides. M. 0–3-septate aerial conidia. N. Chlamydospores. O. Fusiform, mostly 3-septate sporodochial conidia. Bars: I = 1 mm; J–P = 20 μm.
Fusarium rhizicola Akhmetova, D.G. Knapp, Kovács & Molnár, sp. nov.
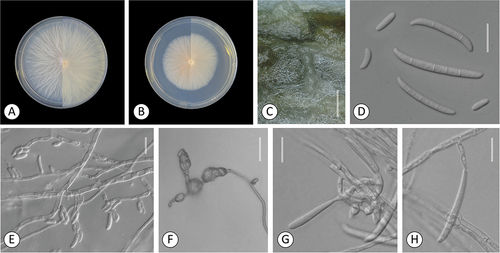
Figure 4. Fusarium rhizicola. A–B. Colony morphology after 8 d growth on PDA using a 12/12 h photoperiod. Colony surface is shown on left half of each plate and colony undersurface on right half. A. KG327. B. KG483. C. Sporodochia of KG327 formed on carnation leaf. D–H. KG483. D. 0-septate and multiseptate aerial conidia. E–F. Microconidia in false heads on monophialides. G–H. Aerial conidia developed on monophialides. Bars: C = 1 mm; D–H =20 μm.
MycoBank MB842149
Typification: REPUBLIC OF KAZAKHSTAN. AKMOLA REGION: Shortandy District, 51°38ʹ36.50″N, 71°01ʹ05.00″E, originally isolated from wheat roots (Triticum aestivum), 12 Aug 2019, G.K. Akhmetova KG483 (holotype 111753BP, a dried culture of KG483, Mycological Collection of the Hungarian Natural History Museum, HNHM-MYC-020595). Ex-holotype culture: KG483 = CBS 148872.
Diagnosis: Distinguished from F. spartum by slower radial growth on PDA at 25 C in dark, absence of chlamydospores, and genealogical exclusivity based on multilocus molecular phylogenetic analyses.
Colony color under alternating dark/fluorescent white and black light cycle at 20 C surface and reverse white. Odor moldy. Aerial mycelium sparse or abundant, commonly forming hyphal bundles, colony margin entire. Optimal growth rate on PDA at 25 C in dark 5.8 mm/d (KG483) or 7 mm/d (KG327).
Conidiophores borne on aerial mycelium unbranched or branched, bearing terminal or lateral monophialides. Aerial conidia hyaline, falcate, thin-walled, straight with even width, apical cells curved, basal cells with blunt end, 1–4-septate, mostly 3-septate, measuring: 1-septate (n = 8) (19.5–)25–30(–49) × (5–)5.2–5.8(–6) μm total range, 28.8 × 5.6 μm average; 2-septate (n = 9) (31.5–)35–49(–51.5) × (5–)5.5–7(–8) μm total range, 42.9 × 6.3 μm average; 3-septate (n = 16) (44.5–)49–58(–71) × (6–)6.2–7.4(–8) μm total range, 54.7 ± 6.7 × 6.9 ± 0.6 μm average ± SD; 4-septate (n = 1) 67.4 × 6.2 μm. Sporodochial conidia typically 3-septate, measuring: 2-septate (n = 4) 31.8–40.5 × 4.6–7.8 μm total range, 35.9 × 6.6 μm average; 3-septate (n = 23) (38–)41–50(–54.5) × (4.9–)5.7–7(–8) μm total range, 46.4 ± 5.3 × 6.6 ± 0.7 μm average ± SD; 4-septate (n = 6) (52–)54–56(–57) × (6.5–)7–8(–8.5) μm total range, 54.9 × 7.5 μm average. Microconidia produced in false heads, 0- or 1-septate, mostly 0-septate, fusiform, ellipsoidal or reniform, measuring: (n = 43) (7.5–)11–18(–20) × (4–)4.5–6(–7.5) μm total range, 13.8 ± 3.4 × 5.2 ± 0.7 μm average ± SD; 1-septate (n = 19) (15.5–)18–21(–22.5) μm total range, 19.6 ± 2.0 × 5.4 ± 0.6 μm average ± SD. Chlamydospores not observed.
Fusarium campestre Akhmetova, D.G. Knapp, Kovács & Molnár, sp. nov.
Figure 5. Fusarium campestre (FTSC 12). A–B. Colony morphology after 8 d growth on PDA using a 12/12 h photoperiod. Colony surface is shown on left half of each plate and colony undersurface on right half. A. KG158. B. KG508. C-G. KG158. C. Sporodochia on carnation leaf. D. Microconidia in false head on a long conidiophore. E. Conidiophores producing citriform microconidia. F. Multiseptate fusiform sporodochial conidia. G. 0- to 1-septate and multiseptate aerial and sporodochial conidia, respectively. Bars: C = 1 mm; D–G = 20 μm.
MycoBank MB842150
Typification: REPUBLIC OF KAZAKHSTAN. AKMOLA REGION: Shortandy District, 51°34ʹ28.8″N, 71°15ʹ53.8″E, originally isolated from the roots of feather or needle grass (Stipa capillata), 13 Sep 2018, G.K. Akhmetova KG158 (holotype 111750BP, a dried culture of KG158, Mycological Collection of the Hungarian Natural History Museum, HNHM-MYC-020592). Ex-holotype culture: KG158 = CBS 148994.
Etymology: campestre, from the Latin noun “campester” (= grassland), referring to the habitat of the isolates.
