ABSTRACT
Objective
Cytotoxic T-lymphocyte antigen 4 (CTLA-4) polymorphisms at positions of +49 and CT60 in donors have been reported to influence clinical outcome following allogeneic hematopoietic stem cell transplantation (allo-HSCT), such as overall survival (OS), disease free survival (DFS), relapse and the risk of graft versus host disease (GVHD). However, the results still remain controversial. Thus, we conducted the first meta-analysis to get a more accurate estimation of the relationship between CTLA-4 genotype and the above end points.
Methods
PubMed, Embase, Web of science and Cochrane Library were searched to select eligible studies, data were extracted and then combined ORs/HRs together with the corresponding 95% confidence intervals (CIs) were calculated. Both the dominant and recessive models were employed to evaluate the associations between genetic variation in donor CTLA-4 and outcome after allo-HSCT.
Results
A total of 15 studies were included the pooled results indicated that +49 GG homozygote in donors was significantly associated with increased risk of chronic GVHD (OR=1.701, 95% CI, 1.124-2.573, P=0.012, I2=34.7%). With regard to CT60 polymorphism, donors with G allele correlated with worse OS (HR = 1.422, 95% CI, 1.080-1.872, P=0.012, I2=0%) and lower susceptibility to severe acute GVHD (HR=0.619, 95% CI, 0.426-0.899, P=0.012, I2=0%). There was no significant association between CTLA-4 polymorphism and DFS or the incidence of relapse.
Conclusions
The present meta-analysis suggests that donors with CT60 G allele might be associated with worse OS but reduced severe aGVHD occurrence, while patients transplanted from donors with GG genotype at position of +49 are more likely to suffer from cGVHD.
Introduction
Allogeneic hematopoietic stem cell transplantation (allo-HSCT) is a curative therapy for several hematological malignancies and nonmalignant diseases, because of its substantial morbidity and mortality, posttransplant survival, relapse and the occurrence of GVHD are issues concerned by both clinicians and researchers. Although varieties of studies [Citation1–4] have reported the potential relationship of single nucleotide polymorphisms (SNPs) in genes encoding minor histocompatibility antigens, cytokine, chemokines and even drug-metabolizing enzymes with transplant outcomes, there are currently no well-established predicted biomarkers of clinical results after allo-HSCT. Therefore, investigations aiming to explore the pretransplantation predictors of genetic variants to assist in the risk assessment of adverse outcomes is critical, which can provide potential targets for novel prevention and treatment strategies.
The success of HSCT relies on sustained engraftment, immune reconstitution, effective graft versus leukemia and control of GVHD [Citation5]. The whole process is regulated by an intricate network of molecular signals, especially the immunomodulatory pathway, polymorphism in these involved genes may influence transplant outcomes. Located on chromosome 2q33, CTLA-4 is a negative regulator of T cell activation [Citation6]. The adenine to guanine nucleotide transitions at positions +49 and CT60 were the most widely identified. There have been meta-analyses illustrating the correlation of certain CTLA-4 genotypes with solid organ transplantation, autoimmune diseases such as systemic lupus erythematosus, Graves’ disease, rheumatoid arthritis, and with some cancers [Citation7–11]. To date, several studies have evaluated the effects of CTLA-4 polymorphisms in donors on clinical outcomes following allo-HSCT, but results remain inconclusive, along with the relative small size of individual studies, the real impact of donor CTLA4 alleles on outcomes is unknown. Therefore, it’s necessary to combine all the available studies to clarify the effects of donors’ CTLA-4 genotypes on posttransplant outcome indicators. In this article, we conducted the first meta-analysis to evaluate the association between CTLA-4 variants in donors and clinical outcomes after HSCT.
Methods
Search strategy
This meta-analysis followed the Preferred Reporting Items for Systematic Reviews and Meta-analyses (PRISMA) criteria [Citation12]. The database of PubMed, EMBASE, Web of Science and Cochrane Library were searched to obtain relevant published articles up until October 2020. Search terms were a combination of the following keywords: (“cytotoxic T-lymphocyte antigen-4” or “CTLA-4”) and (“genetic polymorphism” or “polymorphism” or “SNP” or “genotype” or “allele variant” and (“hematopoietic stem cell transplantation” or “bone marrow transplantation” or “HSCT”). Moreover, manual screen of the reference lists from included papers were conducted to identify additional eligible studies. Limitation on language or publication date did not exist.
