Abstract
The Saihanba Mechanical Forest is an artificial national forest park with a forest-steppe landscape. The flora and fauna have been extensively studied, but a comprehensive understanding of the bacterial community composition and structure of the natural rainwater lakes present in the area has rarely been reported. In this study, the structure and functional characteristics of bacterial communities in lake sediments and water samples in the Saihanba artificial forest were investigated using 16s rRNA high-throughput sequencing. Microbial diversity analyses revealed that Proteobacteria, Acidobacteria, Bacteroidota and Verrucomicrobiota microbiota dominated. The abundance of Proteobacteria and Actinobacteriota was significantly higher (p < .05) in water samples compared to sediment samples. PICRUSt2 functional analysis predicted genes associated with the degradation of xenobiotics and the execution of essential metabolic processes. Here, we report differences in the composition of native bacterial communities in sediments and water under the Saihanba artificial forest and make functional gene predictions. This study provides a reference for further exploring the structure and functional characteristics of microbial communities in water samples and sediment environments of lakes under planted forests.
Introduction
Lakes and rivers are crucial for maintaining biodiversity in forest ecosystems. They play a vital role in climate regulation, landscaping, and facilitating organic matter decomposition [Citation1]. Freshwater lakes are low-salinity lakes, and one of the key factors in ensuring the health of freshwater systems is a sufficient understanding of their bacterial community structure [Citation2]. Planktonic bacteria are involved in key ecological functions such as nutrient cycling, organic matter degradation and water purification in lakes, and their activities are directly related to lake water quality [Citation3–5]. Analyzing microbial diversity, abundance, and functional genes allows to uncover the dynamics of ecological processes in lakes [Citation6]. Additionally, comprehending the structure of bacterial communities aids in identifying potential environmental problems and ecological risks. Microbial communities in lakes are responsive to environmental changes, pollution, and human activities [Citation7]. Monitoring bacterial communities can offer early warnings of potential lake health issues, providing a scientific basis for protecting and managing lake ecosystems. Evaluation of the composition and dynamics of bacterial communities in three temperate lakes with varying trophic statuses on an annual scale revealed significant seasonal variations [Citation8]. Similar changes have been reported in the characteristics in Lake Taihu [Citation9]. Furthermore, bacterial abundance is significantly impacted by human activities, particularly agricultural discharges like pesticides and fertilizers. This leads to increased nutrient concentrations in lake water, influencing changes in bacterial abundance. Additionally, large amounts of organic matter and microorganisms from urban pollutant discharges also affect the bacterial abundance of lakes [Citation10]. Microbial diversity and abundance may also be influenced by the diversity of plant species in the surrounding forest and geographic location. Environments influenced by the same forests share similar resources, leading to comparable bacterial communities, while lakes that are closer or more connected have more similar bacterial communities [Citation11].
Studying microorganisms in freshwater involves traditional microbiological techniques and high-throughput sequencing-based studies. The classical microbiological techniques are still an important part of bacterial diversity studies. However, these methods have limitations in analyzing complex environmental samples, leaving many "uncultured microorganisms" undetected and restricting the overall understanding of the bacterial community. Modern molecular biology techniques have emerged in recent years to address the limitations of traditional culture methods. For example, 16S rRNA sequence analysis techniques can use information from specific genes (16S rRNA genes) to conduct thorough studies on bacterial communities [Citation12,Citation13].
Previously, the Saihanba Mechanical Forest ecosystem faced significant impacts from climate change and human activities, leading to the gradual transformation of the forest into desertified land. Nevertheless, responding to climate change and heightened environmental awareness, a large-scale reforestation initiative has been launched in the Saihanba area. Presently, the forest cover in the Saihanba Mechanical Forest exceeds 82%, dominated by vegetation types such as larch, pinus sylvestris, birch and spruce. Artificial forests have become a common landscape type in the area, playing an important role in soil and water conservation, combating climate change and providing other ecosystem services. Moon Lake, Seven Star Lake and Shenlong Lake, nestled amid vegetation, are three lakes in the Saihanba Mechanical Forest formed by the convergence of natural rainwater. Annually, from July to September, Moon Lake, Seven Star Lake and Shenlong Lake draw a significant number of tourists, establishing themselves as popular attractions. Among the three lakes, Moon Lake attracts the highest number of visitors. Examining the microbial diversity in the surface water and sediments of the three lakes aids in understanding the potential impact of the unique geoclimatic characteristics of the plantation forest and anthropogenic factors resulting from increased tourism on lake microbes.
In this study, 16S rRNA amplicon sequencing was used to assess the diversity and functional characteristics of bacterial populations in Moon Lake, Seven Star Lake and Shenlong Lake, which were formed by natural rainwater aggregation in the Saihanba plantation forest. Functional prediction aspects focused on genes associated with bioremediation potential to shed light on the role of bacteria in lake ecosystems and provide insights into potential contributions to processes such as bioremediation.
