ABSTRACT
In eukaryotes, critical regulation of cell cycle is required to ensure the integrity of cell division. HORMA-containing proteins include various proteins that contain HORMA domain and play important role in the regulation of cell cycle in eukaryotes. Many types of HORMA-containing proteins are found in eukaryotes, but their role in prokaryotes has not been proven. Therefore, we conduct an extensive search in GenBank for HORMA-containing proteins in prokaryotes to compare HORMA domain structure and architecture across eukaryotes and prokaryotes. Strikingly, genome sequencing for many prokaryotic organisms reveals that HORMA domain is present in many bacterial genomes and only two archaeal genomes. We perform sequence alignment and phylogenetic analysis to trace the evolutionary link between HORMA domain in prokaryotes and eukaryotes. HORMA domain in prokaryotes appears to vary in sequence and architecture. Interestingly, seven bacterial HORMA-containing proteins and the two archaeal HORMA-containing proteins showed close relationships with eukaryotic HORMA-containing proteins. Additionally, we uncovered remarkable close relationships between HORMA-containing protein from Chlamydia trachomatis and eukaryotic MAD2 proteins. Our results provide insights into evolutionary relationships between prokaryotic and eukaryotic systems, which facilitate our understanding of the evolution of cell cycle regulation mechanisms.
Introduction
Cell cycle in eukaryotes is a complicated process that involves multiple events performed by many proteins with various functions. In all cellular organisms, organization of the cell cycle and DNA repair is controlled by crucial mechanisms. HORMA domain-containing proteins are a group of proteins that are known to be pivotal in cell division regulation and DNA repair. This group has been identified in Schizosaccharomyces and derives its name from the initial letters of the names of three types: HOP1, REV7 and mitotic arrest deficient 2 (MAD2) [Citation1,Citation2].
HOP1 protein is essential for the synaptonemal complex assembly and chromosome synapsis in meiosis. In Arabidopsis, ASY1 is a homolog to HOP1 in Saccharomyces cerevisiae, which is involved in meiosis in male and female gametophytes [Citation3,Citation4]. In humans, other types of HORMA proteins, HORMAD1 and HORMAD2, are known to be involved in meiotic progression. They are also homologs to Saccharomyces HOP1 [Citation5–Citation7]. Both HORMAD1 and HORMAD2 are important for synapsis surveillance and crucial for the male mid-pachytene checkpoint and the female meiotic prophase checkpoint [Citation6,Citation8].
REV7 is the DNA polymerase zeta (Pol ζ) processivity subunit which interacts with REV3, the catalytic subunit in Pol ζ [Citation9]. REV3 and REV7 are error-prone DNA polymerases involved in translation DNA synthesis polymerases to repair DNA damage during DNA replication [Citation10,Citation11]. Names of some HORMA-containing proteins can be different in the other eukaryotes. For instance, MAD2 is named MAD2L1 or MAD2A in other organisms. REV7 is also known as MAD2L2 or MAD2B. These proteins perform overlapping roles in the mitotic spindle assembly checkpoint, and they delay the initiation of anaphase until all chromosomes are correctly arranged in the cell midline at metaphase. Mutations in MAD2L1 and 2 in humans are related to various types of cancer and numerical chromosomal abnormalities [Citation12,Citation13]. The structure and function of REV7 and MAD2 genes in plants are highly conserved. REV7 in Arabidopsis is known to be involved in damage-tolerance mechanisms through translesion DNA synthesis [Citation14,Citation15].
Specific types of Autophagy-related (ATG) genes that control autophagy in cells have recently been discovered to contain HORMA domain [Citation16]. Autophagy-related 13 is an autophagy factor required for autophagosome formation and mitophagy. This gene was originally identified in yeast, and it has clear orthologs in Arabidopsis [Citation17], Drosophila melanogaster [Citation18], Caenorhabditis elegans [Citation19] and in humans [Citation20].
Although C. elegans has a homolog to MAD2 in yeast, HORMA proteins in the worm appear to be distinct from other eukaryotes [Citation21,Citation22]. C. elegans has four distinct HORMA proteins: Him-Three Paralog 1 (HTP1), HTP2, HTP3 and High Incidence of Males (HIM3). HTP1 and HTP2 are involved in regulation of centriole-centriole cohesion, while HTP3 performs a critical role in meiotic DNA double-strand break formation and synapsis. High Incidence of Males shares overlapping and divergent roles with HTP1, HTP2 and HTP3 in homologous chromosome segregation, DNA break formation and recombination, regulation of centriole-centriole cohesion and synaptonemal complex assembly [Citation21–Citation23].
Cell cycle control in prokaryotes is less complexity than in eukaryotes. However, there are evidences about the existence of cell-cycle regulation in bacteria which cell cycle is arrested before DNA replication initiation or after chromosome segregation. Unsegregated chromosomes block assembly of the FtsZ ring, the essential proteins that triggers the accumulation membrane and cell wall proteins between the dividing bacterial cells [Citation24–Citation27] during cytokinesis in prokaryotes [Citation28]. Additionally, previous studies indicate that some crenarchaea species initiate replication from multiple origins of replication as seen in eukaryotes [Citation29,Citation30]. Moreover, checkpoint-like regulation also is seen in crenarchaea via Cdv proteins that form intracellular structures during constriction. These proteins are induced in normal cell division at the initiation of genome segregation to carry out cytokinesis. In response to DNA damage, Cdv proteins are down-regulated and subsequently inhibit cell division. Cdv-based cell division systems bear some similarity to eukaryotic checkpoint systems, suggesting similar regulation mechanisms [Citation31].
Molecular mechanisms that regulate mitosis or meiosis in eukaryotes are apparently different from that in cell cycle control in prokaryotes. The high level of organization and the complexity of cell division in eukaryotes relative to prokaryotes prompts questions regarding the origin and evolution of eukaryotic cell division mechanisms. Advances in comparative genomics and bioinformatic tools may potentially improve our understanding of the evolution of organisms. It is commonly known that many proteins which are involved in cell division are conserved among prokaryotes and eukaryotes, such as ATPase family proteins [Citation32], several key enzymes of the apoptotic machinery [Citation33] and structural maintenance of chromosomes complex [Citation34]. However, no experimental reports describe the role of HORMA-containing proteins in cell division in prokaryotes. Nevertheless, sequencing for some bacterial genomes demonstrates that a conserved HORMA domain is present in many bacterial genomes [Citation35]. Therefore, we aim to investigate the structure and architecture of HORMA domain through a broad survey in GenBank for HORMA-containing proteins in prokaryotes. Comparison and phylogenetics of prokaryotic and eukaryotic proteins will help in understanding of the evolution of cell division machinery in prokaryotic and eukaryotic systems.