Study selection
The studies included must fulfill the following inclusion criteria: (1) Clinical cohort or case–control study; (2) Investigation should evaluate the relationship between donor CTLA-4 + 49A/G, CT60 polymorphism and outcome in recipients after allo-HSCT; (3) Odds ratio(OR)/hazard ratio(HR) with 95% confidence intervals(CIs) were provided or sufficient information to calculate corresponding results; (4) If serial studies were involved in the same population, we selected the most recent one. Studies reported in only abstracts, reviews and conference proceedings were excluded. Two independent reviewers checked the articles and reached final consensus.
Qualitative assessment
The Newcastle-Ottawa Scale (NOS) was used to assess the methodological quality, which scored studies in three domains: selection, comparability, and exposure [Citation13]. Only trials awarded at least five stars among 9 quality score items were included. Two authors performed the quality evaluation independently and resolved disagreements by discussion.
Data extraction
The following information was extracted independently by two investigators from each eligible study: surname of the first author, year of publication, country, patients’ age, sample size, transplant type, underlying diseases, conditioning intensity, GVHD prophylaxis, genotyping method, conformity with Hardy-Weinberg equilibrium (HWE) and NOS scores. Dissensions were resolved by consultation with a third reviewer.
Statistical analysis
ORs and the corresponding 95% CIs in donors were used to assess the strength of the association between CTLA-4 polymorphism and GVHD risk, while the impact on overall survival (OS), disease-free survival (DFS) and relapse were estimated from with HRs with 95% CIs. More than two studies about the same item were combined in this meta-analysis. Dominant model and recessive model were applied, because most of the trials described results in these two models. The heterogeneity of the included studies was assessed by the Cochrane’s Q statistic and quantified with I2 measures [Citation14]. We carried out fixed-effects to figure out the pooled effect estimates when the I2 is less than 50%, otherwise random-effect model was employed. An estimate of potential publication bias was evaluated by Begg’s funnel plot and Egger’s test in all the available comparisons (of ≥3 studies), P < 0.05 was considered statistically significant. Given the small number of included studies, sensitivity analyses were not implemented. All statistical analyses were conducted with Stata 14.0 software. A two-sided P-value of < 0.05 was considered statistically significant.
Results
Study characteristics
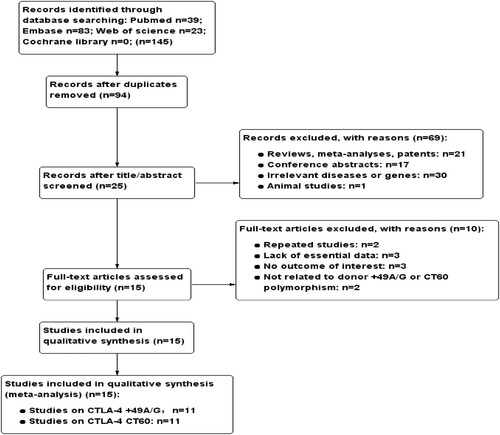
A total of 145 relevant records were obtained from PubMed, Embase, Web of Science and Cochrane Library, of which 51 publications were discarded on account of duplication. After screening over titles and abstracts, 69 records of unrelated articles, conference abstracts or reviews were removed. The remaining 25 articles were assessed for eligibility using the specified inclusion criteria by examining the full text and among them 10 records were excluded for the identical population or lacking of essential data. Eventually, we included 15 qualified studies encompassing 3310 donors in our meta-analysis. The flowchart of literature search and selection was presented in in standard PRISMA flow diagram style. The baseline characteristics of the literatures enrolled were summarized in , eleven studies reported the association between donor CTLA-4 + 49A/G genotypes and different clinical outcome, including GVHD risk, OS or relapse. Similarly, a total of eleven records investigated the influence of CT60 polymorphism in donors have on above-mentioned results. The whole eligible studies were assessed by NOS scale. The quality scores ranged from 6 to 8, indicating that the methodological quality was acceptable.
Table 1. Characteristics of studies included in the meta-analysis.