Diagnosis: Distinguished from sister species, F. tricinctum, by 5-septate sporodochial conidia compared with those of F. tricinctum that are typically 3-septate, and genealogical exclusivity based on multilocus molecular phylogenetic analyses.
Colony color under alternating dark/fluorescent white and black light cycle at 20 C surface white, rosy, and yellow or ochrous, reverse ochrous and light brown. Odor moldy. Aerial mycelium abundant throughout the colony, sparse or loosely floccose. Colony margin irregular. Optimal growth rate on PDA at 25 C in dark 4.5 mm/d (KG508) or 8.5 mm/d (KG158). Sporodochia orange, formed on carnation leaves on CLA after 3 wk (KG158) or not observed (KG508).
Conidiophores borne on aerial mycelium unbranched or branched, bearing terminal or lateral monophialides. Aerial conidia hyaline, falcate, thin-walled, curved, usually with a foot-shaped basal cell, 3–6-septate, mostly 3-septate, measuring: 3-septate (n = 16) (28–)30–41(–44.5) × (5–)5.5–7(–8.5) μm total range, 36 ± 5.7 × 6.2 ± 0.9 μm average ± SD; 4-septate (n = 5) (46–)48–59(–60.5) × (6–)6.5–7.5(–8) μm total range, 53.5 × 6.9 μm average; 5-septate (n = 9) (52–)55–59(–69) × (5.5–)6–6.7(–7) μm total range, 58.8 × 6.4 μm average; 6-septate (n = 1) 61.8 × 6.9 μm. Sporodochial conidia hyaline, thin-walled, long and slender, distinctly curved to lunate. Apical cells long, slightly curved and tapering, basal cells often foot-shaped. Sporodochial conidia 3–5-septate, mostly 5-septate, measuring: 3-septate (n = 6) (51.5–)53–61(–69) × (6–)6.7–7.7(–8.5) μm total range, 60.1 × 7.3 μm average; 4-septate (n = 23) (61–)67–71(–77) × (6.5–)7–8.3(–9.5) μm total range, 69 ± 3.9 × 7.7 ± 0.8 μm average ± SD; 5-septate (n = 47) (60.5–)67–75(–80.5) × (6–)7–9(–9.5) μm total range, 70.8 ± 4.3 × 7.9 ± 0.8 μm average ± SD. Microconidia usually 0- or 1-septate, occasionally 2- or 3-septate, oval or fusoid but usually napiform or citriform in false heads on monophialides, measuring: 0-septate oval-fusoid (n = 8) (13–)14–17(–19) × (4–)5–7(–7.5) μm total range, 16 × 5.5 μm average; 0-septate napiform (n = 31) (10.5–)14–18(–20) × (6.5–)7–9(–10) μm total range, 15.8 ± 1.8 × 8 ± 1.1 μm average ± SD; 1-septate oval-fusoid (n = 14) (22.5–)25–28(–31.5) × (5–)5.5–6.5(–7) μm total range, 27 ± 2.8 × 5.9 ± 0.5 μm average ± SD; 1-septate napiform (n = 33) (14–)16–20(–22) × (6.5–)7.5–9.5(–11) μm total range, 17.7 ± 1.9 × 8.4 ± 0.9 μm average ± SD; 2-septate oval-fusoid (n = 1) 39.5 × 6.7 μm; 2-septate napiform (n = 12) (17.5–)20–27(–31) × (8.5–)8.8–10(–12) μm total range, 24.4 ± 3.9 × 9.6 ± 1.1 μm average ± SD; 3-septate napiform (n = 3) 21.5–30.3 × 7.1–11.5 μm total range, 26.6 × 8.7 μm average. Chlamydospores not observed.
Fusarium steppicola Akhmetova, D.G. Knapp, Kovács & Molnár, sp. nov.
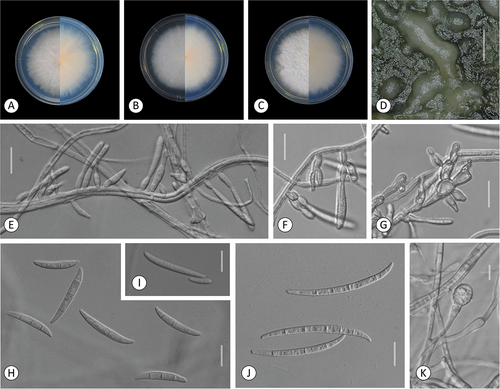
Figure 6. Fusarium steppicola. A–C. Colony morphology after 8 d growth on PDA using a 12/12 h photoperiod. Colony surface shown on left half of each plate and colony undersurface on right half. A. KG107. B. KG142. C. KG192. D–K. KG160. D. Sporodochia formed on carnation leaf. E–G. Aerial conidia formed on monophialides. H–I. Fusiform multiseptate aerial conidia. J. Fusiform multiseptate sporodochial conidia. K. Terminal chlamydospore. Bars: D = 0.5 mm; E–K =20 μm.
MycoBank MB842151
Typification: REPUBLIC OF KAZAKHSTAN. AKMOLA REGION: Shortandy District: 51°38ʹ12.2″N, 71°01ʹ16.7″E, originally isolated from the roots of wheat (Triticum aestivum), 12 Sep 2018, G.K. Akhmetova KG107 (holotype 111752BP, a dried culture of KG107, Mycological Collection of the Hungarian Natural History Museum, HNHM-MYC-020594). Ex-holotype culture: KG107 = CBS 148867.
Etymology: steppicola refers to the steppe habitat.