Experimental procedures
HORMA-containing proteins were retrieved by sequence homology searches in NCBI (http://www.ncbi.nlm.nih.gov/genome) and the UniProt database (http://www.uniprot.org/). The HORMA protein sequences of human (MAD2, NP_002349.1; REV7, NP_006332.3; HORMAD1, NP_001186758.1; HORMAD2, NM_001329457.1), S. cerevisiae (HOP1, NP_012193.3; REV7, NP_012127.1; MAD2, NP_012504.3; ATG13, AJV99010.1), Arabidopsis thaliana (REV7, NM_101522.4; MAD2, NM_001203049.1; ASY1, NM_101522.4; ASY2, NM_119372.3, ATG13, NM_114819.3) were utilized as query sequences in NCBI BlastP and UniProt to retrieve HORMA-containing proteins in eukaryotes and prokaryotes. Among all hits found, HORMA-containing proteins from specific taxonomically-representative eukaryotic organisms were selected to investigate HORMA domain structure. HORMA-containing proteins from the worms; C. elegans, the brown algae; Ectocarpus siliculosus, the euglena; Trypanosoma cruzi and the amoeba; Tieghemostelium lacteum were selected to represent eukaryotic organisms in addition to humans, yeast and Arabidopsis. All HORMA-containing proteins discovered in these eukaryotes were selected for further investigation.
Relative to NCBI, more HORMA-containing proteins in prokaryotic genomes were found in the UniProt database. From 111 hits found in bacterial genomes in UniProt, 20 bacteria were selected to represent the large taxonomic divisions of bacteria. These include HORMA-containing protein from each of the following: Chlamydia trachomatis, Polaribacter dokdonensis DSW-5, Clostridium sp. IBUN13A, Pseudomonas aeruginosa, Bacteroidales bacterium, Desulfovibrio africanus, Candidatus Delongbacteria bacterium, Streptomyces purpurogeneiscleroticus, Achromobacter spanius, Sulfitobacter geojensis, Serratia marcescens subsp. marcescens, Roseomonas rhizosphaerae, Parvibaculum lavamentivorans, Rubricoccus marinus, Sphingopyxis bauzanensis, Planctomycetaceae bacterium, Streptomyces sp., Rickettsiales bacterium, Cytophagales bacterium and Flavobacterium granuli. Only one HORMA-containing protein was found in each of the selected bacteria. A homology search in UniProt revealed only two HORMA-containing proteins in archaea. Both Halorubrum ezzemoulense and Halorientalis regularis contained only one protein. lists accession numbers for all selected proteins.
Table 1. HORMA-containing proteins from taxonomically representative eukaryotic and prokaryotic organisms. HORMA-containing proteins from human, yeast and Arabidopsis were used as query sequences in NCBI BlastP and UniProt to retrieve HORMA-containing proteins in eukaryotes and prokaryotes. Twenty-nine eukaryotic proteins were selected from NCBI and 22 prokaryotic proteins were selected from UniPro for phylogenetic analysis and investigation of HORMA-containing proteins structure and architecture (Accession numbers shown in the 3rd column).
All protein sequences were aligned by MUSCLE version 3.8 [Citation36]. The multiple sequence alignment results were imported to the UniGene software package [Citation37]. The alignment results were assessed to remove the repeated and partial sequences. The phylogenetic tree was constructed based on the alignment of 29 eukaryotic, 20 bacterial and two archaeal HORMA-containing protein sequences. MEGA 7.0.26 [Citation38] was used to construct phylogenetic trees through neighbor joining (NJ) [Citation39] and maximum likelihood (ML) [Citation40] methods using default settings. The constructed phylogenetic tree was also visualized by MEGA 7.0.26.
To compare HORMA domain structure, HORMA domain sequences from each of the selected proteins were determined using the InterPro database [Citation41] and aligned using MUSCLE version 3.8. HORMA domain architecture was retrieved in all protein sequences using ScanProsite [Citation42]. In prokaryotic proteins, HORMA domain architecture did not appear through ScanProsite searches except Chlamydia trachomatis, Cytophagales bacterium and Rickettsiales bacterium. Therefore, HORMA domain architecture in other prokaryotes was drawn using MyDomains- Image Creator (https://prosite.expasy.org/cgi-bin/prosite/mydomains/) [Citation43] based on protein structure information from InterPro.
Results and discussion
Phylogenetic relationships
All selected eukaryotic HORMA-containing proteins belong to one of the following HORMA-containing protein types; HOP1, REV7, MAD2, ATG13, ASY1, ASY2, HORMAD1 and HORMAD2, except one HORMA-containing protein from each of the following protozoa: Trypanosoma cruzi and Tieghemostelium lacteum; these were uncharacterized. All prokaryotic HORMA-containing proteins were uncharacterized except glycine dehydrogenase (GLDC) from Polaribacter dokdonensis and glycosyl transferase (GTR) protein from Bacteroidales bacterium. To clearly investigate phylogenetic relationships, proteins which are known by multiple names in NCBI or UniProt were assigned a unified name for this study. For instance, MAD2A and MAD2L1 were named “MAD2”, whereas MAD2L2 and MAD2B were named “REV7”. Protein sequences that are present in NCBI without clear protein names or which are named “hypothetical protein or HORMA protein” were assigned the names from UniProt for this study. Other proteins without clear protein names in both NCBI and UniProt were referred to as HORMA-containing proteins. To investigate the relationships between HORMA-containing proteins in prokaryotic and eukaryotic systems, we conducted a phylogenetic analysis for 29 HORMA-containing proteins from eukaryotes and 22 HORMA-containing proteins from prokaryotes using two methods, NJ and ML.