Results of meta-analysis
The CTLA-4 + 49a/G polymorphism in donors
Overall, eight and seven studies mentioned the effects of genotypes at position +49 on transplantation outcomes in the form of dominant (GG + GA versus AA) and recessive (GG versus GA + AA) model, respectively. In this meta-analysis, we detected that patients who received a graft from donors with GG genotype had an increased risk of developing cGVHD, the aggregated OR was greater than 1 (P=0.012; OR=1.701, 95% CI, 1.124-2.573; ) under fixed effect model. And there was no obvious heterogeneity in this analysis with I2 < 50%. However, as shown in , we did not find any significant association of donor +49A/G genetic variants with aGVHD, OS or relapse. Further subgroup analyses according to ethnicity or donor type were not performed because of the limited number of eligible records.
Figure 2. Forest plots of CTLA-4 + 49A/G polymorphism with cGVHD risk in recessive model(GG vs GA + AA).
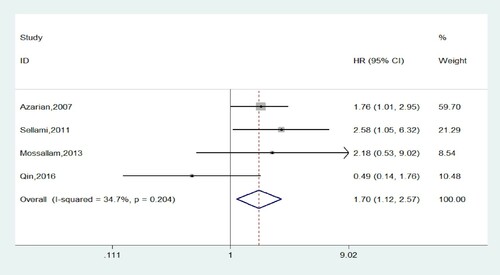
Table 2. Meta-analyses results of the association between CTLA-4 polymorphisms and transplant outcome.
The CTLA-4 CT60 polymorphism in donors
In a dominant model analysis, seven studies were involved assessing the correlation of CTLA-4 CT60 polymorphism with posttransplant outcome variables including OS, DFS, relapse, cGVHD and severity of aGVHD. In this meta, we revealed that donor CTLA-4 CT60 AG + GG showed distinct association with lower OS and reduced incidence of III-IV aGVHD compared to AA genotype (P=0.026; HR=1.440, 95%CI, 1.045-1.984; (a) and P=0.012; HR=0.619, 95%CI, 0.426-0.899; (b), respectively). The I2 values of both of them were 0% and no heterogeneity existed. Nevertheless, our results did not observe any statistical evidence of influence the CT60 G allele have on other ending indicators (see details in ).
Figure 3. (a) Forest plots of CTLA-4 CT60 polymorphism and overall survival in dominant model(GG + GA vs AA). (b) Forest plots of CTLA-4 CT60 polymorphism and the occurrence of III-IV aGVHD in dominant model (GG + GA vs AA). (c) Forest plots of CTLA-4 CT60 polymorphism and overall survival in recessive model (GG vs GA + AA).
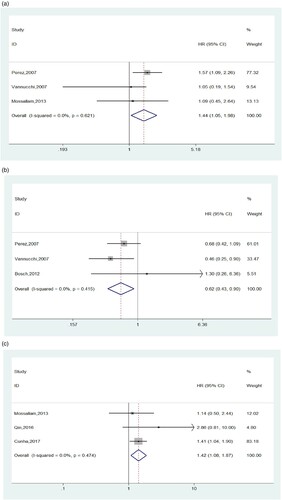
A total of three studies presented the association of CT60 polymorphism in the CTLA-4 gene with different outcome events in a recessive model analysis. As listed in , except for OS, we identified an absence of link between CT60 genetic variations and the susceptibility to cGVHD, relapse or DFS rate post allo-HSCT. With respect to OS, the combined HR suggested that recipients transplanted from donors with CT60 GG homozygote tended to have a shorter survival than those who received grafts from donors possessing A allele (P=0.012, HR=1.422, 95%CI, 1.080-1.872, (c)), which is consistent with the results under a dominant model. Meanwhile, the I2 was 0%, indicating there was no heterogeneity. Due to the limited included studies, we failed to carry out subgroup analyzed on the basis of ethnicity or transplant pattern.
Publication bias
Begg’s funnel plot and Egger’s test were performed to assess publication bias in all the available comparisons (of ≥3 studies). Funnel plot asymmetry was assessed by the method of Egger’s linear regression test. As shown in , the results of Begg’s and Egger’s test suggested that no obvious publication bias existed (P > 0.05).