Materials and methods
Site description
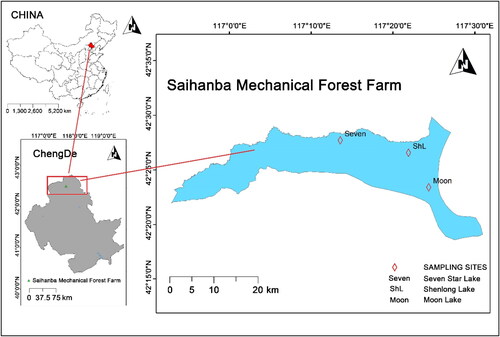
The Saihanba Mechanical Forest is situated at the intersection of the southeastern edge of the Inner Mongolian Plateau and the northern mountainous region of Hebei Province, China. It spans 9,219 km2 with an average elevation of 1500 m. Established 61 years ago, it stands as the largest artificial forest in China. Experiencing snow cover for seven months annually, the peak tourist season at the Saihanba Mechanical Forest occurs from July to September. Lakes are frequented by tourists compared to other landscapes in the Saihanba artificial forest. The study area (Seven Star Lake, Moon Lake and Shen Long Lake) is located in the Saihanba Mechanical Forest Farm, formed by natural rainwater gathering, with a special cold-temperate continental monsoon climate. Detailed geographic information on the three lakes can be found in Supplemental Table S1. The three lakes can represent the typical characteristics of lakes in the Saihanba Mechanical Forest.
Sample collection
A total of 18 sediment samples and 18 water samples collected at two different time points (September and December) were used in this study. The samples were collected from Moon Lake, Seven Star Lake, and Shenlong Lake (), which are formed by natural rainwater collection in the Saihanba Mechanical Forest, and sediment samples and water samples were taken from 3 different locations in each lake (MnSF: fall sediment sample from Moon Lake, MnSW: winter sediment sample from Moon Lake, MnWF: fall water sample from Moon Lake, MnWW: winter water sample from Moon Lake, STSF: fall sediment sample from Seven Star Lake, STSW: winter sediment sample from Seven Star Lake, STWF: fall water sample from Seven Star Lake, STWW: winter water sample from Seven Star Lake, SLSA: fall sediment sample from Shenlong Lake, SLSW: winter sediment sample from Shenlong Lake, SLWF: fall water sample from Shenlong Lake, SLWW: winter water sample from Shenlong Lake). Sediment samples were collected from the bottom of the lake using a homemade plastic pipette tied to a sterile metal spoon, approximately 30 g at a time, and the sediment samples were collected directly into sterile polyethylene bags. Water samples were collected in 3 L of water in a plastic bucket rinsed with lake water at a distance of 20 cm below the surface water body in deep water. Winter water samples were collected in subglacial water and sediment was collected from the bottom of the lake. The collected samples were immediately stored in ice and transported back to the laboratory and stored at −80 °C within 24 h. The samples were thawed before DNA extraction. Water samples were filtered through 0.22 μm microporous membranes, which were stored in sterile test tubes. DNA was extracted from sediment and water samples within 48 h of sample collection processing and stored at −20 °C. Three replicate groups were set up for each water and sediment sample during sampling to ensure the accuracy of the experiment. Moon Lake, Seven Stars Lake and Shenlong Lake can be used as representatives of local lakes.
DNA extraction and PCR amplification
DNA was extracted from water and sediment samples using the CTAB method with reagents from Nobleryder (China). The CTAB method is based on a CTAB cationic decontaminant in a salt solution of high ionic strength, which allows the CTAB to form complexes with proteins and polysaccharides, thus effectively removing impurities [Citation14]. After DNA extraction was completed, the quality and quantity of extracted DNA was checked using 1%(w/v) agarose gel electrophoresis. A portion of each DNA sample was taken in a centrifuge tube and was diluted to 1 ng/μL using sterile water. The V4 variable region of the 16S rRNA gene was subjected to PCR amplification using primers (515 F primer (5ʹ- GTGCCAGCMGCCGCGGTAA −3ʹ) and 806 R primer (5ʹ- GGACTACHVGGGTWTCTAAT-3ʹ)) [Citation15]. In the PCR amplification system (30 μL), 15 μL of Phusion® High-Fidelity PCR Master Mix (M0531S, New England Biolabs), 1 μL each of forward and reverse primers, 10 μL of DNA template, and 2 μL of sterile ultrapure water were used. Details of all primers are given in Supplemental Table S2. Thermal cycling (T100, Bio-Rad, USA) consisted of initial denaturation at 98 °C for 1 min, followed by 30 cycles of denaturation at 98 °C for 10 s, annealing at 50 °C for 30 s, and elongation at 72 °C for 30 s; and a final step of 72 °C for 5 min. Finally, aliquots of the PCR products mixture were mixed with the same volume of 1XTAE buffer and the amplification products were separated via 2% agarose gel electrophoresis. PCR products were mixed in equidensity ratios. Then, the PCR products were purified using the Universal DNA purification kit (TianGen, China).
Illumina NovaSeq sequencing
As recommended by the manufacturer, TruSeq® DNA PCR-Free Sample Preparation Kit (Illumina, USA) was used to generate sequencing libraries. The library quality was assessed on the Agilent 5400 fragment analyser (Agilent Technologies Co Ltd., USA). Finally, the library was sequenced on an Illumina NovaSeq 6000 platform and 250 bp paired-end reads were generated.
Raw sequences were deposited in the National Center for Biotechnology Information (NCBI) Sequence Read Archive (SRA) under the project accession number PRJNA895964 (http://www.ncbi.nlm.nih.gov/bioproject/895964).