Diagnosis: Sporodochial conidia are similar in shape to those formed by F. avenaceum but are much longer. It produces abundant chlamydospores, whereas F. avenaceum does not. It demonstrates genealogical exclusivity based on multilocus molecular phylogenetic analyses.
Colony color under alternating dark/fluorescent white and black light cycle at 20 C surface white, reverse pale orange. Odor moldy. Aerial mycelium abundant throughout the colony, loosely (KG107) or densely (KG142, KG192) floccose, colony margin entire. Optimal growth rate on PDA in the dark 5.5 mm/d for KG142, 6.3 mm/d for KG192, and 6.7 mm/d for KG107 at 25 C. KG107 sporulated abundantly, KG142 occasionally, conidia of KG192 not observed. Cream-colored sporodochia observed only by KG107 after 2 wk on carnation leaves.
Conidiophores borne on aerial mycelium unbranched or branched, bearing terminal or lateral monophialides. Aerial conidia hyaline, ellipsoidal to falcate, thin-walled, dorsal side more curved than the ventral, typically widest in the upper half, 0–4-septate, mostly 3-septate, measuring: 0-septate (n = 6) (13–)14–19(–22) × (5–)5.6–6.5(–7) μm total range, 17.3 × 6 μm average; 1-septate (n = 36) (17–)20–32(–47) × (4.5–)5.5–6.5(–8) μm total range, 26.4 ± 6.4 × 6.5 ± 0.9 μm average ± SD; 2-septate (n = 5) (37–)41–44(–45) × (6.5)7–8(–9) μm total range, 42 × 7.5 μm average; 3-septate (n = 62) (39–)46–60(–66) × (6.5–)8–9.5(–10) μm total range, 54.1 ± 5.5 × 8.5 ± 0.9 μm average ± SD; 4-septate (n = 2) 64–69.5 × 9.2–9.8 μm total range, 66.7 × 9.5 μm average. Sporodochial conidia hyaline, thin-walled, long and slender, slightly curved, widest in the upper half. Apical cells long, slightly curved. Basal cells notched. Sporodochial conidia 3–5-septate, mostly 4-septate, measuring: 3-septate (n = 8) (65–)73–84(–85.5) × (6–)7–8(–8.5) μm total range, 78.2 ± 7.7 × 7.3 ± 0.9 μm average ± SD; 4-septate (n = 34) (81–)90–98(–107.5) μm total range, 93.3 ± 6.1 × 8.4 ± 0.9 μm average ± SD; 5-septate (n = 17) (87.5–)92–100(–108.5) μm total range, 96.0 ± 6.3 × 8.9 ± 1.0 μm average ± SD. Chlamydospores produced abundantly on SNA and CLA after 7 d mostly terminal, formed singly, in pairs or occasionally in chains. Chlamydospores globose to subglobose with slightly roughened surface, measuring (n = 24) (10–)11–16(–19) × (10–)11–15(–19) μm total range, 13 × 15 μm average ± SD.
DISCUSSION
When root-colonizing endophytic fungi of agricultural and nonagricultural plant roots from the northern steppe region of Kazakhstan were surveyed, 290 Fusarium isolates representing 16 different species constituting eight known species complexes and one distinct lineage were recovered and identified using partial TEF1 sequence data. Four novel species were discovered and formally described, including F. steppicola, which represents a phylogenetically divergent lineage that is strongly supported as sister to F. nurragi + the F. tricinctum and F. heterosporum species complexes. Although Fusarium species are frequently isolated from asymptomatic root tissues of different gramineous hosts in grassland communities worldwide (Khidir et al. Citation2010; Knapp et al. Citation2012, Citation2019), prior to the present study, knowledge of Fusarium species in Kazakhstan was limited to two surveys of cereals (Kaumenov et al. Citation2017; Nugmanov et al. Citation2018). In the present survey, most of the Fusarium isolates (276/290, 95.2%) belonged to four species complexes: FTSC (95/290 = 32.8%), FIESC (77/290 = 26.6%), FSAMSC (72/290 = 24.8%), and FRSC (32/290 = 11%).
Fusarium acuminatum accounted for most of the FTSC strains (80/95) and was isolated at the majority of the sampling sites. This species is considered a saprobic endophyte of wild grasses (Özer et al. Citation2020a; Szécsi et al. Citation2013; Tunali et al. Citation2008). Other species of the FTSC recovered in our survey included F. avenaceum, F. iranicum, F. gamsii, and F. campestre, which is formally described here from wheat and feather grass. This species was previously reported as Fusarium sp. FTSC 12 (Laraba et al. Citation2022; O’Donnell et al. Citation2009; Senatore et al. Citation2021), as sister to F. tricinctum. Fusarium iranicum and F. gamsii were recently described as fungicolous species on Agaricus bisporus (Torbati et al. Citation2019), whereas F. iranicum was isolated as an endophyte of wheat and feather grass and F. gamsii from oat. Bottalico and Perrone (Citation2002) reported that F. avenaceum, F. graminearum, and F. culmorum were the dominant species in Fusarium head blight (FHB)-affected wheat and other small-grain cereals. Szécsi et al. (Citation2013) isolated F. avenaceum from the base of asymptomatic wild grasses, whereas it was recovered from feather and barley grass in the present study.