Discussion
A number of previous studies have widely investigated the relationship of CTLA-4 + 49A/G and CT60 genotype with several end points following allo-HSCT, but some with controversial results, so we performed the present meta-analysis hoping to obtain relatively accurate conclusions. To our knowledge, this is the first meta-analysis focusing on such an association. We summarized the effects of donor CTLA-4 + 49A/G and CT60 genetic variations on multiple outcomes including OS, DFS, relapse and the risk of GVHD after allo-HSCT in both dominant and recessive model though we failed to detect partial outcome indicators in a certain model owing to unavailable detailed data. We concluded that patients receiving stem cells from a donor who was in possession of at least 1 G allele in position CT60 conferred inferior overall survival, but the case was not for +49, which was in line with the reports by Perez et al. [Citation15] in a multivariant analysis, the involvement of CT60 was significant while +49A/G was not identified as an independent risk factor for OS. Cunha et al. [Citation27] also revealed the same conclusion regarding the CTLA-4 CT60 polymorphism, suggesting the GG genotype reduced the OS in patients after CBT. As for +49 A/G, Qin et al. [Citation26] supported the above-mentioned results, whereas the contrary findings were presented by Piccioli et al. [Citation18], who observed a longer OS in patients transplanted from a donor that was homozygote G/G. Several other studies did not observe any significant influence of donor +49 or CT60 genetic polymorphism on OS in different cohorts [Citation16, Citation17, Citation24].
When examined the susceptibility to GVHD, the two CTLA-4 SNPs in donors showed different results. To be more specific, cGVHD were more frequent in recipients accepting stem cells from donors with GG genotype at position of +49, whereas the G allele of CT60 acted as a protective factor against the incidence of severe aGVHD (grade III-IV). We failed to find any statistical significantly impact of the +49 polymorphism on aGVHD and CT60 genotype on cGVHD, respectively, possibly because of the limited number of suitable studies involved and relatively higher heterogeneity, so the results were not very reliable to some extent, more studies with larger sample size were needed to further define the associations in the future. After browsing relevant literatures about issues in this aspect, we found discrepant results as well. As for the polymorphism of +49 at exon 1, the same tendency to cGVHD was observed in studies by Azarian et al. [Citation16] and Sellami et al. [Citation19], conversely, Mossalam et al. [Citation24], Vannucchi et al. [Citation17] and Qin et al. [Citation26] found that +49 G allele in donor was not associated with cGVHD risk. In addition, the data provided by Perez-Garcia et al. [Citation15] in a Spanish cohort indicated that the presence of donor CT60 GG + AG resulted in a reduction of moderate GVHD (II-IV) incidence, which is completely contradicted with that concluded by Xiao et al. [Citation23]. Of course, consistent with our result, a lack of correlation between CT60 G allele and moderate aGVHD risk was presented in other studies [Citation20,Citation21,Citation24]. Two researches [Citation15,Citation20] concerning the role of CT60 GG genotype in III-IV aGVHD development evidenced that there were no significant difference, on the contrary, Vannucchi et al. [Citation17] reported that grade III-IV aGVHD had a tendency to occur in patients whose donors were CT60 AA genetic type, the final result of our current meta supported the positive finding of the latter study.
It is known that CTLA-4 consists of four exons and is translated into two isoforms, a soluble form (sCTLA-4) lacking of exon 3 and a full-length isoform(flCTLA-4), which functions through inducing cell cycle arrest and blocking cytokines secretion [Citation30]. It has been suggested that CT60 is of great value in splicing and producing sCTLA-4 efficiently, and the allele A is responsible for greater level of sCTLA-4 in patients with autoimmune diseases and healthy controls [Citation15,Citation31]. Moreover, the sCTLA-4 could competitively block the B7–flCTLA-4 combination, thereby enhancing T-cell reactivity and graft versus leukemia (GVL) effects by preventing the transduction of inhibitory signals that lead to lymphocyte inactivation [Citation15,Citation31]. Taken together, these previous findings could explain our result stated above, the levels of sCTLA-4 in donors with CT60 G allele are lower, resulting in a weaker competitive with flCTLA-4 for the B7 ligand, T cells tend to be inactivated, consequently, lower GVL effects are obtained, perhaps this is the reason why CT60 G allele is associated with worse OS but lower susceptibility to severe aGVHD. Traditionally, the A allele has been identified as protective against greater susceptibility to autoimmune diseases [Citation10,Citation11,Citation32], which is in conformity with our result describing that the donor GG homozygote might result in an increased risk of cGVHD for +49. One hypothesized reason underlying this behavior may trace to the characteristics of +49, located on exon1, which encodes the leader peptide of the protein and participates in CTLA-4 trafficking to endoplasmic reticulum. Although the localization where CTLA-4 exerts its inhibitory capacity is unclear, it is conceivable that A to G transition in the leader sequence may attribute to altered rates of endocytosis or surface trafficking, thus generating different effects on T cell responses [Citation32]. Further investigations are needed to clarify the exact mechanism.