In NJ phylogeny ()), HORMA-containing proteins were divided into two major groups. The first group was composed primarily of eukaryotic proteins, with two archaeal proteins and five bacterial proteins. This group was divided into two subgroups and the first subgroup was split into two branches. The first branch consists of all eukaryotic REV7 and MAD2 proteins, in addition to the Tieghemostelium lacteum uncharacterized HORMA-containing protein and two bacterial HORMA-containing proteins from the Chlamydia trachomatis and Rickettsiales bacterium. The Chlamydia trachomatis HORMA-containing protein descended from an interior node with S. cerevisiae MAD2 protein with a bootstrap value of 59%. The Rickettsiales bacterium formed a separate branch, which was supported with a 78% bootstrap value from the first branch that included all MAD2 and REV7 proteins. Arabidopsis thaliana ASY1 and E. siliculosus HOP1 descended together from second subgroup of the first branch. The second branch consisted of Trypanosoma cruzi uncharacterized HORMA-containing protein, S. cerevisiae HOP1, C. elegans HTP1, Arabidopsis thaliana ASY2 and ATG13, and E. siliculosus ATG13. The second subgroup consisted of HORMA-containing proteins from the two archaea, Halorubrum ezzemoulense and Halorientalis regular, along with human HORMAD1 and HORMAD2 and C. elegans HIM3 with three bacterial HORMA-containing proteins from Clostridium sp., Candidatus Delongbacteria and Flavobacterium granuli.
Figure 1. Phylogenetic phenogram tree produced from the alignment of 29 eukaryotic and 22 prokaryotic HORMA-containing proteins. Multiple sequence alignment was constructed by MUSCLE 3.8 and phylogenetic trees generated using MEGA 7.0.26 software. (a) Phylogenetic tree constructed by the NJ method, (b) Phylogenetic tree constructed by ML method. Numbers on nodes are bootstrap percentages supporting a given partitioning. The proteins are designated by the genus name followed by the name of protein. Uncharacterized proteins are named by genus name followed by “HORMA”. Streptomyces purpurogeneiscleroticus # A0A0M8ZCY4 named as “StreptomycesHORMA”, Streptomyces sp. # A0A2A2Z569 named as “StreptomycesHORMA1”. Accession numbers for all proteins are listed in . Red, blue and green circles indicate for eukaryotic, bacterial and archaeal HORMA-containing proteins, respectively.
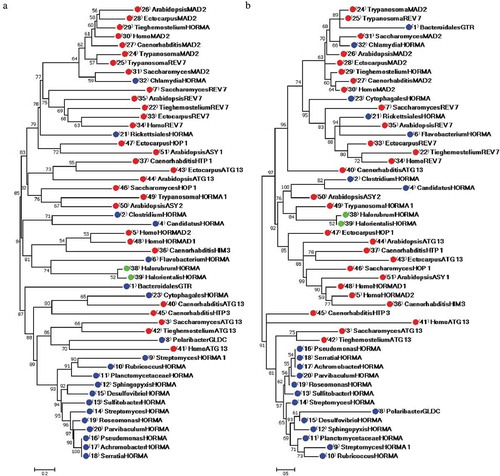
The second major group in the NJ tree consisted of 15 bacterial and five eukaryotic HORMA-containing proteins. This group contained two subgroups. The first subgroup was comprised of C. elegans ATG13 and HORMA-containing protein from the Cytophagales bacterium, which descended from an interior node supported with a 65% bootstrap value. C. elegans HTP3 protein comprised a separate branch from this subgroup, with a bootstrap support value of 80%. The second subgroup branched into two clusters. The first cluster consisted of three eukaryotic ATG13 proteins from humans, S. cerevisiae and Tieghemostelium lacteum with the bacterial protein GLDC from Polaribacter dokdonensis. The second cluster consisted of 12 bacterial proteins, including Rubricoccus marinus, Planctomycetaceae bacterium, Sphingopyxis bauzanensis, Desulfovibrio africanus, Sulfitobacter geojensis, Roseomonas rhizosphaerae, Parvibaculum lavamentivorans, Pseudomonas aeruginosa, Achromobacter spanius, Serratia marcescens, Streptomyces sp. and Streptomyces purpurogeneiscleroticus. Bacteroidales bacterium GTR formed a separate branch from the second major group in the NJ tree, with an 87% bootstrap support value.
The same relationships between prokaryotic and eukaryotic HORMA-containing proteins were nearly confirmed by the ML phylogenetic tree ()). The first group was divided into two subgroups, which demonstrated the same clustering for eukaryotic, bacterial and archaeal proteins, with some exceptions. Unlike the NJ tree, the two bacterial proteins; Bacteroidales bacterium GTR and Cytophagales bacterium HORMA-containing protein, with C. elegans ATG13 clustered with the first group in ML tree instead of clustering in the second group in NJ tree. Bacteroidales bacterium GTR and the Cytophagales bacterium HORMA-containing protein were clustered with REV7 and MAD2 proteins in the ML tree, whereas C. elegans ATG13 formed a separate branch from the first subgroup in the ML tree. Trypanosoma cruzi REV7 and MAD2 were the closest proteins to Bacteroidales bacterium GTR, with bootstrap support value of 89% in the ML tree. Additionally, the HORMA-containing protein from Flavobacterium granuli appeared to be the closest protein to the archaeal proteins in the NJ tree, while it clustered with the eukaryotic REV7 in the ML tree. However, the HORMA-containing protein from Trypanosoma cruzi was the closest to the two archaea in the ML tree. Clustering of Polaribacter dokdonensis GLDC also varied between the NJ and ML trees. In the NJ tree, Polaribacter dokdonensis was the closest to human ATG13 protein, while in the ML tree, this protein clustered with the 12 bacterial proteins in the second group. Unlike the NJ tree, Arabidopsis thaliana ASY1 and E. siliculosus HOP1 clustered with the second subgroup in the ML tree rather than clustering with proteins in the first subgroup in the first group of the NJ tree.