Bioinformatics analysis of species composition and diversity
This analysis was conducted using the "Atacama soil microbiome tutorial" from Qiime2docs and a custom program script (https://docs.qiime2.org/2019.1) [Citation16]. After quality filtering and trimming of the de-multiplicated sequences of each sample, denoising, merging and identifying and removing the chimeric sequences using the DADA2 plugin of QIIME2, the mean Amplicon Sequence Variant (ASV) tables were obtained [Citation17]. The ASV sequences were aligned to a pre-trained SILVA database (trimmed to the V4 region bound by the 515 F/806R primer pair) using the QIIME2 feature-classifier plugin to generate the taxonomy table. The QIIME2 feature-table plugin was used to filter out mitochondrial and chloroplast sequences that may have contaminated the data. For identifying bacteria with different abundances between samples and groups, appropriate methods were used, including Kruskal–Wallis [Citation18–20]. Alpha diversity metrics were calculated using the core-diversity plugin within QIIME2. As a means to investigate structural changes in the bacterial communities of the different samples, beta diversity distances were determined using Bray-Curtis distances [Citation21] and then visualized by principal coordinate analysis (PCoA) and non-metric multidimensional scaling (NMDS).
Bioinformatics analysis for function prediction
PICRUSt2 v2.3.0 (Phylogenetic Investigation of Communities by Reconstruction of Unobserved States) software was used to evaluate the gene function profiles of common ancestors of bacteria and archaea identified by 16S rRNA sequences [Citation22]. PICRUSt2 upgrades PICRUSt1 by updating the previous Greengenes database to an IMG database and allowing users to add custom databases. Compared to the PICRUSt1 software, PICRUSt2 has increased the number of bacteria and archaea measured by nearly 20 times. PICURSt2 software predicts the metagenome in the sample based on the ASV abundance tables. Functional predictions are classified into six KEGG levels I, namely: Metabolism, Genetic Information Processing, Cellular Processing, Human Diseases, Organismal Systems and Environmental Information Processing.
Three tools are required to use the PICRUSt2 software: HMMER (http://www.hmmer.org) to place the table of ASV features to be tested; EPA-ng8 to determine the best position of the ASV in the reference phylogeny; and GAPPA9 to output a new phylogenetic tree containing the ASV display positions. PICRUSt2 software uses sequencing data based on 16S rRNA gene sequences and infers the functional genomes in the samples by comparing them to databases of known genomes. In this study, we analyzed KEGG (Kyoto Encylopaedia of Genes and Genomes) levels I, II and III functional relative abundance using the PICRUSt2 tool. In addition, we further analyzed KEGG orthology groups (KO) top ranked enzymes for analyzing comparative microbial metabolic pathways.
Results
Diversity analysis based on 16S rRNA sequencing
After performing 16S rRNA sequencing, a total of 1,810,567 splice reads were obtained after quality control of all sequences from all samples using the DADA2 plugin in the QIIME2 software, which generated ASVs (Amplicon Sequence Variants). Further alpha and beta diversity analyses were conducted using the QIIME2 diversity plugin.
The sparsity curves indicate ample sequencing depth, affirming the acquisition of a satisfactory number of sequences for alpha analysis (). In , the Chao1 diversity index, Shannon diversity index, and Simpson diversity index were calculated for both water and sediment samples. The Chao1 diversity index measures the species richness of the community, providing an estimation of the total number of species present. The Shannon and Simpson diversity indices assess the evenness or homogeneity of the samples, taking into account both species richness and abundance. To determine whether there was a significant difference in alpha diversity between the water and sediment samples, the Kruskal–Wallis method was employed for statistical comparison.
Figure 2. Alpha and beta diversity analysis based on 16S rRNA sequencing results. (A) Rarefaction curves were generated to assess the sequencing depth of water samples and sediment samples collected from Moon Lake, Seven Star Lake and Shen Long Lake. (B) Chao1 diversity index, (C) Shannon diversity index and (D) Simpson diversity index, which measures species richness, were calculated for both water samples and sediment samples. Sediment samples compared with the water samples, *p < .05. (E) NMDS (Non-Metric Multidimensional Scaling) analysis was performed based on Bray-Curtis distances to visualize the dissimilarities among samples. (F) PCoA (Principal Coordinates Analysis) was also conducted using Bray-Curtis distances to assess the variation and clustering patterns of samples. Specific labels were used to indicate different sample types: ST: Seven Star Lake; SL: Shenlong Lake; Mn: Moon lake; SF: sediment samples collected in fall; SW: sediment samples collected in winter; WF: water samples collected in fall; WW: water samples collected in winter.
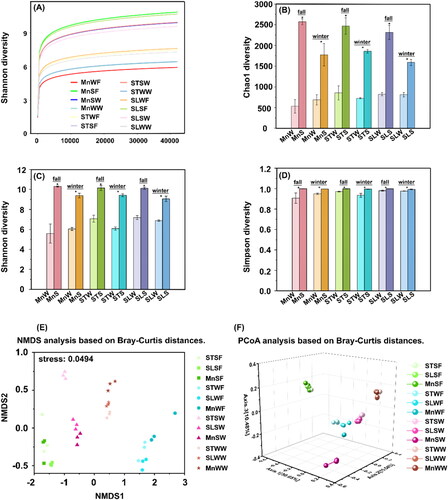
The results indicate that water samples from the same lake exhibited lower homogeneity and species abundance compared to sediment samples. In terms of fall samples, the sediment samples from Moon Lake (MnS) showed the highest Chao1 diversity index with a value of 2574, followed by the sediment samples from Seven Star Lake (STS) with a value of 2471, and finally the sediment sample from Shen Long Lake (SLS) with a value of 2314. The order of magnitude of the Chao1 diversity index across all samples was: MnS > STS > SLS > STW > STW > MnW. This order was consistent for both the Shannon and Simpson diversity indices as well. For winter samples, the sediment sample from Seven Star Lake exhibited the highest Chao1 diversity index with a value of 1860, followed by the sediment sample from Moon Lake with a value of 1775, and finally the sediment sample from Shen Long Lake with a value of 1590. Similar to the fall samples, the overall trend for the Chao1 diversity index across all samples was: STS > MnS > SLS > SLW > STW > MnW.