The FIESC was the second-most common species complex recovered in the present study, represented by three species accounting for 77/290 (= 26.6%) of the Fusarium isolates. These included Fusarium clavum (61/77; Fusarium sp. FIESC 5 sensu Villani et al. Citation2019), F. equiseti (15/77; Fusarium sp. FIESC 15 sensu Villani et al. Citation2019), and F. gracilipes (1/77; Fusarium sp. FIESC 13 sensu Villani et al. Citation2019). The F. clavum isolates were recovered from cereals and wild grasses, whereas previous studies identified this cosmopolitan species from wheat, barley, oat, and rice (Villani et al. Citation2016), Solanum tuberosum (Xia et al. Citation2019), and humans (O’Donnell et al. Citation2009). The host range and geographic distribution of F. equiseti should be interpreted with caution because this name has been misapplied to the sequences of many species within the FIESC deposited in NCBI GenBank (O’Donnell et al. Citation2015, Citation2022). Our 15 F. equiseti isolates were from wheat, barley, barley grass, and feather grass. The single isolate of F. gracilipes was recovered from barley grass, whereas previous studies identified F. equiseti from diverse cereals (i.e., barley, wheat, maize, and oat), humans (O’Donnell et al. Citation2009), and soil, S. tuberosum, and Daphne mezereum (Villani et al. Citation2016; Xia et al. Citation2019).
The FSAMSC was represented by 63 isolates of F. culmorum and nine of F. sporotrichioides, which are known to produce type B and A trichothecene mycotoxins, respectively (Laraba et al. Citation2021). Fusarium culmorum was isolated from asymptomatic flax, wheat, barley, oat, barley, and feather grass. This species is the main causal agent of FHB of wheat and other small-grain cereals in cooler regions of Europe (Bottallico and Perrone Citation2002), and FHB and Fusarium crown rot in Algeria (Laraba et al. Citation2017a). The F. sporotrichioides isolates in our survey were isolated from barley and feather grass. Szécsi et al. (Citation2013) isolated this species from the asymptomatic stems of two wild grasses (Hordeum murinum and Sesleria budensis). Members of the Sporotrichioides Clade colonize a wide range of cereals, wild grasses, soil, and debris (Laraba et al. Citation2021) and are less aggressive secondary invaders of FHB-diseased cereal heads (Bottalico and Perrone Citation2002).
Two species within the FRSC were recovered in our survey, including 30 isolates of F. redolens from wheat, barley, and lentil. In previous studies, F. redolens was reported as a pathogen of diverse plants, causing wilts, seedling damping-off, and cortical rot (Baayen et al. Citation2001; Gargouri et al. Citation2020), whereas Su et al. (Citation2010) discovered F. redolens to be an endophyte of asymptomatic Stipa grandis roots in the Inner Mongolia steppe of China. The novel species within the FRSC was described as F. rhizicola, which was represented by two isolates from wheat and lentil. This species was strongly supported as sister to F. spartum, which is an endophyte of needle grass in Tunisia (Gargouri et al. Citation2020).
Although F. oxysporum is the most reported endophytic partner of cultivated and noncultivated plants, including the roots of many poaceous species (Edel et al. Citation1997; Maciá-Vicente et al. Citation2008; Su et al. Citation2010), it was only represented by two isolates from feather grass and wheat in our collection. Members of the FOSC are considered one of the top ten pathogens in plant pathology (Dean et al. Citation2012), causing wilt and stem rot diseases of many agriculturally important plants (e.g., Panama disease of banana and vascular wilt of tomato; O’Donnell et al. Citation1998b; Summerell Citation2019).
Fusarium kazakhstanicum, described here as a novel species within the FBURSC from the Akmola Region of northern Kazakhstan, was also isolated from diseased wheat roots in the Karaganda and Almaty regions in a separate pathogen survey in collaboration with the International Maize and Wheat Improvement Center (CIMMYT). Fusarium kazakhstanicum was strongly supported as sister to F. algeriense, which is a crown rot pathogen of durum wheat in Algeria (Laraba et al. Citation2017b) and winter wheat in Azerbaijan (Özer et al. Citation2020b). The three other species within the FBURSC (F. burgessii, F. beomiforme, and Fusarium sp. Morphotype B) were isolated from soils or soil debris (Laurence et al. Citation2011; LeBlanc et al. Citation2017; Nelson et al. Citation1987) and are not known to be phytopathogenic.
One isolate of Fusarium vanettenii, formerly known as F. solani f. sp. pisi and Nectria haematococca mating population MPVI (Coleman et al. Citation2009), from lentil was the sole representative of the FSSC. Comparative phylogenomic analyses strongly support the inclusion of the FSSC in a monophyletic Fusarium (Geiser et al. Citation2021; O’Donnell et al. Citation2020).
One of the more important results of the present study was discovery of the monotypic Fusarium steppicola lineage, which we posit will be recognized as a novel species complex once a closely related sister species is recovered and characterized in future endophyte and pathogen surveys. Currently, it was resolved as a phylogenetically divergent sister to F. nurragi + the F. tricinctum and F. heterosporum species complexes. This novel species was represented by three isolates, two from wheat roots (KG107, KG142) and the third from feather grass (KG192).
Future studies are needed to determine whether F. steppicola and the three other novel species described herein are pathogenic to cereals and assess their potential to produce mycotoxins in vitro and via comparative genomics (Kim et al. Citation2020; Laraba et al. Citation2021). Based on what is known about the toxin potential of their near relatives, we hypothesize that F. steppicola and F. campestre might produce aurofusarin, enniatins, and moniliformin (Laraba et al. Citation2022; O’Donnell et al. Citation2018), F. rhizicola beauvericin and enniatins (Gargouri et al. Citation2020), and F. kazakhstanicum beauvericin, bikaverin, and fusarin (Laraba et al. Citation2017a).