The close relationships between the Chlamydia trachomatis HORMA-containing protein and S. cerevisiae MAD2 protein in the NJ tree were also confirmed in the ML tree with a bootstrap value of 66%. The close relationship between HORMA-containing proteins from the two archaea, Halorubrum ezzemoulense and Halorientalis regular, was confirmed by both the NJ and ML tree. These proteins descended from an interior node with a bootstrap support value of 52% in the NJ and ML trees. Similarly, the relationships between the two bacterial HORMA-containing proteins from Clostridium sp. and Candidatus Delongbacteria, and the two archaeal proteins were confirmed by both phylogenetic trees. These bacterial proteins descended from an interior node with bootstrap support value of 71% in the NJ and 82% in the ML tree. However, clustering of eukaryotic proteins in the subgroup that included these bacterial and archaeal proteins varied between the two phylogenetic trees. In the NJ tree, this subgroup included C. elegans HIM3 and human HORMAD1 and HORMAD2, in addition to Flavobacterium granuli. Conversely, the same subgroup in the ML tree included Arabidopsis ASY2 and HORMA-containing protein from Trypanosoma cruzi, together with Clostridium sp., Candidatus Delongbacteria and the two archaeal proteins.
HORMA domain architecture
The HORMA domain architecture in all the proteins investigated was retrieved and drawn through PROSITE. illustrates HORMA domain architecture for some eukaryotic and prokaryotic proteins. In eukaryotes, HORMA domain size ranged from 177 amino acids in Trypanosoma cruzi MAD2 protein, to 264 amino acids in C. elegans HTP3 ()). Domain architecture varied across various eukaryotic HORMA-containing proteins. MAD2 and REV7 proteins appeared to be composed entirely of HORMA domain, except Tieghemostelium lacteum REV7, which contained HORMA domain in its C-terminal region. Other eukaryotic HORMA-containing proteins were longer than MAD2 and REV7 and had different architecture, which contained HORMA domain in the N-terminal region. ASY2 protein from Arabidopsis was unique HORMA-containing protein (1399 aa length). The NCBI and UniProt BLAST searches did not reveal any homologs for the ASY2 protein in eukaryotes or prokaryotes. The Arabidopsis ASY1 protein contained SWIRM domain with 99 amino acids in addition to HORMA domain. SWIRM domain presents in proteins that are involved in chromatin modifications and remodeling. Previous studies have reported that SWIRM domain may be involved in the assembly of chromatin-protein complexes [Citation44]. ATG13 in eukaryotes also contained ATG13 domain, which overlapped with HORMA domain in all ATG13 proteins in the N-terminal region. ATG13 HORMA appears to has structural plasticity even at the N terminus of the protein. This plasticity is due to the role of ATG13 in regulating spatiotemporal assembly of the components in the autophagy induction complex [Citation45].
Figure 2. HORMA domain architecture in some HORMA-containing proteins from (a) eukaryotes and (b) prokaryotes. In eukaryotes, HORMA domain size ranged from 177 to 264 aa. Prokaryotic HORMA-containing proteins were classified based on domain size and multiple sequence alignment results into three groups. HORMA-containing proteins in the first group ranged in size from 165 to 167 aa and HORMA domains in these group ranged from 148–159 aa. The second group contained larger HORMA domains (from 196 to 224 aa), while the third group contained smaller HORMA domains (from 52 to 145 aa). ScanProsite search used to retrieve domain architecture in all eukaryotic HORMA-containing proteins and three bacterial proteins; Chlamydia trachomatis, Cytophagales bacterium and Rickettsiales bacterium. HORMA domains architecture in other prokaryotic proteins were drawn using MyDomains- Image Creator based on proteins structure information in InterPro.
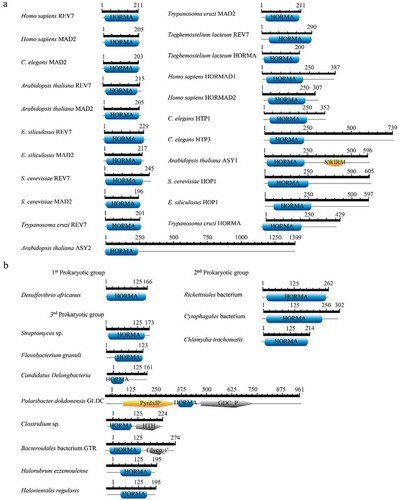
Based on HORMA domain size and multiple sequence alignment results, prokaryotic HORMA-containing proteins were classified into three groups. The first group contained 11 proteins ranging in size from 165 to 167 amino acids, while HORMA domains in these group ranged from 148 to 159 amino acids. This group contained 11 HORMA-containing proteins: Pseudomonas aeruginosa, Desulfovibrio africanus, Streptomyces purpurogeneiscleroticus, Achromobacter spanius, Sulfitobacter geojensis, Serratia marcescens subsp. marcescens, Roseomonas rhizosphaerae, Parvibaculum lavamentivorans, Rubricoccus marinus, Sphingopyxis bauzanensis and Planctomycetaceae bacterium. Bacterial taxonomy demonstrated that all members of this group belong to Proteobacteria except Rubricoccus marinus which belong to Rhodothermaeota and Planctomycetaceae bacterium which belong to Planctomycetes, Verrucomicrobia and Chlamydiae (PVC) superphylum [Citation46]. Because of HORMA domain in the first prokaryotic group the first group shares similar size and architecture, it is represented in ) by one protein (Desulfovibrio africanus protein). The second group consisted of larger proteins and larger HORMA domains. As displayed in , proteins in the second group ranged from 214 to 302 amino acids, while HORMA domains ranged from 196 to 224 amino acids. This group was comprised of Chlamydia trachomatis, Rickettsiales bacterium and Cytophagales bacterium. These three bacteria belong to the three taxonomic divisions of bacteria; PVC, Proteobacteria and Bacteroidetes, respectively [Citation46]. The third group contained smaller HORMA domains (from 52 to 145 amino acids) and varied protein sizes and domain architectures. This group contained five bacterial proteins and the two archaeal proteins. The bacterial proteins in the third group included Flavobacterium granuli, Candidatus Delongbacteria, Streptomyces sp., Clostridium sp., GLDC from Polaribacter dokdonensis and GTR from Bacteroidales bacterium. The last three bacteria are members of Bacteroidetes, Streptomyces sp. is member of Actinobacteria, and Clostridium sp. is member of Firmicutes, where Candidatus Delongbacteria is an unclassified bacterium [Citation46]. Clostridium sp. HORMA-containing protein, GLDC and GTR were multiple protein domains. HORMA-containing protein from Clostridium sp. contained the helix-turn-helix domain, the DNA-binding motif that exists in transcription regulatory proteins in many bacteria [Citation47]. In addition to HORMA domain, GTR Bacteroidales bacterium contained a domain of galactosyltransferase, which is involved in the biosynthesis of different glycoconjugates and saccharide structures [Citation48]. Stemphylium lycopersici bacterium owns GTR gene that appears to contain also HORMA domain beside the domain of galactosyltransferase. Polaribacter dokdonensis GLDC contained two domains; glycine cleavage system P-protein and pyridoxal phosphate-dependent transferase, in addition to HORMA domain. Glycine cleavage system P-protein domain catalyzes the degradation of glycine. It participates in glycine, serine and threonine metabolism [Citation49]. In eukaryotes, GLDC gene is a mitochondrial gene that shares conserved sequences with its bacterial homologs [Citation50,Citation51]. In contrast to Polaribacter dokdonensis GLDC, the homologs GLDC genes in eukaryotes or other prokaryotes appear not to contain HORMA domain.
