ABSTRACT
The domain of unknown function DUF936 family consists of proteins with an average length of around 328 residues that have no characterized function in rice. In this study, we report nine OsDUF936 (OsDUF936.1–OsDUF936.9) family members in rice Nipponbare with three distinct motifs. All the OsDUF936 proteins were classified into four major groups (I, II, III and IV) by phylogenetic analysis. The expression level of OsDUF936.3, OsDUF936.5 and OsDUF936.6 displayed significant increase under salt stress conditions. Overexpression of OsDUF936.6 in Escherichia coli significantly improved the tolerance to salt, while overexpression of OsDUF936.3 and OsDUF936.5 in E. coli did not have such an effect. The results presented here would be helpful for further study on the function of the OsDUF936 family in rice.
Introduction
Abiotic stresses, including drought, salinity and extreme temperatures, have a significant impact on plant productivity and crop quality throughout the world [Citation1,Citation2]. Among these stresses, salinity is a major constraint to rice production worldwide and the salt-affected soils currently account for about 20% of the total rice planting area [Citation3]. Salinity, leading to the disruption of homeostasis and ion distribution in the cell, is considered an important osmotic stress [Citation4–6]. The responses of plants to salinity stress are controlled by abscisic acid (ABA) [Citation7,Citation8], reactive oxygen species (ROS) [Citation9–11], ion transport [Citation12,Citation13] and some transcription factors [Citation14]. In recent years, some success in improving the salt stress tolerance of plants has been achieved [Citation15–22]. However, the regulatory mechanisms involved in salt stress tolerance are not fully understood.
Despite the completion of sequencing and annotation of the genome of some species, a number of genes have not yet been characterized at the biochemical and biological function level. There are about 3000 DUF (domains of unknown functions) families within the Pfam database, representing over 20% of known protein families [Citation23]. In recent years, it was reported that some DUF-containing proteins could enhance the tolerance to abiotic stress in Escherichia coli and plants [Citation24–30].
The DUF936 family consists of plant proteins of unknown function with an average length of around 328 residues. So far, the OsDUF936 family has not been functionally identified in rice. In this study, we analysed the OsDUF936 family in rice and studied the expression profile of nine OsDUF936 members and their expression pattern under various stresses and ABA treatment. Furthermore, we overexpressed gene OsDUF936.6 in E. coli and the results showed that OsDUF936.6 could improve the tolerance to salt in transgenic E. coli.
Materials and methods
Database searches
In order to obtain the sequences of putative OsDUF936 family proteins in the rice genome, we searched the rice genome annotation project (RGAP version 7) database (http://rice.plantbiology.msu.edu/) using domain number PF06075. All the corresponding protein sequences of the putative OsDUF936 family members were then downloaded from RGAP and confirmed with the SMART database [Citation31,Citation32] (http://smart.embl-heidelberg.de/smart/batch.pl). If there were several gene models at one locus, only the complete gene model was selected for further sequence analysis.
Sequence analysis
Information of each OsDUF936 member about the chromosomal localization, amino acid (aa) length and full-length cDNA accessions was obtained from RGAP and the Knowledge-Based Oryza Molecular Biological Encyclopedia (KOME) (http://cdna01.dna.affrc.go.jp/cDNA). The protein localization of OsDUF936 members was predicted using online protein location prediction software PSORT (http://psort.hgc.jp/form.html). MEME version 4.9.1 (http://meme.nbcr.net/meme/) was used to predict the motifs of OsDUF936 family members [Citation33]. MEGA4 was used to construct the phylogenetic tree [Citation34]. The unrooted tree was generated using MEGA4.0 program by the maximum-likelihood method. Bootstrap testing was performed with 1000 resamplings.
Plant growth and various stress and ABA treatment
Rice (Oryza sativa L. subsp. japonica cv. Nipponbare) seeds were germinated for 3 days, and then seedlings were grown hydroponically in a 30-L nutrient solution vessel [Citation35]. In order to examine the expression patterns of OsDUF946 family members, 15 different tissues of rice Nipponbare seedlings were sampled, as follows: Lb1, leaf blade at four-leaf stage; Lb2, leaf blade from plants with four tillers; Lb3, leaf blade at ripening stage; Ls1, leaf sheath at four-leaf stages; Ls2, leaf sheath from plants with four tillers; Rt1, root at four-leaf stages; Rt2, root at from plants with four tillers; St1, stem from plants with four tillers; St2, stem at ripening stage; An, 1.2–1.5 mm anther; Pi, pistil from 10–14 cm inflorescence; Em1, embryo at 7 days after flowering; Em2, embryo at 28 days after flowering; En1, endosperm at 7 days after flowering; En2, endosperm at 28 days after flowering.