Limitations of the current meta-analysis should be acknowledged. Firstly, apparent heterogeneity was observed among some outcome indictors, especially relapse and DFS, the heterogeneity between trials may come from distinct transplantation settings, ethnic background, GVHD prophylaxis and even differences in genetic detection methods; Secondly, few studies evaluated the influence of CTLA-4 genotype on some outcome we focused on and the sample size of incorporated studies were relatively small due to a lack of detailed data, so we failed to complete subgroup analyses according to the sources of heterogeneity, this restricted the strength of our study; In addition, owing to the absence of detailed original data, we could not conduct appropriate analyses under other genetic models, such as allele model, homozygote model or heterozygote model; Finally, We have no chance to carry out sensitivity analysis as a result of small size of studies included again.
In summary, despite the limitations, this meta-analysis still suggests that donors with CT60 G allele might be associated with worse OS but reduced severe aGVHD occurrence, while patients transplanted from donors with GG genotype at position of +49 are more likely to suffer from cGVHD. This study contributes towards improving patient outcomes after HSCT by providing insight and rationale in developing novel, individualized treatment strategies considering genetic determinants. More future studies with good methodology design and larger-scale population are warranted to confirm our findings.
Disclosure statement
No potential conflict of interest was reported by the author(s).
References
- Loeffler J, Ok M, Morton OC, et al. Genetic polymorphisms in the cytokine and chemokine system: their possible importance in allogeneic stem cell transplantation. Curr Top Microbiol Immunol. 2010;341:83–96. Epub 2010/04/17. doi:https://doi.org/10.1007/82_2010_22. PubMed PMID: 20397074.
- Martinez-Laperche C, Buces E, Aguilera-Morillo MC, et al. A novel predictive approach for GVHD after allogeneic SCT based on clinical variables and cytokine gene polymorphisms. Blood Adv. 2018;2(14):1719–1737. Epub 2018/07/22. doi:https://doi.org/10.1182/bloodadvances.2017011502. PubMed PMID: 30030270; PubMed Central PMCID: PMCPMC6058238.
- Hamadeh IS, Zhang Q, Steuerwald N, et al. Effect of CYP3A4, CYP3A5, and ABCB1 polymorphisms on Intravenous tacrolimus exposure and adverse events in Adult allogeneic stem cell transplant patients. Biology of Blood and Marrow Transplantation: Journal of the American Society for Blood and Marrow Transplantation. 2019;25(4):656–663. Epub 2019/01/01. doi:https://doi.org/10.1016/j.bbmt.2018.12.766. PubMed PMID: 30597277.
- Larsen ME, Kornblit B, Larsen MV, et al. Degree of predicted minor histocompatibility antigen mismatch correlates with poorer clinical outcomes in nonmyeloablative allogeneic hematopoietic cell transplantation. Biology of Blood and Marrow Transplantation: Journal of the American Society for Blood and Marrow Transplantation. 2010;16(10):1370–1381. Epub 2010/04/01. doi:https://doi.org/10.1016/j.bbmt.2010.03.022. PubMed PMID: 20353833.
- Hansen JA, Petersdorf EW, Lin MT, et al. Genetics of allogeneic hematopoietic cell transplantation. role of HLA matching, functional variation in immune response genes. Immunol Res. 2008;41(1):56–78. Epub 2007/11/09. doi:https://doi.org/10.1007/s12026-007-0043-x. PubMed PMID: 17989941.
- Teft WA, Kirchhof MG, Madrenas J. A molecular perspective of CTLA-4 function. Annu Rev Immunol. 2006;24:65–97. Epub 2006/03/23. doi:https://doi.org/10.1146/annurev.immunol.24.021605.090535. PubMed PMID: 16551244.
- Hu P, Liu Q, Deng G, et al. The prognostic value of cytotoxic T-lymphocyte antigen 4 in cancers: a systematic review and meta-analysis. Sci Rep. 2017;7:1-10, Epub 2017/02/18. doi:https://doi.org/10.1038/srep42913. PubMed PMID: 28211499; PubMed Central PMCID: PMCPMC5314410.
