Abstract
We report the first complete mitogenome (Mt) sequence of Aedes japonicus japonicus (Diptera: Culicidae). The sequence was extracted from one adult from the Big Island of Hawai’i Island. The length of the Ae. japonicus japonicus Mt was 16,528bp with 78.1% AT content. Its sequence is most similar to the Mt sequence of Aedes koreicus with 90.81% sequence identity. This is the first full Mt sequence available for this species and provides important genetic resource for studying population genetics and dynamics of this important invasive mosquito species.
Introduction
Aedes (Hulecoeteomyia) japonicus (Theobald Citation1901), commonly known as the Asian bush mosquito (), was first documented in Japan in 1901 as Culex japonicus (Theobald Citation1901). Aedes j. japonicus has been described as a subspecies of Ae. japonicus and can be challenging to morphologically distinguish from other subspecies within Ae. japonicus (Tanaka et al. Citation1979). On the other hand, four subspecies are genetically quite distinct from each other (Cameron et al. Citation2010). Based on the re-examination of morphological characteristics, evidence of molecular divergence, and apparent allopatric distributions of the subspecies of Ae. japonicus, Wilkerson et al. suggested elevating Ae. j. japonicus and other subspecies to species rank recently (Wilkerson et al. Citation2022).
Figure 1. Aedes japonicus reference image. Courtesy of David Pecor at the Walter Reed Biosystematics Unit (WRBU) (Matsunaga et al. Citation2019). (A) Habitus in lateral view. (B) Thorax in dorsal view. The scutum is characterized by golden stripes with distinctive lyre-shaped strips, two sub-median and a median strip, which is one of the features used for species identification.

Aedes j. japonicus is more common than other subspecies, and its native distribution area is Palearctic region while other subspecies is found in Oriental region of Japan (Tanaka et al. Citation1979; Cameron et al. Citation2010). Interaction of Ae. j. japonicus with international commerce has been implicated in its invasion and spread in North America and Europe (Cameron et al. Citation2010). Its slow expansion in the United States of America could be explained by its limited tolerance for higher ambient temperatures in coastal habitats (Kaufman and Fonseca Citation2014), although this species has been established in subtropical Florida (Riles et al. Citation2017) and the tropical Hawaiian Islands (Larish and Savage Citation2005; Larish et al. Citation2010; Matsunaga et al. Citation2019).
Ecological impacts of its expansion are recorded in its interspecific associations with other container-inhabiting mosquitoes including disease vector (Armistead et al. Citation2008; Andreadis and Wolfe Citation2010), and therefore may indirectly affect the local disease dynamics. In North America, Aedes j. japonicus has been found naturally infected with several pathogens (Harris et al. Citation2015; Silaghi et al. Citation2017; Yang et al. Citation2018; DeCarlo et al. Citation2020) including Cache Valley virus, Heartworm, La Crosse Virus, and West Nile virus. However, its role in transmission can be considered ambiguous and requires further investigations. In vector competence experiments, Ae. j. japonicus has been determined to be an efficient laboratory vector due to its susceptibility to various arboviruses crossing salivary gland barriers (Turell et al. Citation2013; Abbo et al. Citation2020).
Materials and methods
The adult Aedes japonicus specimen was collected from Hawai’i (19.0853°N, 155.7757°W) using a BG-sentinel mosquito trap (Biogents, Regensburg, Germany) baited with BG-Sweetscent Attractant. DNA extraction and library preparations were conducted as previously described (Chen et al. Citation2021; Kelly et al. Citation2021). Extracted DNA sample (voucher accession number: Ae21OCNV006, contact person: Yoosook Lee, [email protected]) was kept in the Florida Medical Entomology Laboratory at the University of Florida.
The library was sequenced for 150 bp paired-end reads using a NovaSeq 6000 instrument (Illumina, San Diego, CA) at the University of Florida Interdisciplinary Center for Biotechnology Research (UF ICBR) NextGen DNA Sequencing Core. Raw sequencing reads were trimmed using fastp version 0.20.1 (Chen et al. Citation2018). Mt contigs were assembled using NOVOPlasty version 4.2 (Dierckxsens et al. Citation2017). Following the methods used for Ae. busckii and Ae. taeniorhynchus (Cornel et al. Citation2020), automatic annotation of mitogenome (Mt) was conducted with the MITOS website (Bernt et al. Citation2013) under default settings and the invertebrate genetic code for mitochondria. The Jukes-Cantor model was used to build the phylogenetic tree with the maximum likelihood method using MEGA11 software version 11.0.13 (Kearse et al. Citation2012).