At the emergence of the fourth leaf, the seedlings of rice Nipponbare were subjected to various stresses and ABA treatment as described previously [Citation36]. For high salinity treatment, NaCl solution was added to achieve a final concentration of about 200 mmol/L. Drought stress was modelled by putting intact plants in the air without water supply. For cold stress, vigorous seedlings were transferred to a growth chamber at 4 °C (12-h light/12-h dark cycle). For heat stress, the seedlings were subjected to 42 °C heat shock treatments. For ABA treatment, 100 μmol/L ABA was sprayed onto the leaves of seedlings. Seedlings were sampled at 0, 4, 8 and 16 h after treatment.
RNA extraction and real time polymerase chain reaction (PCR)
Total RNA was extracted with Trizol reagent (GIBCO, Burlington, ON, USA) according to the manufacturer's instructions. Then, the total RNA was treated with RNase-free DNase I (Invitrogen, Carlsbad, CA, USA) for 15 min to degrade any potentially contaminated DNA and the resulting RNA was used for first-strand cDNA synthesis. Real-time PCR was performed as described [Citation36]. The rice Actin1 gene (LOC_Os03g50885) was used as an internal control with primers 5'-TGGCATCTCTCAGCACATTC-3' and 5'-TGCACAATGGATGGGCCAGA-3'. The primers used in the OsDUF936 gene family members’ expression analysis are listed in Supplementary Table S1. Data analyses of the relative expression levels with the 2−ΔΔCt method were performed as described previously [Citation37]. All the quantitative PCR products have been confirmed by sequencing.
Assay for abiotic stress tolerance of E. coli transformants
In order to construct the expression vector pET32a-OsDUF936.3, pET32a-OsDUF936.5 and pET32a-OsDUF936.6, designed specific primers were used (Supplementary Table S2). The amplified products were cloned into a pET32a vector at the BamHI–HindIII site to express the fusion protein, which has a Trx•Tag™ thioredoxin at the N-terminus [Citation38]. The transformed E. coli Rosetta cells (Sanborn, MN, USA) were grown in Luria–Bertani (LB) liquid medium containing 100 µg/mL of ampicillin at 37 °C overnight. Isopropylthio-β-D-galactoside (IPTG) was added into the cultures to induce the expression of the transformed gene. The concentrations of all induced cell cultures were adjusted to an OD600 value of 1.0.
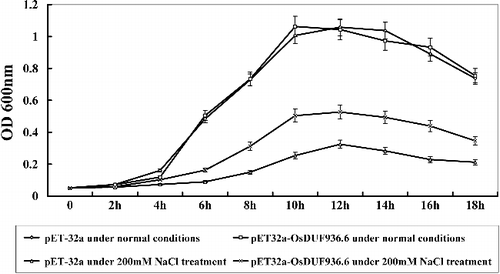
For the assay of the salt tolerance, the growth curve of E. coli recombinants was studied. The OD600 values of E. coli recombinants were measured at 0, 2, 4, 6, 8, 10, 12, 14, 16 and 18 h under normal conditions and 200 mmol/L NaCl treatment.
Data analysis
Three biological replicates were performed for all the experiments. Data are mean values from three independent experiments, with standard error of the means (±SEM). Statistical analysis was performed using the Student's t-test by Microsoft EXCEL version 2003.
Results and discussion
Sequence analysis of OsDUF936 family
Nine OsDUF936 gene family members were identified () by searching the rice genome annotation project with domain number PF06075. Then, the presence of these OsDUF936 genes in the rice genome was confirmed with the SMART database (Supplementary Figure S1). All these genes were named as OsDUF936.1 to OsDUF936.9 according to their positions on pseudomolecules (). The nine members of the rice OsDUF936 are distributed on chromosome 1 (OsDUF936.1), 3 (OsDUF936.2), 4 (OsDUF936.3), 7 (OsDUF936.4) 8 (OsDUF936.5), 9 (OsDUF936.6), 11 (OsDUF936.7 and OsDUF936.8) and 12 (OsDUF936.9), respectively.