Multilocus GCPSR-based and detailed morphological analyses are crucial in biodiversity research and applied science, especially in Fusarium because it represents the most important genus of mycotoxigenic plant pathogens (Geiser et al. Citation2021). Fusarium is currently estimated to comprise at least 450 phylospecies (O’Donnell et al. Citation2022), but this likely represents only a fraction of their global phylogenetic diversity based on estimates that fewer than 10% of fungi on planet Earth have been discovered (Blackwell Citation2011; Hawksworth and Lücking Citation2017). These estimates appear to be supported by our discovery and formal recognition of four novel fusaria among the culturable endophytes of healthy roots of agricultural and nonagricultural plants in Kazakhstan.
Supplemental Material
Download Zip (4.3 MB)ACKNOWLEDGMENTS
The authors thank the A. I. Barayev “Scientific Production Centre for Grain Farming” for the official permission to collect samples from the fields and to conduct this research. Sampling in the Karaganda and Almaty provinces was supported by the International Maize and Wheat Improvement Center (CIMMYT) with assistance from Rauan Zhapayev (CIMMYT-Kazakhstan) and Abdelfattah A. S. Dababat (CIMMYT-Turkey). The authors thank Andras Molnár for preparing the photographic plates.
DISCLOSURE STATEMENT
No potential conflict of interest was reported by the author(s).
SUPPLEMENTARY MATERIAL
Supplemental data for this article can be accessed online at https://doi.org/10.1080/00275514.2022.2119761.
Additional information
Funding
LITERATURE CITED
- Akhmetova G, Knapp DG, Ashrafi S, Maier W, Molnár O, Kovács GM. 2021. Fungal root endophytes from Northern Kazakhstan – novel lineages and dominant core members. Acta Microbiol Immunol Hung. 68:54.
- Audenaert K, Van Broeck R, Bekaert B, De Witte F, Heremans B, Messens K, Höfte M, Haesaert G. 2009. Fusarium head blight (FHB) in Flanders: population diversity, inter-species associations and DON contamination in commercial winter wheat varieties. Eur J Plant Pathol. 125:445–458.
- Baayen RP, O’Donnell K, Breeuwsma S, Geiser DM, Waalwijk C. 2001. Molecular relationships of fungi within the Fusarium redolens–F. hostae clade. Phytopathology. 91:1037–1044.
- Blackwell M. 2011. The fungi: 1, 2, 3 … 5.1 million species? Am J Bot. 98:426–438.
- Bottalico A, Perrone G. 2002. Toxigenic Fusarium species and mycotoxins associated with head blight in small-grain cereals in Europe. Eur J Plant Pathol. 108:611–24.
- Coleman JJ, Rounsley SD, Rodriguez-Carres M, Kuo A, Wasmann CC, Grimwood J, Schmutz J, Taga M, White GJ, Zhou S, et al. 2009. The genome of Nectria haematococca: contribution of supernumerary chromosomes to gene expansion. PLoS Genet. 5(8):e1000618.
- Dean R, van Kan JAL, Pretorius ZA, Hammond-Kosack KE, Di Pietro A, Spanu PD, Rudd JJ, Dickman M, Kahmann R, Ellis J, et al. 2012. The top 10 fungal pathogens in molecular plant pathology. Mol Plant Pathol. 13:414–430.
- Deng W, Maust BS, Nickle DC, Learn GH, Liu Y, Heath L, Kosakovsky Pond SL, Mullins JI. 2010. DIVEIN: a web server to analyze phylogenies, sequence divergence, diversity, and informative sites. BioTechniques. 48:405–408.
- Edel V, Steinberg C, Gautheron N, Alabouvette C. 1997. Populations of nonpathogenic Fusarium oxysporum associated with roots of four plant species compared to soilborne populations. Phytopathology. 87:693–697.
- FAOSTAT. 2021. Food and Agriculture Organization of the United Nations Statistics Division. [ accessed 2021 Jul 11]. http://faostat3.fao.org/download/Q/QC/E.
- Fisher NL, Burgess LW, Toussoun TA, Nelson PE. 1982. Carnation leaves as a substrate and for preserving cultures of Fusarium species. Phytopathology. 72:151–153.
- Gardes M, White TJ, Fortin JA, Bruns TD, Taylor JW. 1991. Identification of indigenous and introduced symbiotic fungi in ectomycorrhizae by amplification of nuclear and mitochondrial ribosomal DNA. Can J Bot. 69:180–190.
- Gargouri S, Balmas V, Burgess L, Paulitz T, Laraba I, Kim H-S, Proctor RH, Busman M, Felker FC, Murray T, et al. 2020. An endophyte of Macrochloa tenacissima (esparto or needle grass) from Tunisia is a novel species in the Fusarium redolens species complex. Mycologia. 112:792–807.
- Geiser DM, Al-Hatmi A, Aoki T, Arie T, Balmas V, Barnes I, Bergstrom GC, Bhattacharyya MKK, Blomquist CL, Bowden R, et al. 2021. Phylogenomic analysis of a 55.1 kb 19-gene dataset resolves a monophyletic Fusarium that includes the Fusarium solani species complex. Phytopathology. 111:1064–1079.
- Geiser DM, Jiménez-Gasco MM, Kang S, Makalowska I, Veeraraghavan N, Ward TJ, Zhang N, Kuldau GA, O’Donnell K. 2004. FUSARIUM-ID v. 1.0: a DNA sequence database for identifying Fusarium. Eur J Plant Pathol. 110:473–479.