Results
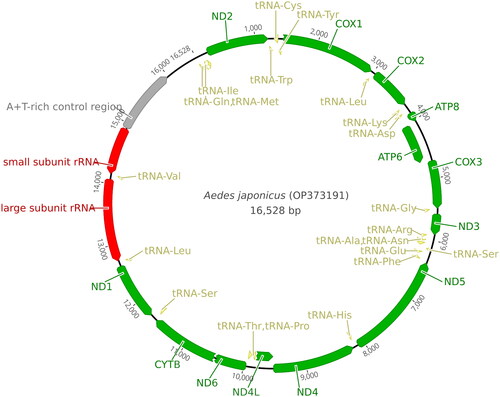
Our NovoPlasty run yielded a single circular DNA contig of Ae. j. japonicus mitochondrion. The average organelle coverage was 3298X. The mitochondrial genome was 0.55% of the extracted mosquito genomic DNA. The length of the Ae. j. japonicus Mt (GenBank accession number: OP373191) was 16,528 bp and the percentage of A + T was 78.1% (). The cytochrome c oxidase I (COI) fragment spanning 1458–2166 bp of Ae. j. japonicus sequence was 99.49% (±0.2 SD) identical to the COI sequences of Ae. j. japonicus deposited in GenBank. Some of automatic annotations did not end with stop codon. In such cases, we revised the end points manually to properly align with stop codons. Annotation information was also deposited to the GenBank with the genome sequence. Genomic map is provided in .
Figure 2. Mitogenome map of Ae. j. japonicus. Green ribbons indicate genes; yellow ribbons indicate tRNA-encoding regions; red regions indicate rRNA-encoding regions; and gray ribbon indicates an AT-rich region. The ribbon orientation depicts the direction of transcription.
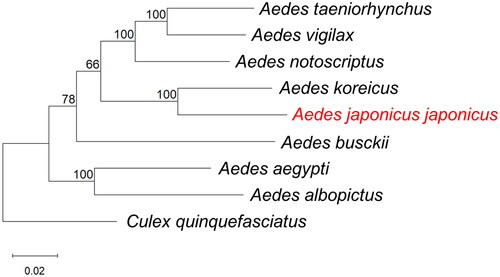
A phylogenetic tree including other disease vector mosquito species is shown in . In the phylogenetic tree, the closest match to the whole Mt sequence of Ae. j. japonicus was Ae. koreicus (GenBank accession number: MT093832) collected from South Korea (Shin and Jung Citation2020) with 90.81% sequence identity.
Figure 3. Phylogenetic tree constructed using maximum-likelihood (ML) based on complete mitogenome sequences of related mosquitoes. The following sequences were used: Aedes taeniorhynchus MN626442 (Cornel et al. Citation2020), Ae. vigilax KP995260 (Hardy et al. Citation2016), Ae. notoscriptus KM676219 (Hardy et al. Citation2016), Ae. koreicus MT093832 (Shin and Jung Citation2020), Ae. japonicus japonicus OP373191 (this study), Ae. busckii MN626443 (Cornel et al. Citation2020), Ae. aegypti MH348176 (Schmidt et al. Citation2018), Ae. albopictus NC_006817 (Ho et al. Citation2021), Culex quinquefasciatus HQ724617 (Atyame et al. Citation2016). Numbers at nodes indicate bootstrap frequencies out of 500 replicates. Culex quinquefasciatus was considered as an outgroup. The scale bar indicates relative nucleotide difference (0.02 = 2% nucleotide difference).
Discussion and conclusions
Here, we sequenced and annotated the first complete Mt sequence of wild caught Ae. j. japonicus from Hawai’i. The phylogenetic analysis of the whole mitochondria sequence showed that Ae. j. japonicus had the closet match to Ae. koreicus with 90.81% sequence identify, comparing with any of other co-occurring and morphologically confusing invasive species such as Ae. albopictus and Ae. aegypti (Wilkerson et al. Citation2022). This close relationship is consistent with one earlier study that revealed the surprising monophyletic grouping among all subspecies of Ae. japonicus and Ae. koreicus by using two mitochondrial markers and a nuclear marker (Cameron et al. Citation2010). Given that these two taxa are invasive sibling species but are difficult to separate morphologically (Tanaka et al. Citation1979), the results of this study may provide mitochondrial genetic markers for molecular species identification. Data presented here could also have beneficial impacts on future genetic and evolutionary studies on Ae. japonicus.
Ethics statement
The study involves collection of mosquito specimen in public places (beach parks) or private properties. The verbal consent from homeowners was acquired prior to conducting collections in private properties. No permit or ethical approval is required for this study.
Author contributions
YL, CMJ, and OSA conceived experiments; CMJ and YL conducted field collection and species identification; SS, ALR-W, and VTN contributed to sequence generation; SS, ALR-W, VTN, TCC, XW, and MTR contributed to data analysis; SS, XW, MTR, and YL contributed to drafting the manuscript; all authors contributed to revision of manuscript.