Table 1 . OsDUF936 gene family and their predicted protein structure information.
Information about the OsDUF936 family members’ amino acid length, full-length cDNA accessions, molecular weight (MW) and isoelectric points (pI) is presented in . The length of the OsDUF936 proteins was predicted to vary from 177 (OsDUF936.3) to 760 (OsDUF936.6). Three distinct motifs were identified using the MEME motif search tool (http://meme.sdsc.edu/meme/) (Supplementary Figure S2A). Motif 2 was not found in the predicted OsDUF936.4 sequence. Interestingly, in the predicted OsDUF936.2 sequence, only motif 1 was found, while in the predicted OsDUF936.1 and OsDUF936.8 sequences, only motif 3 was found (Supplementary Figure S2B).
Phylogenetic analysis was performed to examine the evolutionary relationships among the nine OsDUF936 family members in rice using MEGA4. Here seven Arabidopsis DUF936 members were included as reference sequences. The results showed that the OsDUF936 family members could be classified into four major groups (I, II, III and IV) (). Group I contained seven members (including OsDUF936.4, OsDUF936.5, OsDUF936.6 and OsDUF936.8), group II contained three members (including OsDUF936.7 and OsDUF936.9), group III contained two members (including OsDUF936.2) and group IV contained three members (including OsDUF936.1).
Figure 1. Phylogenetic relationship of OsDUF936 members in rice and seven DUF936 members in Arabidopsis.
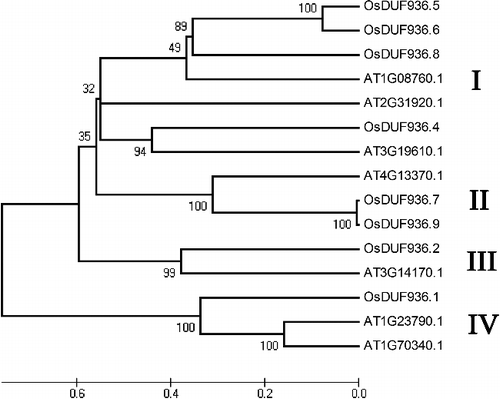
In this study, nine OsDUF936 members that contain the PF06075 signature sequence in rice were identified. The nine members of this OsDUF936 family are distributed on eight chromosomes. The rice genome includes 10 duplicated blocks accounting for 45% of its total size [Citation39]. Therefore, genome duplication might be the mechanism that underlies the distribution of OsDUF936 family members in different chromosomes.
Expression patterns of OsDUF936
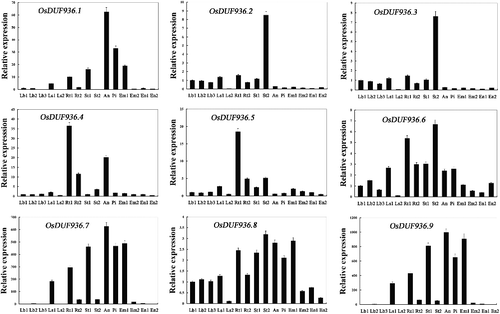
We examined the expression profile of OsDUF936 family members in 15 tissues by real-time PCR to acquire more information. For OsDUF936.2 and OsDUF936.3, they possessed a similar expression profile in 15 tissues and the highest expression was in stem at the ripening stage (St2), whereas the lowest expression was in the leaf sheath from plants with four tillers (Ls2). The other members of the OsDUF936 family showed very different expression patterns in different tissues ().
Figure 2. Real-time PCR analysis of OsDUF936 gene family members in different tissues of rice Nipponbare. Fifteen representative tissues are as follows: Lb1, leaf blade at four-leaf stage; Lb2, leaf blade from plants with four tillers; Lb3, leaf blade at ripening stage; Ls1, leaf sheath at four-leaf stages; Ls2, leaf sheath from plants with four tillers; Rt1, root at four-leaf stages; Rt2, root at from plants with four tillers; St1, stem from plants with four tillers; St2, stem at ripening stage; An, 1.2–1.5 mm anther; Pi, pistil from 10 to 14 cm inflorescence; Em1, embryo at 7 days after flowering; Em2, embryo at 28 days after flowering; En1, endosperm at 7 days after flowering; En2, endosperm at 28 days after flowering.