- Hawksworth DL, Lücking R. 2017. Fungal diversity revisited: 2.2 to 3.8 million species. Microbiol Spectr. 5:1-17.
- Jumpponen A, Herrera J, Porras-Alfaro A, Rudgers J. 2017. Biogeography of root-associated fungal endophytes. In:Tedersoo L, editor. Biogeography of Mycorrhizal Symbiosis. Cham, Switzerland: Springer; p. 195–222.
- Katoh K, Standley DM. 2013. MAFFT: multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol. 30:772–780.
- Kaumenov NS, Beyshova IS, Kovalchuk AM. 2017. Identification of Fusarium fungi in grain crops in Northern Kazakhstan by real time PCR (In Russian), and Kotsur VP, ed. XLІІ International Scientific and Practical Internet Conference “Problems and Prospects of Scientific Development at the Beginning of the Third Millennium in European and Asian countries,” Pereyaslav-Khmelnytskyi, Ukraine, Sept 29–30. p. 12–14. http://conferences.neasmo.org.ua/uploads/conference/file/45/conference_29-30.9.2017.pdf
- Kavroulakis N, Doupis G, Papadakis IE, Ehaliotis C, Papadopoulou KK. 2018. Tolerance of tomato plants to water stress is improved by the root endophyte Fusarium solani FsK. Rhizosphere. 6:77–85.
- Kerényi Z, Moretti A, Waalwijk C, Oláh B, Hornok L. 2004. Mating type sequences in asexually reproducing Fusarium species. Appl Environ Microbiol. 70:4419–4423.
- Khidir HH, Eudy DM, Porras-Alfaro A, Herrera J. 2010. A general suite of fungal endophytes dominate the roots of two dominant grasses in a semiarid grassland. J Arid Environ. 74:35–42.
- Kia SH, Glynou K, Nau T, Thines M, Piepenbring M, Maciá-Vicente JG. 2017. Influence of phylogenetic conservatism and trait convergence on the interactions between fungal root endophytes and plants. ISME J. 11:777–790.
- Kim H-S, Lohmar JM, Busman M, Brown DW, Naumann TA, Divon HH, Lysøe E, Uhlig S, Proctor RH. 2020. Identification and distribution of gene clusters required for synthesis of sphingolipid metabolism inhibitors in diverse species of the filamentous fungus Fusarium. BMC Genomics. 21:510.
- Klein I, Gessner U, Kuenzer C. 2012. Regional land cover mapping and change detection in Central Asia using MODIS time-series. Appl Geogr. 35:219–234.
- Knapp DG, Imrefi I, Boldpurev E, Csíkos S, Akhmetova G, Berek-Nagy PJ, Otgonsuren B, Kovács GM. 2019. Root-colonizing endophytic fungi of the dominant grass Stipa krylovii from a Mongolian steppe grassland. Front Microbiol. 10:2565.
- Knapp DG, Németh JB, Barry K, Hainaut M, Henrissat B, Johnson J, Kuo A, Lim JHP, Lipzen A, Nolan M, et al. 2018. Comparative genomics provides insights into the lifestyle and reveals functional heterogeneity of dark septate endophytic fungi. Sci Rep. 8:6321.
- Knapp DG, Pintye A, Kovács GM. 2012. The dark side is not fastidious– dark septate endophytic fungi of native and invasive plants of semiarid sandy areas. PLoS One. 7:e32570.
- Kovács GM, Rudnóy S, Vágvölgy C, Lásztity D, Rácz I, Bratek Z. 2001. Intraspecific invariability of the ITS region of rDNA of Terfezia terfezioides in Europe. Folia Microbiol. 46:423–426.
- Kuldau GA, Yates IE. 2000. Evidence for Fusarium endophytes in cultivated and wild plants. In: Bacon CW, White JF, editors. Microbial endophytes. New York, NY: Marcel Dekker; p. 85–120.
- Kumar S, Stecher G, Tamura K. 2016. MEGA7: molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol Biol Evol. 33:1870–1874.
- Laraba I, Boureghda H, Abdallah N, Bouaicha O, Obanor F, Moretti A, Geiser DM, Kim H-S, McCormick SP, Proctor RH, et al. 2017a. Population genetic structure and mycotoxin potential of the wheat crown rot and head blight pathogen Fusarium culmorum in Algeria. Fungal Genet Biol. 103:34–41.
- Laraba I, Busman M, Geiser DM, O’Donnell K. 2022. Phylogenetic diversity and mycotoxin potential of emergent phytopathogens within the Fusarium tricinctum species complex. Phytopathology. 112:1284–1298.
- Laraba I, Keddad A, Boureghda H, Abdallah N, Vaughan MM, Proctor RH, Busman M, O’Donnell K. 2017b. Fusarium algeriense, sp. nov., a novel toxigenic crown rot pathogen of durum wheat from Algeria is nested in the Fusarium burgessii species complex. Mycologia. 109:935–950.
- Laraba I, McCormick SP, Vaughan MM, Geiser DM, O’Donnell K. 2021. Phylogenetic diversity, trichothecene potential, and pathogenicity within Fusarium sambucinum species complex. PLoS ONE. 16:e0245037.
- Laurence MH, Summerell BA, Burgess LW, Liew ECY. 2011. Fusarium burgesii sp. nov. representing a novel lineage in the genus Fusarium. Fungal Div. 49:101–12.