Bacteria from PVC superphylum and some archaebacteria appear to have a distinct cell division mechanisms that seem to be exceptions to the dominant mode of prokaryotic cell division by binary fission [Citation52]. Cell division on Planctomycetes, Chlamydia and many archaebacteria is atypical since it occurs in the absence of a sequence homologue of FtsZ [Citation53] and peptidoglycan [Citation54,Citation55]. Moreover, both Planctomycetes and Chlamydia has condensed DNA [Citation56]. Nevertheless, Chlamydiales generally divide by binary fission [Citation57] while Planctomycetes divide by budding process [Citation58]. Recently, Chlamydia has been shown to divide by a polarized cell division similar to the budding process observed in Planctomycetes [Citation59]. Chlamydia trachomatis is sexually transmitted human pathogen. Cell division of this pathogen interests the scientists as the understanding of Chlamydia cell cycle may result in novel therapeutic antimicrobial compounds. Planctomycetes phylum possess a distinct intracellular compartmentalization and unique metabolism patterns apparently comparable to the eukaryotic mitochondrion [Citation60]. The exact mechanisms of cell division in Chlamydia and Planctomycetes are still unknown yet. Some biologists currently claim that bacteria of PVC group are considered evolutionary intermediates in the prokaryotes to eukaryotes transition [Citation56].
HORMA-containing proteins in the first and second prokaryotic groups were almost entirely composed of HORMA domain, except Cytophagales bacterium. This protein with Candidatus Delongbacteria, Clostridium sp., Streptomyces sp. and Bacteroidales bacterium contained HORMA domain in its N-terminal region. However, Flavobacterium granuli and the two archaeal proteins contained HORMA domain in C-terminal region. Polaribacter dokdonensis GLDC enzyme had a unique architecture, in which HORMA domain existed in the central region between the other two domains (glycine cleavage system P-protein and pyridoxal phosphate-dependent transferase) ().
HORMA domain multiple sequence alignment
To provide insights into the structure of HORMA domain, sequence alignment was performed exclusively for HORMA domain sequences from all investigated proteins (). Multiple sequence alignment revealed that the sequence of HORMA domain was highly conserved within the first prokaryotic group. Chlamydia trachomatis HORMA-containing also demonstrated a high degree of sequence conservation with the eukaryotic MAD2 proteins, Trypanosoma cruzi REV7 and HORMA-containing proteins from Tieghemostelium lacteum. Limited conservation was displayed by HORMA domain sequences from all C. elegans proteins, all ATG13 proteins and HORMA domain from each of; Clostridium sp.,Bacteroidales bacterium GTR, Candidatus Delongbacteria, Polaribacter dokdonensis GLDC and Flavobacterium granuli. These proteins also revealed less conservation with HORMA domain sequences from other prokaryotes and eukaryotes. HORMA domains from the two archaeal proteins also demonstrated a lower degree of conservation with bacterial and eukaryotic proteins. Remarkably, the sequence of the HORMA domain from Chlamydia trachomatis was closely related to the sequences of eukaryotic MAD2 proteins. Therefore, further sequence alignment was performed for HORMA domain from MAD2 proteins and HORMA domain from Chlamydia trachomatis to closely compare the HORMA domain structure. As shown in , the HORMA domain sequences of Chlamydia trachomatis was highly conserved with MAD2 protein sequences from unicellular and multicellular eukaryotes. The NCBI BLASTP search revealed a 43% sequence identity between HORMA domain from Chlamydia trachomatis and S. cerevisiae MAD2, and a 42% sequence identity between HORMA domain from Chlamydia trachomatis and Arabidopsis MAD2.
Figure 3. Multiple sequence alignments of HORMA domains from 29 eukaryotic, 20 bacterial and two archaeal HORMA-containing proteins. The proteins are designated by the genus name followed by the name of protein. Uncharacterized proteins are named by genus name followed by “HORMA”. Streptomyces purpurogeneiscleroticus # A0A0M8ZCY4 named as “StreptomycesHORMA”, Streptomyces sp. # A0A2A2Z569 named as “StreptomycesHORMA1”. Alignment was performed using MUSCLE 3.8 and visualized by UniGene.

Figure 4. Multiple sequence alignments of conserved HORMA domain in some eukaryotic MAD2 protein sequences along with HORMA domain from Chlamydia trachomatis protein. The proteins are designated by the genus name followed by the name of protein. Alignment was performed using MUSCLE 3.8 and visualized by UniGene.