The expression of the OsDUF936 family was also examined under various stress conditions and ABA treatment (). The expression level of OsDUF936.1, OsDUF936.4 and OsDUF936.9 showed a significant drop under drought and salt conditions, whereas the expression level of OsDUF936.3, OsDUF936.5 and OsDUF936.6 displayed significant increase under salt conditions. The expression of OsDUF936.2 was significantly upregulated under drought conditions, yet downregulated under cold conditions. As for OsDUF936.7 and OsDUF936.8, their expression level was approximately constant under various stress and ABA conditions. ().
Figure 3. Relative expression levels of OsDUF936 gene family members in rice Nipponbare seedlings under various stress and ABA conditions. D0, D1, D2 and D3: drought stress treatment for 0, 4, 8 and 16 h, respectively; S0, S1, S2 and S3: salt stress treatment for 0, 4, 8 and 16 h, respectively; C0, C1, C2 and C3: cold stress treatment for 0, 4, 8 and 16 h, respectively; H0, H1, H2 and H3: heat stress treatment for 0, 4, 8 and 16 h, respectively; A0, A1, A2 and A3: ABA treatment for 0, 4, 8 and 16 h, respectively.
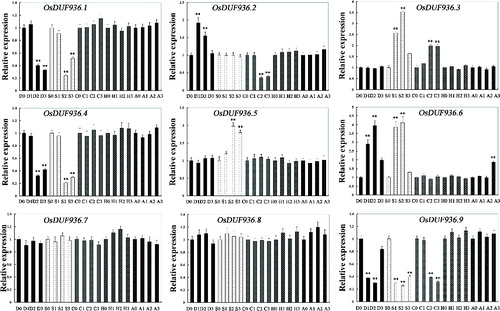
Some gene family members usually display a more or less common significant increase or decrease under various stress conditions [Citation40–42]. All the OsDUF936 members in rice displayed very different expression patterns under various stress conditions and ABA treatment (). Thus, it was suggested that OsDUF936 family members play different roles in response to various stresses.
OsDUF936.6 improved the tolerance to salt in transgenic E. coli
Based on the fact that the expression level of OsDUF936.3, OsDUF936.5 and OsDUF936.6 was significantly elevated under salt conditions, we speculated that they might be involved in salt tolerance. These genes were overexpressed in E. coli and then various stress resistance of recombinants was examined.
Under normal conditions, the growth rate of E. coli transformed with pET32a-OsDUF936.3, pET32a-OsDUF936.5 and pET32a-OsDUF936.6 was the same as that of E. coli transformed with pET-32a (control). Under salt conditions, the growth rate in E. coli transformed with pET32a-OsDUF936.6 was much higher than that of the control (Rosetta/pET-32a) from 2 to 18 h (), whereas there was no evident difference in the growth rate between transgenic lines (E. coli transformed with pET32a-OsDUF936.3 and pET32a-OsDUF936.5) and the control (Supplementary Figure S3). The results indicated that overexpression of OsDUF936.6 in E. coli significantly improved the tolerance to salt, while overexpression of OsDUF936.3 and \OsDUF936.5 in E. coli did not have such an effect.
Production of ROS is a frequent event in organisms subjected to abiotic stresses, such as drought and salinity [Citation43,Citation44]. ROS production causes oxidative damage on membrane lipids and perturbation of cell membrane function [Citation45]. It is therefore very important to enhance the activities of ROS-scavenging enzymes in cells in response to abiotic stress in organisms [Citation46]. At present, the specific mechanism by which salt tolerance is enhanced in transgenic E. coli is unclear. Maybe the salt stress signal is strengthened in E. coli recombinants transformed with OsDUF936.6, and the enhanced stress signal promotes the increased expression of a number of downstream salt-stress-related genes, including genes corresponding to ROS-scavenging enzymes. Thus, the salt tolerance in transgenic E. coli was improved as a result.
Conclusions
This study provides not only important information regarding the OsDUF936 family members, but also showed the expression patterns of OsDUF936 family members in different tissues, and under various abiotic stress and ABA treatment. Importantly, OsDUF936.6 could improve the resistance to salt in transgenic E. coli. These data can provide important reference for further studies of the functions of OsDUF936 family.
Supplemental_Data.pdf
Download PDF (870.7 KB)Disclosure statement
No potential conflict of interest was reported by the authors.