- Zhai JX, Zou LW, Zhang ZX, et al. CTLA-4 polymorphisms and systemic lupus erythematosus (SLE): a meta-analysis. Mol Biol Rep. 2013;40(9):5213–5223. Epub 2013/08/08. doi:https://doi.org/10.1007/s11033-012-2125-7. PubMed PMID: 23922195.
- Zhu CL, Huang Q, Liu CH, et al. Polymorphisms in the cytotoxic T-lymphocyte antigen 4 gene and acute rejection risk in transplant recipients. Mol Biol Rep. 2012;39(9):8701–8708. Epub 2012/06/20. doi:https://doi.org/10.1007/s11033-012-1727-4. PubMed PMID: 22711308.
- Si X, Zhang X, Tang W, et al. Association between the CTLA-4 + 49A/G polymorphism and Graves’ disease: A meta-analysis. Exp Ther Med. 2012;4(3):538–544. Epub 2012/11/28. doi:https://doi.org/10.3892/etm.2012.618. PubMed PMID: 23181132; PubMed Central PMCID: PMCPMC3503798.
- Li G, Shi F, Liu J, et al. The effect of CTLA-4 A49G polymorphism on rheumatoid arthritis risk: a meta-analysis. Diagn Pathol. 2014;9:157, Epub 2014/08/17. doi:https://doi.org/10.1186/s13000-014-0157-0. PubMed PMID: 25128482; PubMed Central PMCID: PMCPMC4160544.
- Moher D, Liberati A, Tetzlaff J, et al. Preferred reporting items for systematic reviews and meta-analyses: the PRISMA statement. PLoS Med. 2009;6(7):e1000097.
- Wells GA, Shea BJ, O’Connell D, et al. The Newcastle–Ottawa scale (NOS) for assessing the quality of non-randomized studies in meta-analysis. Appl Eng Agric. 2014;18:727–734.
- Cochran WG. The combination ofestimates from different experiments. Biometrics. 1954;10:101–129.
- Perez-Garcia A, De la Camara R, Roman-Gomez J, et al. CTLA-4 polymorphisms and clinical outcome after allogeneic stem cell transplantation from HLA-identical sibling donors. Blood. 2007;110(1):461–467. Epub 2007/03/27. doi:https://doi.org/10.1182/blood-2007-01-069781. PubMed PMID: 17384200.
- Azarian M, Busson M, Lepage V, et al. Donor CTLA-4 + 49 A/G*GG genotype is associated with chronic GVHD after HLA-identical haematopoietic stem-cell transplantations. Blood. 2007;110(13):4623–4624. Epub 2007/12/07. doi:https://doi.org/10.1182/blood-2007-08-106385. PubMed PMID: 18056853.
- Vannucchi AM, Guidi S, Guglielmelli P, et al. Significance of CTLA-4 and CD14 genetic polymorphisms in clinical outcome after allogeneic stem cell transplantation. Bone Marrow Transplant. 2007;40(10):1001–1002. Epub 2007/09/12. doi:https://doi.org/10.1038/sj.bmt.1705850. PubMed PMID: 17846597.
- Piccioli P, Balbi G, Serra M, et al. CTLA-4 + 49A > G polymorphism of recipients of HLA-matched sibling allogeneic stem cell transplantation is associated with survival and relapse incidence. Ann Hematol. 2010;89(6):613–618. Epub 2009/12/19. doi:https://doi.org/10.1007/s00277-009-0885-5. PubMed PMID: 20020126.
- Sellami MH, Bani M, Torjemane L, et al. Effect of donor CTLA-4 alleles and haplotypes on graft-versus-host disease occurrence in Tunisian patients receiving a human leukocyte antigen-identical sibling hematopoietic stem cell transplant. Hum Immunol. 2011;72(2):139–143. Epub 2010/11/18. doi:https://doi.org/10.1016/j.humimm.2010.11.008. PubMed PMID: 21081144.
- Bosch-Vizcaya A, Perez-Garcia A, Brunet S, et al. Donor CTLA-4 genotype Influences clinical outcome after T cell-Depleted allogeneic hematopoietic stem cell transplantation from HLA-identical sibling donors. Biol Blood Marrow Transplant. 2012;18(1):100–105. doi:https://doi.org/10.1016/j.bbmt.2011.05.021. PubMed PMID: WOS:000303140200012.