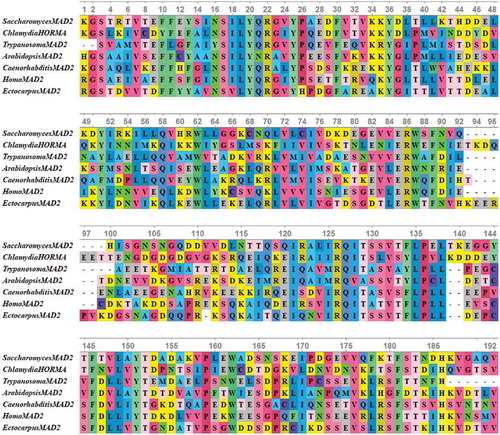
MAD2 functions as an effective molecule in the mitotic spindle assembly checkpoint. It ensures that all chromosomes are correctly attached to kinetochores before the initiation of anaphase. This function can be performed through switching between two states for MAD2. In the presence of unattached kinetochores, MAD2 protein is in its active state and can arrest cell cycle in metaphase through establishing an inhibitory mitotic checkpoint complex, which binds to the anaphase-promoting complex activator Cdc20 and locks it. In the inactive state of MAD2, it releases Cdc20 and allows the initiation of anaphase [Citation61]. The conserved sequence and architecture of Chlamydia trachomatis HORMA domain with eukaryotic MAD2 proteins introduce other evidence to the uniqueness of cell cycle of Chlamydia trachomatis. This evidence indicates for existence of a similar checkpoint before segregation of Chlamydial chromosome unlike dominant bacterial division where the chromosome separates during replication. This supports the suggestion that the eukaryotic mitotic spindle checkpoint might evolve from a similar cell cycle control mechanism during Chlamydial cell division. Alignment of eukaryotic HOP1, REV7 and MAD2 sequences, along with gene co-expression and protein–protein interaction data, provide evidence that the MAD2 gene may have evolved earlier than the REV7 and HOP1 genes, which the single-celled eukaryotes only possess the MAD2 gene [Citation62].
Altogether, the classification of prokaryotic HORMA-containing proteins into three groups was nearly exposed by phylogenetic trees. The 11 bacterial proteins in the first prokaryotic group, which shared similar HORMA domain sequences and architecture, clustered together in the NJ and ML trees. The second prokaryotic group appeared to be the closest bacterial proteins to the eukaryotic proteins. They shared similar sequences and architecture with eukaryotic HORMA domains and clustered with eukaryotic proteins in both the NJ and ML trees. Interestingly, HORMA domain from Chlamydia trachomatis was the most similar prokaryotic sequence relative to eukaryotic MAD2 proteins in both phylogenetic trees. This relationship is supported by previous comparative genomics that revealed that chlamydiae genomes are most similar to many plant proteins [Citation63]. Furthermore, these similar proteins are obtained from horizontal gene transfer between chlamydiae and primary photosynthetic eukaryotes [Citation64]. This similarity suggests a close common ancestry among plants, cyanobacteria and Chlamydia [Citation65]. 16S ribosomal RNA data has revealed that Chlamydiales and the early eukaryotes (Parachlamydia amoebophila) diverged from the common ancestor about 700 million years ago [Citation66]. The third prokaryotic group was composed of diverse HORMA domain sequences and architecture. Nevertheless, bacterial and archaeal proteins in this group demonstrated close phylogenetic relationships with specific eukaryotic HORMA-containing proteins. Our results suggest evolutionary relations between prokaryotic proteins in the second and third groups and the eukaryotic HORMA-containing proteins. The position of the two archaeal proteins with bacterial and eukaryotic proteins reflect the evolutionary relationships among the three domains; Bacteria, Archaea and Eukarya. These relationships were previously supported by many studies [Citation67–Citation69]. Nevertheless, the minimal information about HORMA domain in the sequenced archaeal genomes limits our knowledge about the evolution of HORMA domain from archaeal to eukaryotes. Therefore, the role of HORMA domain in cell division in archaea and bacteria must be explained through experimental evidence.
Conclusion
HORMA domain presents in a wide range of proteins that are involved in regulating the cell cycle. Its role in the cell cycle has been investigated extensively in eukaryotes. However, the role of HORMA domain in cell division in prokaryotes is still unclear. Recently, sequencing of many bacterial and archaeal genomes has revealed that HORMA domain is present in many prokaryotes. By exploiting bioinformatic tools and HORMA-containing protein sequences from NCBI and UniProt databases, this study investigates the phylogenetic relationships between prokaryotic and eukaryotic HORMA-containing proteins, with an emphasis on the structure and architecture of prokaryotic HORMA-containing proteins. Our results indicate that many prokaryotic genomes contain HORMA-containing proteins. One HORMA-containing protein was found in each of bacterial and archaeal genome. Interestingly, prokaryotic HORMA domain varied in sequences and architectures between the different organisms. Furthermore, phylogenomic analysis reveals an evolutionary connection between some bacterial and eukaryotic HORMA-containing proteins. Relationships between Chlamydia and six other bacterial proteins with eukaryotic HORMA-containing proteins provides convincing evidence for this evolutionary link. Further experimental research is necessary to explain the functional diversification of HORMA domain in prokaryotes; this will potentially facilitate the construction of the evolutionary scenario of HORMA-containing genes.
Disclosure statement
No potential conflict of interest was reported by the author.
References
- He X, Patterson TE, Sazer S. The Schizosaccharomyces pombe spindle checkpoint protein mad2p blocks anaphase and genetically interacts with the anaphase-promoting complex. Proc Natl Acad Sci U S A [Internet]. 1997;94(15):7965–7970. [ cited 2018 Jun 9]. Available from http://www.ncbi.nlm.nih.gov/pubmed/9223296
- Aravind L, Koonin EV. The HORMA domain: a common structural denominator in mitotic checkpoints, chromosome synapsis and DNA repair. Trends Biochem Sci [Internet]. 1998;23:284–286. [ cited 2018 Jun 9]. Available from http://www.ncbi.nlm.nih.gov/pubmed/9757827
- Caryl AP, Armstrong SJ, Jones GH, et al. A homologue of the yeast HOP1 gene is inactivated in the Arabidopsis meiotic mutant asy1. Chromosoma [Internet]. 2000;109:62–71.
- Sanchez-Moran E, Osman K, Higgins JD, et al. ASY1 coordinates early events in the plant meiotic recombination pathway. Cytogenet Genome Res 2008;120:302–312.
- Pangas SA, Yan W, Matzuk MM, et al. Restricted germ cell expression of a gene encoding a novel mammalian HORMA domain-containing protein. Gene Expr Patterns 2004;5:257–263.
- Chen Y-T, Venditti CA, Theiler G, et al. Identification of CT46/HORMAD1, an immunogenic cancer/testis antigen encoding a putative meiosis-related protein. Cancer Immun 2005;5:9.
- Xie W, Yang X, Xu M, et al. Structural insights into the assembly of human translesion polymerase complexes. Protein Cell 2012;3:864–874.
- Liu M, Chen J, Hu L, et al. HORMAD2/CT46.2, a novel cancer/testis gene, is ectopically expressed in lung cancer tissues. Mol Hum Reprod 2012;18:599–604.