The beta diversity analysis was conducted using Bray-Curtis distances, and the results were visualized through PCoA and NMDS analyses (). In the NMDS plot, a stress value of less than 0.05 indicates that the clustering accurately represents the data. The bacterial communities of sediment and water in the three lakes formed by natural rainwater aggregation under artificial forest differed more between groups than within groups, and the three lakes showed good clustering among themselves, with the water samples clustering best in winter. However, the sediment samples differed significantly (p = .049) from the water samples under the same season. In the PCoA plot, the sediment samples of the three lakes in the fall were relatively clustered and the water samples were relatively dispersed, with Moon Lake being distant from the water samples of the other two lakes.
Analysis of bacterial community using QIIME2
To study the species composition of the samples, we selected representative sequences of ASVs and compared them with the SILVA database to obtain species annotation information. Based on the species annotation information, ASVs and their contained sequences annotated as chloroplasts, mitochondria and those that could not be annotated to the phylum level were removed. Based on the absolute abundance of ASVs and annotation information, the samples were classified into archaea and bacteria, and a total of 9894 (archaea) and 1,505,922 (bacteria) valid sequences were obtained, which bacteria were further divided into phyla, classes, orders, families and genera. Additionally, a statistical analysis of the number of sequences at the taxonomic level was performed to more effectively assess the species annotation resolution of the samples. Unclassified ASVs were those that had not been fully taxonomically determined.
At the phylum level (), Proteobacteria were dominant in all water samples, with Alphaproteobacteria and Betaproteobacteria being the primary representative classes (). In addition, the abundance of Bacteroidetes, Actinobacteria and Firmicutes was also high in all three water samples (Supplemental Figure S1A). The dominant phyla in the sediments were Proteobacteria, Acidobacteriota, Bacteroidota, Actinobacteriota, Desulfobacterota and Chloroflexi. A total of 30 phyla were detected in the winter water samples, while 65 phyla were detected in the winter sediments. The relative abundance of Firmicutes was higher in the winter sediment samples (Supplemental Figure S1B). Proteobacteria and Actinobacteria were more abundant in the water samples compared to the sediment samples. Acidobacteriota was unique to the sediment samples (Supplemental Figure S1).
Figure 3. Classification of bacteria based on 16S rRNA gene sequences. The graphs are plotted with Origin v. 2021. (A) At the phylum level; (B) at the class level; (C) at the genus level. MnW: water samples from Moon Lake; MnS: sediment samples from Moon Lake; STW: water samples from Seven Star Lake; STS: sediment samples from Seven Star Lake; SLW: water samples from Shenlong Lake; SLS: sediment samples from Shenlong Lake.
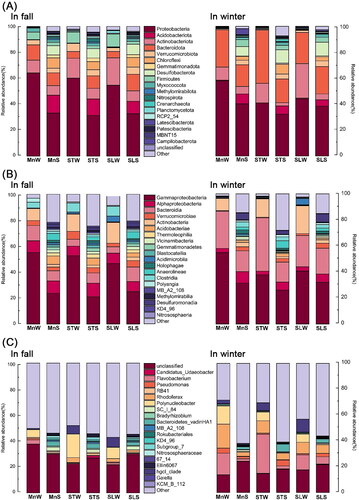
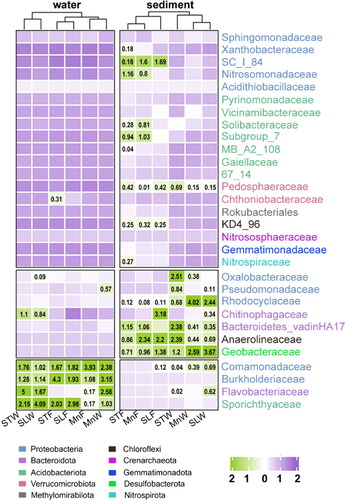
A total of 178 families were observed in all samples, among the water samples, Comamonadaceae, Burkholderiaceae (Proteobacteria), Flavobacteriaceae (Bacteroidota) and Sporichthyaceae (Acidobacteriota) were in higher abundance. The abundance of Oxalobacteraceae, Pseudomonadaceae, Rhodocyclaceae (Proteobacteria), Chitinophagaceae, Bacteroidetes_vadinHA17 (Bacteroidota), Anaerolineaceae (Chloroflexi) and Geobacteraceae (Desulfobacterota) was high in the sediment samples. Family Pedosphaeraceae (Verrucomicrobiota) was specific to the sediment samples.
To analyze the correlation between the samples (water, sediment), a cluster heat map () was made at the family level. The clustering analysis showed a strong seasonal correlation. However, the hierarchical cluster analysis method shows there was a weak correlation between sediment samples and water samples.
Figure 4. Cluster tree of samples constructed by cluster analysis. Family level heat maps are shown in the picture. The graphs are plotted with R software version 4.1.3. MnW: water samples from Moon Lake; MnS: sediment samples from Moon Lake; STW: water samples from Seven Star Lake; STS: sediment samples from Seven Star Lake; SLW: water samples from Shenlong Lake; SLS: sediment samples from Shenlong Lake; F: fall; W: winter.