- LeBlanc N, Kinkel L, Kistler HC. 2017. Plant diversity and plant identity influence Fusarium communities in soil. Mycologia. 109:128–139.
- Leslie JF, Summerell BA. 2006. The Fusarium laboratory manual. Iowa, USA: Blackwell Professional Publishing; p. 388.
- Li X, He X, Hou L, Ren Y, Wang S, Su F. 2018. Dark septate endophytes isolated from a xerophyte plant promote the growth of Ammopiptanthus mongolicus under drought condition. Sci Rep. 8:7896.
- Liu YJ, Whelen S, Hall BD. 1999. Phylogenetic relationships among ascomycetes: evidence from an RNA polymerase II subunit. Mol Biol Evol. 16:1799–1808.
- Maciá-Vicente JG, Jansson H-B, Abdullah SK, Descals E, Salinas J, Lopez-Llorca LV. 2008. Fungal root endophytes from natural vegetation in Mediterranean environments with special reference to Fusarium spp. FEMS Microbiol Ecol. 64:90–105.
- Mandyam K, Loughin T, Jumpponen A. 2010. Isolation and morphological and metabolic characterization of common endophytes in annually burned tallgrass prairie. Mycologia. 102:813–821.
- Mermut AR. 1999. Impact of management systems on crop production in the Eurasian steppes of Kazakhstan. In: Karabayev M, Satybaldin A, Benites J, Friedrich T, Pala M, Payne T, eds. Conservation tillage: a viable option for sustainable agriculture in Central Asia. Proceedings of an international workshop Shortandy-Astana, Republic of Kazakhstan, September 19-24.
- Murray MG, Thompson WF. 1980. Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res. 8:4321–4326.
- Nelson PE, Toussoun TA, Burgess LW. 1987. Characterization of Fusarium beomiforme sp. nov. Mycologia. 79:884–889.
- Newsham KK. 2011. A meta-analysis of plant responses to dark septate root endophytes. New Phytol. 190:783–793.
- Nirenberg HI, O’Donnell K. 1998. New Fusarium species and combinations within the Gibberella fujikuroi species complex. Mycologia. 90:434–458.
- Nugmanov A, Beishova I, Kokanov S, Lozowicka B, Kaczynski P, Konecki R, Snarska K, Wołejko E, Sarsembayeva N, Abdigaliyeva T. 2018. Systems to reduce mycotoxin contamination of cereals in the agricultural region of Poland and Kazakhstan. Crop Prot. 106:64–71.
- O’Donnell K, Al-Hatmi AMS, Aoki T, Brankovics B, Cano-Lira JF, Coleman JJ, de Hoog GS, Di Pietro A, Frandsen RJN, Geiser DM, et al. 2020. No to Neocosmospora: phylogenomic and practical reasons for continued inclusion of the Fusarium solani species complex in the genus Fusarium mSphere. mSphere. 5:e00810–20. DOI:10.1128/mSphere.00810-20.
- O’Donnell K, Cigelnik E. 1997. Two divergent intragenomic rDNA ITS2 types within a monophyletic lineage of the fungus Fusarium are nonorthologous. Mol Phylogenet Evol. 7:103–116.
- O’Donnell K, Cigelnik E, Nirenberg HI. 1998a. Molecular systematics and phylogeography of the Gibberella fujikuroi species complex. Mycologia. 90:465–493.
- O’Donnell K, Kistler HC, Cigelnik E, Ploetz RC. 1998b. Multiple evolutionary origins of the fungus causing Panama disease of banana: concordant evidence from nuclear and mitochondrial gene genealogies. Proc Nat Acad Sci USA. 95:2044–2049.
- O’Donnell K, McCormick SP, Busman M, Proctor RH, Ward TJ, Doehring G, Geiser DM, Alberts JF, Rheeder JP. 2018. Marasas et al. 1984 “Toxigenic Fusarium species: identity and Mycotoxicology” revisited. Mycologia. 110:1058–1080.
- O’Donnell K, Nirenberg HI, Aoki T, Cigelnik E. 2000. A multigene phylogeny of the Gibberella fujikuroi species complex: detection of additional phylogenetically distinct species. Mycoscience. 41:61–78.
- O’Donnell K, Sutton DA, Rinaldi MG, Gueidan C, Crous PW, Geiser DM. 2009. Novel multilocus sequence typing scheme reveals high genetic diversity of human pathogenic members of the Fusarium incarnatum-equiseti and F. chlamydosporum species complexes within the United States. J Clin Microbiol. 47:3851–3861.
- O’Donnell K, Sutton DA, Rinaldi MG, Sarver BAJ, Balajee SA, Schroers H-J, Summerbell RC, VARG R, Crous PW, Zhang N, et al. 2010. Internet-accessible DNA sequence database for identifying fusaria from human and animal infections. J Clin Microbiol. 48:3708–3718.
- O’Donnell K, Ward TJ, Robert VARG, Crous PW, Geiser DM, Kang S. 2015. DNA sequence-based identification of Fusarium: current status and future directions. Phytoparasitica. 43:583–595.
- O’Donnell K, Whitaker BK, Laraba I, Proctor RH, Brown DW, Broders K, Kim H-S, McCormick SP, Busman M, Aoki T, et al. 2022. DNA sequence-based identification of Fusarium: a work in progress. Plant Dis. 106:1597–1609.