- Gan GN, Wittschieben JP, Wittschieben B, et al. DNA polymerase zeta (pol ζ) in higher eukaryotes. Cell Res 2008;18:174–183.
- Baynton K, Bresson-Roy A, Fuchs RPP. Distinct roles for Rev1p and Rev7p during translesion synthesis in Saccharomyces cerevisiae. Mol Microbiol. 1999;34:124–133.
- Murakumo Y. The property of DNA polymerase zeta: REV7 is a putative protein involved in translesion DNA synthesis and cell cycle control. Mutat Res. [Internet]. 2002;510:37—44.
- Wang M, Chen L, Chen S, et al. Alleviation of cadmium-induced root growth inhibition in crop seedlings by nanoparticles. Ecotoxicol Environ Saf. 2012;79:48–54.
- Byrne T, Coleman HG, Cooper JA, et al. The association between MAD2 and prognosis in cancer: a systematic review and meta-analyses. Oncotarget [Internet]. 2017;8;102223–102234. Available from http://www.oncotarget.com/fulltext/18414
- Takahashi S, Sakamoto A, Sato S, et al. Roles of Arabidopsis AtREV1 and AtREV7 in translesion synthesis. Plant Physiol Internet] 2005; 138:870–881. Available from. ;:. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1150404/pdf/pp1380870.pdf%0Ahttp://www.ncbi.nlm.nih.gov/pubmed/15908599
- Caillaud M-C, Paganelli L, Lecomte P, et al. Spindle assembly checkpoint protein dynamics reveal conserved and unsuspected roles in plant cell division. PLoS One [Internet]. 2009;4:e6757.
- Jao CC, Ragusa MJ, Stanley RE, et al. A HORMA domain in Atg13 mediates PI 3-kinase recruitment in autophagy. Pnas [Internet]. 2013;110:5486–5491.
- Suttangkakul A, Li F, Chung T, et al. The ATG1/ATG13 protein kinase complex is both a regulator and a target of autophagic recycling in arabidopsis. Plant Cell [Internet]. 2011;23:3761–3779.
- Nezis IP, Shravage BV, Sagona AP, et al. Autophagic degradation of dBruce controls DNA fragmentation in nurse cells during late Drosophila melanogaster oogenesis. J Cell Biol 2010;190:523–531.
- Tian E, Wang F, Han J, et al. epg-1 functions in autophagy-regulated processes and may encode a highly divergent Atg13 homolog in C. elegans. Autophagy 2009;5:608–615.
- Mercer CA, Kaliappan A, Dennis PB. A novel, human Atg13 binding protein, Atg101, interacts with ULK1 and is essential for macroautophagy. Autophagy 2009;5:649–662.
- Zetka MC, Kawasaki I, Strome S, et al., Synapsis and chiasma formation in Caenorhabditis elegans require HIM-3, a meiotic chromosome core component that functions in chromosome segregation. Genes Dev [Internet]. 1999;13: 2258–2270. [ cited 2018 Jun 10].
- Kim Y, Rosenberg SC, Kugel CL, et al. The chromosome axis controls meiotic events through a hierarchical assembly of HORMA domain proteins. Dev Cell [Internet]. 2014;31:487–502.
- Vader G, Musacchio A. HORMA domains at the heart of meiotic chromosome dynamics. Dev Cell [Internet]. 2014;31:389–391.
- Autret S, Levine A, Holland IB, et al. Cell cycle checkpoints in bacteria. Biochimie [Internet]. 1997;79;549–554. Available from http://www.sciencedirect.com/science/article/pii/S0300908497820020
- Ward D, Newton A. Requirement of topoisomerase IV parC and parE genes for cell cycle progression and developmental regulation in Caulobacter crescentus. Mol Microbiol 1997;26:897–910.
- Britton RA, Powell BS, Dasgupta S, et al. Cell cycle arrest in Era GTPase mutants: A potential growth rate-regulated checkpoint in Escherichia coli. Mol Microbiol. 1998;27:739–750.
- Trusca D, Scott S, Thompson C. Bacterial SOS checkpoint protein SulA inhibits polymerization of purified FtsZ cell division protein. J Bacteriol.1998;180:3946–3953. Available from https://jb.asm.org/content/180/15/3946.long
- Bi E, Lutkenhaus J. FtsZ ring structure associated with division in Escherichia coli. Nature [Internet]. 1991;354:161.
- Lundgren M, Andersson A, Chen L, et al. Three replication origins in Sulfolobus species: synchronous initiation of chromosome replication and asynchronous termination. Proc Natl Acad Sci USA [Internet]. 2004;101;7046–7051. Available from http://www.pnas.org/cgi/content/long/101/18/7046
- Robinson NP, Dionne I, Lundgren M, et al. Identification of two origins of replication in the single chromosome of the archaeon sulfolobus solfataricus. Cell. 2004;116:25–38.
- Samson RY, Obita T, Freund SM, et al. A role for the ESCRT system in cell division in archaea. Sceince 2008;322:1710–1713.
- Fagan M, Saier M. P-type ATPases of eukaryotes and bacteria: sequence analyses and construction of phylogenetic trees. J Mol Evol. 1994;38:57–99.
- Koonin E, Aravind L. Koonin EV, Aravind L. Origin and evolution of eukaryotic apoptosis: the bacterial connection. Cell Death Differ. 2002;9:394–404.
- Leman AR, Noguchi E. Linking chromosome duplication and segregation via sister chromatid cohesion. Methods Mol Biol [Internet]. 2014;1170:75–98.
- Burroughs AM, Zhang D, Schäffer DE, Iyer LMAravind L. Comparative genomic analyses reveal a vast, novel network of nucleotide-centric systems in biological conflicts, immunity and signaling. Nucleic Acids Research. 2015;43:10633–10654. doi:10.1093/nar/gkv1267
- Katoh K, Standley DM. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol. 2013;30:772–780.
- Agarwala R, Barrett T, Beck J, et al. Database resources of the national center for biotechnology information. Nucleic Acids Res. 2018;46:D8–13.
- Kumar S, Stecher G, Tamura K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger d atasets. Mol Biol Evol. 2016;33:1870–1874.
- Saitou N, Nei M. The neighbor-joining method - a new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987;4:406–425.