In this study, a total of 1185 genera were detected in all the samples, of which unclassified genera accounted for 50% of the total (). The relative abundance of unclassified genera was higher in fall samples compared to winter samples. The genus with higher abundance in the water samples was Polynucleobacter, and the genus with higher abundance in the sediment samples was Bacteroidetes_vadinHA17. The genus Hgcl_clade was specific to the water samples.
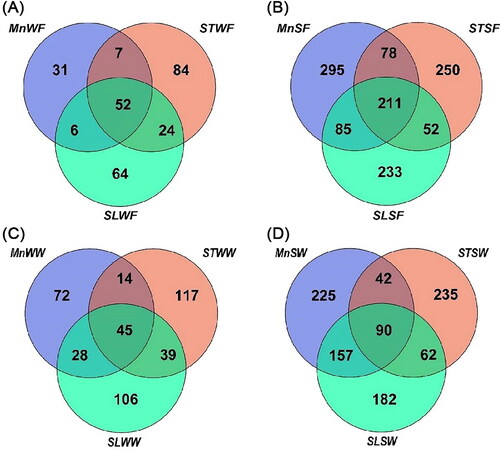
At the family level, 52 shared ASVs counts were observed in the fall water samples and 211 shared ASV counts were observed in the sediment samples (). The fall and winter water samples from Seven Star Lake (84,117) contained the largest number of unique ASVs (). Fall Moon Lake sediment samples (295) contain the most unique ASVs (). Winter Seven Star Lake sediment samples (235) contain the most unique ASVs ().
Figure 5. Venn diagram demonstrating unique and shared ASVs at the family level. The graph is built using the Wekemo Bioincloud (https://www.bioincloud.tech). (A) Among the water samples in fall, (B) among the sediment samples in fall, (C) among the water samples in winter and (D) among the sediment samples in winter of the three lakes. MnW: water samples from Moon Lake; MnS: sediment samples from Moon Lake; STW: water samples from Seven Star Lake; STS: sediment samples from Seven Star Lake; SLW: water samples from Shenlong Lake; SLS: sediment samples from Shenlong Lake; F: fall; W: winter.
Bacterial functional genes predicted by PICRUSt2 software
PICRUSt2 software was employed to predict the functional genes present in the samples, utilizing the abundance tables of ASVs. Following quality control on all sequences across all samples using the DADA2 plugin in QIIME2 software, a total of 2,171,558 non-chimeric sequences were obtained.
KEGG pathways classification
The PICRUSt2 software predicted functions showed the highest percentage of gene sequences for "Metabolism", followed by "Genetic Information Processing". Supplemental Figure S3 shows the comparison of significant differences in KEGG level I of the three lakes in different seasons of water samples. The results show that the metabolic functions of the same lake showed variability across seasons, with winter samples of the three lakes showing significantly higher metabolic functions than fall samples (p < .05, Supplemental Figure S2) ().
Figure 6. Bacterial community functions predicted by PICRUSt2 at level III which related to xenobiotics degradation and metabolism.
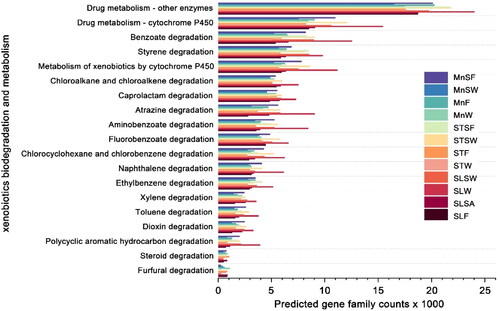
At level II, many sequences exhibit amino acid metabolism and metabolism of cofactors and vitamins capacity (Supplemental Figure S4). Most aquatic life organisms including bacteria, are exposed to these xenobiotics, because the lake is afflicted with water pollution as a result of numerous sources of agricultural pollution, as well as tourists throwing pollutants into the water. Consequently, 19 KEGG pathways related to xenobiotic biodegradation and metabolism were chosen and exhibited higher relative abundance. At the top of the list were Drug metabolism – other enzymes, followed by Drug metabolism – cytochrome P450. Benzoate degradation, Styrene degradation and Metabolism of xenobiotics by cytochrome P450 were close in abundance.
In both sediment and water samples, a total of 7629 KEGG orthology groups (KOs) were identified. In detail, sediment samples yielded 7203 KOs, and water samples in fall yielded 7261 KOs.
For winter samples, sediment samples produced 7309 KOs, and water samples resulted in 7128 KOs. The most abundant KO was K03088 () associated with signal transduction genes, which corresponds to the RNA polymerase sigma-70 factor of the ECF (extracellular function) subfamily. Next were the ABC-2 type transport system-related genes (e.g. K01990, K01992, K02003, K02004).
Table 1. Enzyme classes corresponding to the more abundant KEGG orthology groups (KOs).
Discussion
In this study, the bacterial diversity and functions in lakes formed by natural rainfall in the Saihanba plantation forest were investigated for the first time. The Saihanba artificial forest has been regarded as a symbol of pioneering and protecting the environment in China, and more attention has been paid to the study of its cultural significance, while the study of microorganisms in the lakes has lacked sufficient attention and in-depth scientific investigation. We still have limited understanding of the species diversity, community structure, ecological functions of microorganisms in these lakes and their influence on the stability of lake ecosystems. This study provides new insights into the structure and function of bacterial populations in lakes formed by natural rainwater aggregation in artificial forests through the use of culture-independent molecular investigation methods.