- Özer G, İmren M, Paulitz TC, Bayraktar H, Muminjanov H, Dababat AA. 2020b. First report of crown rot caused by Fusarium algeriense on wheat in Azerbaijan. Plant Dis. 104:582.
- Özer G, Paulitz TC, Imren M, Alkan M, Muminjanov H, Dababat AA. 2020a. Identity and pathogenicity of fungi associated with crown and root rot of dryland winter wheat in Azerbaijan. Plant Dis. 104:2149–2157.
- Pavlova VN, Varcheva SE, Bokusheva R, Calanca P. 2014. Modelling the effects of climate variability on spring wheat productivity in the steppe zone of Russia and Kazakhstan. Ecol Modell. 277:57–67.
- Porras-Alfaro A, Bayman P. 2011. Hidden fungi, emergent properties: endophytes and microbiomes Annu. Rev Phytopathol. 49:291–315.
- Porras-Alfaro A, Herrera J, Sinsabaugh RL, Odenbach KJ, Lowrey T, Natvig DO. 2008. Novel root fungal consortium associated with a dominant desert grass. Appl Environ Microbiol. 74:2805–2813.
- Reeb V, Lutzoni F, Roux C. 2004. Contribution of RPB2 to multilocus phylogenetic studies of the euascomycetes (Pezizomycotina, Fungi) with special emphasis on the lichen-forming Acarosporaceae and evolution of polyspory. Mol Phylogenet Evol. 32:1036–60.
- Rodriguez RJ, White JF Jr, Arnold AE, Redman RS. 2009. Fungal endophytes: diversity and functional roles. New Phytol. 182:314–330.
- Ronquist F, Huelsenbeck JP. 2003. MrBayes 3: bayesian phylogenetic inference under mixed models. Bioinformatics. 19:1572–1574.
- Schoch CL, Seifert KA, Huhndorf S, Robert V, Spouge JL, Levesque CA, Chen W, Bolchacova E, Voigt K, Crous PW, et al. 2012. Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proc Nat Acad Sci USA. 109:6241–6246.
- Senatore MT, Ward TJ, Cappelletti E, Beccari G, McCormick SP, Busman M, Laraba I, O’Donnell K, Prodi A. 2021. Species diversity and mycotoxin production by members of the species complex associated with Fusarium head blight of wheat and barley in Italy. Int J Food Microbiol. 358(2021):109298.
- Sieber TN, Grünig CR. 2013. Fungal root endophytes. In: Wasel Y, Eshel A, Kafkafi U, editors. Plant roots: the hidden half. New York, NY: Marcel Dekker; p. 1–49.
- Silvestro D, Michalak I. 2012. raxmlGUI: a graphical front-end for RAxML. Org Divers Evol. 12:335–337.
- Staden R, Beal KF, Bonfield JK. 2000. The staden package, 1998. In: Misener S, Krawetz SA, editors. Bioinformatics methods and protocols. Totowa (Florida): Humana Press; p. 115–30.
- Stamatakis A. 2014. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics. 30:1312–1313.
- Su YY, Guo LD, Hyde KD. 2010. Response of endophytic fungi of Stipa grandis to experimental plant function group removal in Inner Mongolia steppe, China. Fungal Diver. 43:93–101.
- Summerell BA. 2019. Resolving Fusarium: current status of the genus. Annu Rev Phytopathol. 57:323–339.
- Szécsi Á, Magyar D, Tóth S, Szőke C. 2013. Poaceae: a rich source of endophytic fusaria. Acta Phytopathol Entomol Hung. 48:19–32.
- Taylor JW, Jacobson DJ, Kroken S, Kasuga T, Geiser DM, Hibbett DS, Fisher MC. 2000. Phylogenetic species recognition and species concepts in fungi. Fungal Genet Biol. 31:21–32.
- Torbati M, Arzanlou M, Sandoval-Denis M, Crous PW. 2019. Multigene phylogeny reveals new fungicolous species in the Fusarium tricinctum species complex and novel hosts in the genus Fusarium from Iran. Mycol Prog. 18:19–133.
- Tunali B, Nicol JM, Hodson D, Uçkun Z, Büyük O, Erdurmuş D, Hekimhan H, Aktaş H, Akbudak MA, Bağci SA. 2008. Root and crown rot fungi associated with spring, facultative, and winter wheat in Turkey. Plant Dis. 92:1299–1306.
- Vandenkoornhuyse P, Baldauf SL, Leyval C, Straczek J, Young JPV. 2002. Extensive fungal diversity in plant roots. Science. 295:2051.
- Villani A, Moretti A, De Saeger S, Han Z, Di Mavungu JD, Soares CMG, Proctor RH, Venâncio A, Lima N, Stea G, et al. 2016. A polyphasic approach for characterization of a collection of cereal isolates of the Fusarium incarnatum-equiseti species complex. Int J Food Microbiol. 234:24–35.
- Villani A, Proctor RH, Kim H-S, Brown DW, Logrieco AF, Amatulli MT, Moretti A, Susca A. 2019. Variation in secondary metabolite production potential in the Fusarium incarnatum-equiseti species complex revealed by comparative analysis of 13 genomes. BMC Genomics. 20:314.
- White TJ, Bruns T, Lee S, Taylor JW. 1990. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ, editors. PCR protocols: a guide to methods and applications. New York, NY: Academic Press; p. 315–322.
- Xia JW, Sandoval-Denis M, Crous PW, Zhang XG, Lombard L. 2019. Numbers to names – restyling the Fusarium incarnatum-equiseti species complex. Persoonia. 43:186–221.