- Harkensee C, Oka A, Onizuka M, et al. Single nucleotide polymorphisms and outcome risk in unrelated mismatched hematopoietic stem cell transplantation: An exploration study. Blood. 2012;119(26):6365–6372. doi:https://doi.org/10.1182/blood-2012-01-406785.
- Orru S, Orru N, Manolakos E, et al. Recipient CTLA-4*CT60-AA genotype is a prognostic factor for acute graft-versus-host disease in hematopoietic stem cell transplantation for thalassemia. Hum Immunol. 2012;73(3):282–286. Epub 2012/01/17. doi:https://doi.org/10.1016/j.humimm.2011.12.014. PubMed PMID: 22245568; PubMed Central PMCID: PMCPMC3314940.
- Xiao H, Luo Y, Lai X, et al. Genetic variations in T-cell activation and effector pathways modulate alloimmune responses after allogeneic hematopoietic stem cell transplantation in patients with hematologic malignancies. Haematologica. 2012;97(12):1804–1812. Epub 2012/06/27. doi:https://doi.org/10.3324/haematol.2012.066159. PubMed PMID: 22733023; PubMed Central PMCID: PMCPMC3590086.
- Mossallam GI, Samra MA. CTLA-4 polymorphism and clinical outcome post allogeneic hematopoietic stem cell transplantation. Hum Immunol. 2013;74(12):1643–1648. Epub 2013/08/27. doi:https://doi.org/10.1016/j.humimm.2013.08.002. PubMed PMID: 23973330.
- Karabon L, Markiewicz M, Partyka A, et al. A CT60G > A polymorphism in the CTLA-4 gene of the recipient may confer susceptibility to acute graft versus host disease after allogeneic hematopoietic stem cell transplantation. Immunogenetics. 2015;67(5-6):295–304. Epub 2015/05/06. doi:https://doi.org/10.1007/s00251-015-0840-7. PubMed PMID: 25940108; PubMed Central PMCID: PMCPMC4427628.
- Qin XY, Wang Y, Li GX, et al. CTLA-4 polymorphisms and haplotype correlate with survival in ALL after allogeneic stem cell transplantation from related HLA-haplotype-mismatched donor. J Transl Med. 2016;14:100, Epub 2016/04/28. doi:https://doi.org/10.1186/s12967-016-0864-2. PubMed PMID: 27118383; PubMed Central PMCID: PMCPMC4847362.
- Cunha R, Zago MA, Querol S, et al. Impact of CTLA4 genotype and other immune response gene polymorphisms on outcomes after single umbilical cord blood transplantation. Blood. 2017;129(4):525–532. Epub 2016/11/05. doi:https://doi.org/10.1182/blood-2016-06-722249. PubMed PMID: 27811020.
- Hammrich J, Wittig S, Ernst T, et al. CTLA-4 polymorphism rs231775: influence on relapse and survival after allogeneic hematopoietic stem cell transplantation in childhood. Eur J Haematol. 2019;102(3):251–255. Epub 2018/11/23. doi:https://doi.org/10.1111/ejh.13200. PubMed PMID: 30465728.
- Romaniuk DS, Khmelevskaya AA, Drokov MY, et al. Effect of CTLA4 gene polymorphism on relapse probability among patients with acute leukemias after allogenic hematopoietic stem cells transplantation. Oncogematologiya. 2019;14(1):76–82. doi:https://doi.org/10.17650/1818-8346-2019-14-1-76-82.
- Gough SC, Walker LS, Sansom DM. CTLA4 gene polymorphism and autoimmunity. Immunol Rev. 2005;204:102–115. Epub 2005/03/26. doi:https://doi.org/10.1111/j.0105-2896.2005.00249.x. PubMed PMID: 15790353.
- Ueda H, Howson JM, Esposito L, et al. Association of the T-cell regulatory gene CTLA4 with susceptibility to autoimmune disease. Nature. 2003;423(6939):506–511. Epub 2003/05/02. doi:https://doi.org/10.1038/nature01621. PubMed PMID: 12724780.
- Oaks MK, Hallett KM. Cutting Edge: A soluble form of CTLA-4 in patients with autoimmune Thyroid disease. The Journal of Immunology. 2000;164(10):5015, doi:https://doi.org/10.4049/jimmunol.164.10.5015.