- Felsenstein J, Churchill GA. A hidden markov model approach evolution to variation among sites in rate of evolution. Mol Biol Evol. 1996;13:93–104.
- Finn RD, Attwood TK, Babbitt PC, et al. InterPro in 2017-beyond protein family and domain annotations. Nucleic Acids Res. 2017;45:D190–9.
- de Castro E, Sigrist CJA, Gattiker A, et al. ScanProsite: detection of PROSITE signature matches and ProRule-associated functional and structural residues in proteins. Nucleic Acids Res [Internet]. 2006;34:W362–5.
- Hulo N, Bairoch A, Bulliard V, et al. The 20 years of PROSITE. Nucleic Acids Res. 2008;36:245–249.
- Aravind L, Iyer LM. The SWIRM domain: a conserved module found in chromosomal proteins points to novel chromatin-modifying activities. Genome Biol [Internet]. 2002;3;research0039. Available from http://www.ncbi.nlm.nih.gov/pubmed/12186646
- Popelka H, Klionsky DJ. The molecular mechanism of Atg13 function in autophagy induction: what is hidden behind the data? Autophagy [Internet]. 2017;13:449–451.
- Woese CR. Bacterial evolution. Microbiol Rev. [Internet]. 1987;51;221–271. Available from http://www.nrcresearchpress.com/doi/abs/10.1139/m88-093
- Gallegos M-T, Michán C, Ramos JL. The XylS/AraC family of regulators. Nucleic Acids Res. 1993;21:807–810.
- Chang A, Singh S, Phillips G, et al. Glycosyltransferase structural biology and its role in the design of catalysts for glycosylation. Curr Opin Biotechnol. 2012;22:800–808.
- Stauffer LT, Fogarty SJ, Stauffer GV. Characterization of the Escherichia coli gcv operon. Gene [Internet]. 1994;142;17–22. Available from http://www.sciencedirect.com/science/article/pii/0378111994903492
- Kume A, Koyata H, Sakakibara T, et al. The glycine cleavage system. Molecular cloning of the chicken and human glycine decarboxylase cDNAs and some characteristics involved in the deduced protein structures. J Biol Chem. [Internet]. 1991;266;3323—3329. Available from http://europepmc.org/abstract/MED/1993704
- Okamura-Ikeda K, Ohmura Y, Fujiwara K, et al. Cloning and nucleotide sequence of the gcv operon encoding the Escherichia coli glycine-cleavage system. Eur J Biochem. [Internet]. 1993;216:539—548.
- Rivas-Marín E, Canosa I, Devos DP. Evolutionary cell biology of division mode in the bacterial Planctomycetes-verrucomicrobia-Chlamydiae superphylum. Front Microbiol. 2016;7:1–11.
- Archibald JM. The eocyte hypothesis and the origin of eukaryotic cells. Proc Natl Acad Sci. [Internet]. 2008;105:20049–20050.
- Fox A, Rogers JC, Gilbart J, et al. Muramic acid is not detectable in Chlamydia psittaci or Chlamydia trachomatis by gas chromatography-mass spectrometry. Infect Immun. 1990;58:835–837.
- Makarova KS, Yutin N, Bell SD, et al. Evolution of diverse cell division and vesicle formation systems in archaea. Nat Rev Microbiol. [Internet]. 2010;8:731.
- Mcinerney JO, Martin WF, Koonin EV, et al. Planctomycetes and eukaryotes: A case of analogy not homology. BioEssays. 2011;33:810–817.
- Greub G, Raoult D. Crescent bodies of parachlamydia acanthamoeba and its life cycle within acanthamoeba polyphaga: an electron micrograph study. Appl Environ Microbiol. 2002;68:3076–3084.
- Sittig M, Schlesner H. Chemotaxonomic investigation of various prosthecate and/or budding bacteria. Syst Appl Microbiol [Internet]. 1993;16;92–103. Available from http://www.sciencedirect.com/science/article/pii/S0723202011802535
- Abdelrahman Y, Ouellette SP, Belland RJ, et al. Polarized cell division of chlamydia trachomatis. PLoS Pathog. 2016;12:1–20.
- Fuerst JA, Sagulenko E. Beyond the bacterium: planctomycetes challenge our concepts of microbial structure and function. Nat Rev Microbiol [Internet]. 2011;9:403.
- Mapelli M, Musacchio A. MAD contortions: conformational dimerization boosts spindle checkpoint signaling. Curr Opin Struct Biol [Internet]. 2007;17;716–725. Available from http://www.sciencedirect.com/science/article/pii/S0959440X07001182
- Zhang L-Y, Zhu Z, Yang J. Structural and functional diversification of HORMA domain-containing proteins. J Syst Evol [Internet]. 2015;53:321–329.
- Brinkman FSL, Blanchard JL, Cherkasov A, et al. Evidence that plant-like genes in Chlamydia species reflect an ancestral relationship between Chlamydiaceae, cyanobacteria, and the chloroplast. Genome Res. 2002;12:1159–1167.
- Huang J, Gogarten JP. Did an ancient chlamydial endosymbiosis facilitate the establishment of primary plastids? Genome Biol. 2007;8:1–13.
- McCoy AJ, Adams NE, Hudson AO, et al. L,L-diaminopimelate aminotransferase, a trans-kingdom enzyme shared by Chlamydia and plants for synthesis of diaminopimelate/lysine. Pnas [Internet]. 2006;103;17909–17914. Available from http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=1693846&tool=pmcentrez&rendertype=abstract
- Horn M, Collingro A, Schmitz-Esser S, et al. Illuminating the Evolutionary History of Chlamydiae. [Internet]. Science 2004; 304(5671): 728–730. Available from: http://science.sciencemag.org/content/304/5671/728
- Koonin EV, Yutin N. The dispersed archael eukaryome and the complex archael ancestor of eukaryotes. Cold Spring Harb Perspect Biol. [Internet]. 2014;6:a016188.
- Guy L, Saw JH, Ettema TJG. The archaeal legacy of eukaryotes: A phylogenomic perspective. Cold Spring Harb Perspect Biol. 2014;6:1–16.
- Martin WF, Garg S, Zimorski V. Endosymbiotic theories for eukaryote origin. Philos Trans R Soc B Biol Sci. 2015;370:2014330.