Alpha diversity analysis of both water and sediment samples indicated the presence of diverse bacterial communities in the lake ecosystem, even in the face of harsh winter conditions and low temperatures in the Saihanba artificial forest. Sediment samples showed higher diversity compared to water samples. This finding is consistent with previous studies and suggests that in aquatic environments, sediment samples tend to harbor more bacterial species compared to water samples [Citation23,Citation24]. The elevated microbial diversity observed in sediment samples can be attributed to several key factors. First and foremost, sediment environments tend to exhibit greater environmental heterogeneity compared to water samples. This inherent variability provides a wide array of ecological niches for bacterial communities to thrive in, leading to increased species richness. Moreover, sediment samples typically boast higher concentrations of essential nutrients such as nitrogen, phosphorus and organic matter. These nutrient-rich conditions create a favorable environment for microbial growth and diversity, as they provide ample resources to support various microbial species. Furthermore, the substantial variation in the levels of dissolved oxygen within different sediment layers plays a crucial role. This fluctuation in oxygen availability creates niches suitable for a diverse range of bacterial physiological types, allowing them to occupy specific ecological niches tailored to their oxygen preferences.
In terms of seasonal changes in the diversity of the bacterial communities in the water samples, it was observed that the diversity in Moon Lake was higher in winter than in fall. This change could potentially be attributed to the fact that Moon Lake receives a significant number of tourists from July to September each year, resulting in increased anthropogenic disturbances. Previous studies have shown that anthropogenic factors can lead to a decrease in microbial diversity in lakes [Citation25,Citation26]. However, it is worth noting that there was no significant difference in the diversity of water samples between Moon Lake and the other two lakes during winter. This suggests that the bacterial community may possess some self-restoration ability, allowing it to recover and maintain diversity in the face of disturbances [Citation27,Citation28].
Understanding of the distribution of microbial abundance in lake sediments and the factors that regulate it remains limited. In this study, Proteobacteria, Acidobacteriota, Bacteroidota, Actinobacteriota, Desulfobacterota and Chloroflexi were the phyla with higher relative abundance in the lake sediment samples. The sediment communities of the three lakes were dominated by many freshwater bacterial genera and lineages that are common in the sediment of smaller freshwater lakes [Citation28–30].
Since lakes may be disturbed by agricultural pollution as well as litter dropped into the water by tourists, we selected 19 KEGG pathways associated with biodegradation and metabolism of xenobiotics. Among them Drug metabolism - other enzymes had the highest abundance, followed by Drug metabolism - cytochrome P450, Benzoate degradation, Styrene degradation and Metabolism of xenobiotics by cytochrome P450 were close in abundance. Biomarkers represent systemic changes caused by toxic substances and can serve as connectors between pollution and biological effects that can indicate the health of an ecosystem. It is possible that some accumulation of pharmaceuticals may be present in the lakes of the present study, as cytochrome P450 has been significantly correlated with pollutants [Citation31] In addition, the identification of pathways associated with benzoate degradation suggests that catechol may also be degraded. Enzymatic mechanisms enable microorganisms to utilize benzoate or catechol as a source of carbon and energy for growth, and the presence of these degradation pathways in bacteria plays a crucial role in the natural bioremediation of catechols and aromatic compounds in polluted environments [Citation32].
Further analysis of KO abundance in the lake revealed that K03088 corresponds to the RNA polymerase sigma-70 factor of the ECF (extracellular cytoplasmic function) subfamily, which is widely present in water and sediments, and the presence of K03088 in water and sediment samples suggests that microorganisms in these environments possess the ability to express the RNA polymerase sigma-70 factor-related genes. This factor plays a crucial role in coordinating cellular responses and regulating the expression of genes involved in various metabolic and physiological processes [Citation33,Citation34]. Ther was higher abundance with ABC-2 type transport system-related genes (e.g. K01990, K01992, K02003, K02004), which transport toxins into the extracellular matrix and target membrane components [Citation35]. Overall, these findings provide insights into the functional potential of the bacterial communities in the studied lakes, particularly, with regard to their ability to degrade xenobiotics and perform basic metabolic processes. Further research and understanding of these functional pathways could contribute to a better understanding of the ecological roles and activities of microorganisms in ecosystem dynamics.
Limitations
On the one hand, different kits and contamination during sample handling may affect the results from the experiment. Therefore, the efforts to minimize the occurrence of contaminants included standardized sample collection, the CTAB method for sample DNA extraction, controlling the number of amplification cycles and attention to the aseptic operation of the experiments. On the other hand, 16S rRNA sequencing can reveal the microbial community composition of a sample as well as its relative abundance, but it cannot estimate the absolute abundance. Several simulation experiments have demonstrated a weak correlation between 16S rRNA gene abundance and the abundance of real organisms, so the method of estimating the abundance of species present in the initial sample based on the abundance of sequencing reads does have some drawbacks [Citation36–38]. However, 16S rRNA gene sequencing is a low-cost technique; hence, it is still widely applied. Measuring microbial community structure by the abundance of sequencing reads does not always have a significant impact, but a significant impact on inferences about microbial community structure in some datasets was indeed seen. Therefore, in subsequent studies, we will assess the changes in spatial and temporal distribution characteristics in the environment to reduce amplification bias during PCR. It is important to note that the limitations of 16S rRNA gene sequencing technology may restrict our understanding of the full extent of anthropogenic and climatic influences on freshwater lakes near artificial forests. To overcome these limitations and gain a deeper understanding, future studies should consider using full-length sequencing. The full-length sequencing approach could provide a more comprehensive understanding of the bacterial communities in lakes near planted forests and their response to anthropogenic and climatic impacts.