Additional information
Funding
References
- Boyer JS. Plant productivity and environment. Science. 1982;218:443–448.
- Mahajan S, Tuteja N. Cold, salinity and drought stresses: an overview. Arch Biochem Biophys. 2005;444:139–158.
- Gupta B, Huang B. Mechanism of salinity tolerance in plants: physiological, biochemical, and molecular characterization. Int J Genomics. 2014; [cited 2017 Mar 25];2014:701596. DOI: 10.1155/2014/701596
- Serrano R, Rodriguez-Navarro A. Ion homeostasis during salt stress in plants. Curr Opin Cell Biol. 2001;13:399–404.
- Zhu JK. Plant salt tolerance. Trends Plant Sci. 2001;6:66–71.
- Zhu JK. Regulation of ion homeostasis under salt stress. Curr Opin Plant Biol. 2003;6:441–445.
- Park SY, Fung P, Nishimura N, et al. Abscisic acid inhibits type 2C protein phosphatases via the PYR/PYL family of START proteins. Science. 2009;324:1068–1071.
- Fujii H, Zhu JK. Osmotic stress signaling via protein kinases. Cell Mol Life Sci. 2012;69:3165–3173.
- Pei ZM, Murata Y, Benning G, et al. Calcium channels activated by hydrogen peroxide mediate abscisic acid signalling in guard cells. Nature. 2000;406:731–734.
- Kwak JM, Mori IC, Pei ZM, et al. NADPH oxidase AtrbohD and AtrbohF genes function in ROS-dependent ABA signaling in Arabidopsis. EMBO J. 2003;22:2623–2633.
- Ma L, Zhang H, Sun L, et al. NADPH oxidase AtrbohD and AtrbohF function in ROS-dependent regulation of Na(+)/K(+)homeostasis in Arabidopsis under salt stress. J Exp Bot. 2012;63:305–317.
- Zhang X, Miao YC, An GY, et al. K+ channels inhibited by hydrogen peroxide mediate abscisic acid signaling in Vicia guard cells. Cell Res. 2001;11:195–202.
- Sutter JU, Sieben C, Hartel A, et al. Abscisic acid triggers the endocytosis of the Arabidopsis KAT1 K+ channel and its recycling to the plasma membrane. Curr Biol. 2007;17:1396–1402.
- Fujita Y, Fujita M, Shinozaki K, et al. ABA-mediated transcriptional regulation in response to osmotic stress in plants. J Plant Res. 2011;124:509–525.
- Campo S, Baldrich P, Messeguer J, et al. Overexpression of a calcium-dependent protein kinase confers salt and drought tolerance in rice by preventing membrane lipid peroxidation. Plant Physiol. 2014;165:688–704.
- Cai W, Liu W, Wang WS, et al. Overexpression of rat neurons nitric oxide synthase in rice enhances drought and salt tolerance. PLoS ONE. 2015; [cited 2017 Mar 25];10:e0131599. DOI: 10.1371/journal.pone.0131599
- Cui LG, Shan JX, Shi M, et al. DCA1 acts as a transcriptional co-activator of DST and contributes to drought and salt tolerance in rice. PLoS Genet. 2015; [cited 2017 Mar 25];11:e1005617. DOI: 10.1371/journal.pgen.1005617
- Gao Y, Jiang W, Dai Y, et al. A maize phytochrome-interacting factor 3 improves drought and salt stress tolerance in rice. Plant Mol Biol. 2015;87:413–428.
- Hong Y, Zhang H, Huang L, et al. Overexpression of a stress-responsive NAC transcription factor gene ONAC022 improves drought and salt tolerance in rice. Front Plant Sci. 2016; [cited 2017 Mar 25];7:4. DOI: 10.3389/fpls.2016.00004
- Liu Z, Liu P, Qi D, et al. Enhancement of cold and salt tolerance of Arabidopsis by transgenic expression of the S-adenosylmethionine decarboxylase gene from Leymus chinensis. J Plant Physiol. 2017;211:90–99.
- Li P, Li YJ, Zhang FJ, et al. The Arabidopsis UDP-glycosyltransferases UGT79B2 and UGT79B3, contribute to cold, salt and drought stress tolerance via modulating anthocyanin accumulation. Plant J. 2017;89:85–103.