The findings from this study can serve as a valuable case study to guide further investigations into lakes formed by rainwater aggregation near artificial forests. In the meantime, there is a pressing need to prioritize the conservation of plantation forests and their surrounding lake ecosystems. Protecting and restoring these ecosystems will not only support the recovery of bacterial communities but also provide a broader platform for exploring restoration efforts in plantation forests. We can better understand the complex interactions between microbial communities, human activities and environmental factors, ultimately leading to more effective conservation and management strategies.
Conclusions
Presently, there have been limited reports on the use of high-throughput sequencing techniques to predict the microbial population structure and pollutant degradation in the surrounding lake environments under relatively monoculture plant species in the artificial forest. This finding suggests that bacteria in lakes under artificial forests may also have the potential to develop undiscovered valuable natural products. In the future, further identification at the species level using methods such as shotgun metagenomics or quantitative PCR will be necessary.
Authors contributions
XG: Data curation; Investigation; Visualization; Writing – original draft, review & editing; YL: Writing – review & editing; investigation; KL: Data curation; original draft preparation; funding acquisition; project administration; CT, JC: Visualization; SX: Methodology; formal analysis; funding acquisition; project administration; supervision. All authors have read and approved the final version of the paper.
Supplemental Material
Download PDF (1 MB)Disclosure statement
No potential conflict of interest was reported by the author(s).
Data availability statement
The data that support the findings of this study are available from the corresponding authors upon reasonable request.
Additional information
Funding
References
- Bunn SE. Grand challenge for the future of freshwater ecosystems. Front Environ Sci. 2016;4:1. doi:10.3389/fenvs.2016.00021.
- Qi L, Li RW, Wu YD, et al. Spatial distribution and assembly processes of bacterial communities in Northern Florida freshwater springs. Environ Res. 2023;235:116584. doi:10.1016/j.envres.2023.116584.
- Tang VT, Rene ER, Li QH. Application and performance of eco-bag revetment for water purification in lake environments: a case study from rural China. J Water Supply Res Technol-Aqua. 2019;68(2):149–13. doi:10.2166/aqua.2019.110.
- Woszczyk M, Bechtel A, Cieśliński R. Interactions between microbial degradation of sedimentary organic matter and lake hydrodynamics in shallow water bodies: insights from lake sarbsko (Northern Poland). J Limnol. 2011;70(2):293–304. doi:10.4081/jlimnol.2011.293.
- Zhu WT, Liu J, Li QH, et al. Effects of nutrient levels on microbial diversity in sediments of a eutrophic shallow lake. Front Ecol Evol. 2022;10:909983. doi:10.3389/fevo.2022.909983.
- Hahn MW. The microbial diversity of inland waters. Curr Opin Biotechnol. 2006;17(3):256–261. doi:10.1016/j.copbio.2006.05.006.
- Feng WY, Gao JY, Wei YM, et al. Pattern changes of microbial communities in urban river affected by anthropogenic activities and their environmental driving mechanisms. Environ Sci Eur. 2022;34(1):93. doi:10.1186/s12302-022-00669-1.
- Yannarell AC, Kent AD, Lauster GH, et al. Temporal patterns in bacterial communities in three temperate Lakes of different trophic status. Microb Ecol. 2003;46(4):391–405. doi:10.1007/s00248-003-1008-9.
- Zhu CM, Zhang JY, Nawaz MZ, et al. Seasonal succession and spatial distribution of bacterial community structure in a eutrophic freshwater lake, lake Taihu. Sci Total Environ. 2019;669:29–40. doi:10.1016/j.scitotenv.2019.03.087.
- Zheng YH, Su ZG, Dai TJ, et al. Identifying human-induced influence on microbial community: a comparative study in the effluent-receiving areas in Hangzhou Bay. Front Environ Sci Eng. 2019;13(6):90. doi:10.1007/s11783-019-1174-8.
- Navarro MB. Large differences in bacterial community composition of nearby shallow Lakes surrounded by Nothofagus pumilio forest in Patagonia(Argentina). J Plankton Res. 2022;44:350–364. doi:10.1093/plankt/fbac018.
- Pascault N, Roux S, Artigas J, et al. A high-throughput sequencing ecotoxicology study of freshwater bacterial communities and their responses to tebuconazole. FEMS Microbiol Ecol. 2014;90(3):563–574. doi:10.1111/1574-6941.12416.
- Sonthiphand P, Ruangroengkulrith S, Mhuantong W, et al. Metagenomic insights into microbial diversity in a groundwater basin impacted by a variety of anthropogenic activities. Environ Sci Pollut Res Int. 2019;26(26):26765–26781. doi:10.1007/s11356-019-05905-5.
- Nishiguchi MK, Doukakis P, Egan M, et al. DNA isolation procedures. In DeSalle R, Giribet G, Wheeler WC, editors. Techniques in molecular systematics and evolution. Basel: Birkhäuser Verlag. 2002. p. 249–287. doi:10.1007/978-3-0348-8125-8_12.
- Qian Y, Okano K, Kodato M, et al. Dynamics of the prokaryotic and eukaryotic microbial community during a cyanobacterial bloom. Biosci Biotechnol Biochem. 2022;86(1):78–91. doi:10.1093/bbb/zbab179.
- Caporaso JG, Kuczynski J, Stombaugh J, et al. Qiime allows analysis of high-throughput community sequencing data. Nat Methods. 2010;7(5):335–336. doi:10.1038/nmeth.f.303.
- Callahan BJ, McMurdie PJ, Rosen MJ, et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat Methods. 2016;13(7):581–583. doi:10.1038/NMETH.3869.
- Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15(12):550–550. doi:10.1186/s13059-014-0550-8.
- Mandal S, Van Treuren W, White RA, et al. Analysis of composition of microbiomes: a novel method for studying microbial composition. Microb Ecol Health Dis. 2015;26(0):27663. doi:10.3402/mehd.v26.27663.
- Segata N, Izard J, Waldron L, et al. Metagenomic biomarker discovery and explanation. Genome Biol. 2011;12(6):R60. doi:10.1186/gb-2011-12-6-r60.
- Vázquez-Baeza Y, Pirrung M, Gonzalez A, et al. Emperor a tool for visualizing high-throughput microbial community data. Gigascience. 2013;2(1):16. doi:10.1186/2047-217X-2-16.
- Douglas GM, Maffei VJ, Zaneveld JR, et al. PICRUSt2 for prediction of metagenome functions. Nat Biotechnol. 2020;38(6):685–688. doi:10.1038/s41587-020-0548-6.
- Fang JH, Yang RR, Cao QQ, et al. Differences of the microbial community structures and predicted metabolic potentials in the lake, river, and wetland sediments in Dongping lake basin. Environ Sci Pollut Res Int. 2020;27(16):19661–19677. doi:10.1007/s11356-020-08446-4.
- Paul D, Kumbhare SV, Mhatre SS, et al. Exploration of microbial diversity and community structure of Lonar lake: the only hypersaline meteorite crater lake within basalt rock. Front Microbiol. 2015;6:1553. doi:10.3389/fmicb.2015.01553.
- Hu AY, Ju F, Hou LY, et al. Strong impact of anthropogenic contamination on the co-occurrence patterns of a riverine microbial community. Environ Microbiol. 2017;19(12):4993–5009. doi:10.1111/1462-2920.13942.
- Luo JW, Zeng H, Zhou QX, et al. Anthropogenic impacts on the biodiversity and anti-interference ability of microbial communities in Lakes. Sci Total Environ. 2022;820:153264. doi:10.1016/j.scitotenv.2022.153264.
- Camacho A, Peinado R, Santamans AC, et al. Functional ecological patterns and the effect of anthropogenic disturbances on a recently restored Mediterranean coastal lagoon. Needs for a sustainable restoration. Estuar Coast Shelf Sci. 2012;114:105–117. doi:10.1016/j.ecss.2012.04.034.
- Ma TL, Jiang YM, Elbehery AHA, et al. Resilience of planktonic bacterial community structure in response to short-term weather deterioration during the growing season in an alpine lake. Hydrobiologia. 2020;847(2):535–548. doi:10.1007/s10750-019-04118-8.
- Wang HJ, Liu XC, Wang YL, et al. Spatial and temporal dynamics of microbial community composition and factors influencing the surface water and sediments of Urban Rivers. J Environ Sci. 2023;124:187–197. doi:10.1016/j.jes.2021.10.016.
- Zhang JX, Yang YY, Zhao L, et al. Distribution of sediment bacterial and archaeal communities in Plateau freshwater Lakes. Appl Microbiol Biotechnol. 2015;99(7):3291–3302. doi:10.1007/s00253-014-6262-x.
- Bucheli TD, Fent K. Induction of cytochrome-P450 as a biomarker for environmental contamination in aquatic ecosystems. Crit Rev Environ Sci Technol. 1995;25(3):201–268. doi:10.1080/10643389509388479.
- Vera A, Wilson FP, Cupples AM. Predicted functional genes for the biodegradation of xenobiotics in groundwater and sediment at two contaminated naval sites. Appl Microbiol Biotechnol. 2022;106(2):835–853. doi:10.1007/s00253-021-11756-3.
- Higgins CF. Abc transporters – from microorganisms to man. Annu Rev Cell Biol. 1992;8(1):67–113. doi:10.1146/annurev.cellbio.8.1.67.
- ter Beek J, Guskov A, Slotboom DJ. Structural diversity of ABC transporters. J Gen Physiol. 2014;143(4):419–435. doi:10.1085/jgp.201411164.
- Huang H, Chen Y, Yang S, et al. CuO and ZnO nanoparticles drive the propagation of antibiotic resistance genes during sludge anaerobic digestion: possible role of stimulated signal transduction. Environ Sci. 2019;6(2):528–539. doi:10.1016/j.biortech.2022.128253.
- Gold Z, Shelton AO, Casendino HR, et al. Signal and noise in metabarcoding data. PLOS One. 2023;18(5):e0285674. doi:10.1371/journal.pone.0285674.
- Lamb PD, Hunter E, Pinnegar JK, et al. How quantitative is metabarcoding: a meta-analytical approach. Mol Ecol. 2019;28(2):420–430. doi:10.1111/mec.14920.
- Shelton AO, Gold ZJ, Jensen AJ, et al. Toward quantitative metabarcoding. Ecology. 2023;104(2):e3906. doi:10.1002/ecy.3906.