- Wang C, Yang Y, Wang H, et al. Ectopic expression of a cytochrome P450 monooxygenase gene PtCYP714A3 from Populus trichocarpa reduces shoot growth and improves tolerance to salt stress in transgenic rice. Plant Biotechnol J. 2016;14:1838–1851.
- Bateman A, Coggill P, Finn RD. DUFs: families in search of function. Acta Crystallogr F. 2010;66:1148–1152.
- He X, Hou X, Shen Y, et al. TaSRG, a wheat transcription factor, significantly affects salt tolerance in transgenic rice and Arabidopsis. FEBS Lett. 2011;585:1231–1237.
- Kim SJ, Ryu MY, Kim WT. Suppression of Arabidopsis RING-DUF1117 E3 ubiquitin ligases, AtRDUF1 and AtRDUF2, reduces tolerance to ABA-mediated drought stress. Biochem Biophys Res Commun. 2012;420:141–147.
- Luo C, Guo C, Wang W, et al. Overexpression of a new stress-repressive gene OsDSR2 encoding a protein with a DUF966 domain increases salt and simulated drought stress sensitivities and reduces ABA sensitivity in rice. Plant Cell Rep. 2014;33:323–336.
- Wang L, Shen R, Chen LT, et al. Characterization of a novel DUF1618 gene family in rice. J Integr Plant Biol. 2014;56:151–158.
- Guo C, Luo C, Guo L, et al. OsSIDP366, a DUF1644 gene, positively regulates responses to drought and salt stresses in rice. J Integr Plant Biol. 2016;58:492–502.
- Li L, Xie C, Ye T, et al. Molecular characterization, expression pattern, and function analysis of the rice OsDUF866 family. Biotechnol Biotechnol Equip. 2017;31:243–249.
- Li L, Ye T, Xu J, et al. Molecular characterization and function analysis of the rice OsDUF946 family. Biotechnol Biotechnol Equip. 2017;31:477–485.
- Schultz J, Milpetz F, Bork P, et al. SMART, a simple modular architecture research tool: identification of signaling domains. Proc Natl Acad Sci U S A. 1998;95:5857–5864.
- Letunic I, Doerks T, Bork P. SMART: recent updates, new developments and status in 2015. Nucleic Acids Res. 2015;43:D257–D260.
- Bailey TL, Boden M, Buske FA, et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 2009;37:W202–W208.
- Tamura K, Dudley J, Nei M, et al. MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol. 2007;24:1596–1599.
- Li L, Liu C, Lian X. Gene expression profiles in rice roots under low phosphorus stress. Plant Mol Biol. 2010;72:423–432.
- Li L, Ye T, Gao X, et al. Molecular characterization and functional analysis of the OsPsbR gene family in rice. Mol Genet Genomics. 2017;292:271–281.
- Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods. 2001;25:402–408.
- LaVallie ER, DiBlasio EA, Kovacic S, et al. A thioredoxin gene fusion expression system that circumvents inclusion body formation in the E. coli cytoplasm. Biotechnology (N Y). 1993;11:187–193.
- Wang X, Shi X, Hao B, et al. Duplication and DNA segmental loss in the rice genome: implications for diploidization. New Phytol. 2005;165:937–946.
- Dubouzet JG, Sakuma Y, Ito Y, et al. OsDREB genes in rice, Oryza sativa L., encode transcription activators that function in drought-, high-salt- and cold-responsive gene expression. Plant J. 2003;33:751–763.
- Ito Y, Katsura K, Maruyama K, et al. Functional analysis of rice DREB1/CBF-type transcription factors involved in cold-responsive gene expression in transgenic rice. Plant Cell Physiol. 2006;47:141–153.
- Su CF, Wang YC, Hsieh TH, et al. A novel MYBS3-dependent pathway confers cold tolerance in rice. Plant Physiol. 2010;153:145–158.
- Miller G, Shulaev V, Mittler R. Reactive oxygen signaling and abiotic stress. Physiol Plant. 2008;133:481–489.
- Mittler R. Oxidative stress, antioxidants and stress tolerance. Trends Plant Sci. 2002;7:405–410.
- Gechev TS, Van Breusegem F, Stone JM, et al. Reactive oxygen species as signals that modulate plant stress responses and programmed cell death. Bioessays. 2006;28:1091–1101.
- Mittler R, Vanderauwera S, Gollery M, et al. Reactive oxygen gene network of plants. Trends Plant Sci. 2004;9:490–498.