Acknowledgements
We thank UF ICBR for providing sequencing services. We appreciate the support from Gracelda Simmons from Hawai’i Department of Health toward our project and the assistance from Logan Takasaki, Tyler Gibo, Cory Shiraishi, Guy Tsutsui, and Bruce Mackey from the Hawai’i Department of Health for field collection.
Disclosure statement
The authors report no conflicts of interest. The authors alone are responsible for the content and writing of this article. This publication was developed under Assistance Agreement No. 84020401 awarded by the U.S. Environmental Protection Agency to Akbari. It has not been formally reviewed by EPA. The views expressed in this document are solely those of authors and do not necessarily reflect those of the Agency. EPA does not endorse any products or commercial services mentioned in this publication.
Data availability statement
The data that support the findings of this study are openly available in GenBank of NCBI at https://www.ncbi.nlm.nih.gov/ under the accession number OP373191. The associated BioProject, SRA, and Bio-Sample numbers are PRJNA882781, SRR21658404, and SAMN30950993, respectively.
Additional information
Funding
References
- Abbo SR, Visser TM, Wang H, Göertz GP, Fros JJ, Abma-Henkens MHC, Geertsema C, Vogels CBF, Koopmans MPG, Reusken CBEM, et al. 2020. The invasive Asian bush mosquito Aedes japonicus found in the Netherlands can experimentally transmit Zika virus and Usutu virus. PLoS Negl Trop Dis. 14(4):e0008217.
- Andreadis TG, Wolfe RJ. 2010. Evidence for reduction of native mosquitoes with increased expansion of invasive Ochlerotatus japonicus japonicus (Diptera: Culicidae) in the northeastern United States. J Med Entomol. 47(1):43–52.
- Armistead JS, Arias JR, Nishimura N, Lounibos LP. 2008. Interspecific larval competition between Aedes albopictus and Aedes japonicus (Diptera: Culicidae) in northern Virginia. J Med Entomol. 45(4):629–637.
- Atyame C, Weill M, Duron O. 2016. Culex quinquefasciatus from USA mitochondrion, complete genome [Internet] [accessed 2022 Dec 13]. http://www.ncbi.nlm.nih.gov/nuccore/HQ724617.1.
- Bernt M, Donath A, Jühling F, Externbrink F, Florentz C, Fritzsch G, Pütz J, Middendorf M, Stadler PF. 2013. MITOS: improved de novo metazoan mitochondrial genome annotation. Mol Phylogenet Evol. 69(2):313–319.
- Cameron EC, Wilkerson RC, Mogi M, Miyagi I, Toma T, Kim H-C, Fonseca DM. 2010. Molecular phylogenetics of Aedes japonicus, a disease vector that recently invaded Western Europe, North America, and the Hawaiian islands. J Med Entomol. 47(4):527–535.
- Chen S, Zhou Y, Chen Y, Gu J. 2018. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics. 34(17):i884–i890.
- Chen T, Vorsino AE, Kosinski KJ, Romero-Weaver AL, Buckner EA, Chiu JC, Lee Y. 2021. A magnetic-bead-based mosquito DNA extraction protocol for next-generation sequencing. J Vis Exp. 170:e62354.
- Cornel AJ, Bargielowski IE, Collier TC, Weakley AM, Blosser EM, Lanzaro GC, Hulshof K, Braks MAH, Lee Y. 2020. Complete mitogenome sequences of Aedes (Howardina) busckii and Aedes (Ochlerotatus) taeniorhynchus from the Caribbean Island of Saba. Mitochondrial DNA B Resour. 5(2):1163–1164.
- DeCarlo CH, Campbell SR, Bigler LL, Mohammed HO. 2020. Aedes japonicus and West Nile Virus in New York. J Am Mosq Control Assoc. 36(4):261–263.
- Dierckxsens N, Mardulyn P, Smits G. 2017. NOVOPlasty: de novo assembly of organelle genomes from whole genome data. Nucleic Acids Res. 45(4):e18.
- Hardy CM, Court LN, Morgan MJ, Webb CE. 2016. The complete mitochondrial DNA genomes for two lineages of Aedes notoscriptus (Diptera: Culicidae). Mitochondrial DNA A DNA Mapp Seq Anal. 27(3):2024–2025.
- Hardy CM, Court LN, Morgan MJ. 2016. The complete mitochondrial DNA genome of Aedes vigilax (Diptera: Culicidae). Mitochondrial DNA A DNA Mapp Seq Anal. 27(4):2552–2553.
- Harris MC, Dotseth EJ, Jackson BT, Zink SD, Marek PE, Kramer LD, Paulson SL, Hawley DM. 2015. La Crosse Virus in Aedes japonicus japonicus mosquitoes in the Appalachian Region, United States. Emerg Infect Dis. 21(4):646–649.
- Harwood JF, Fiorenzano JM, Gerardo E, Black T, LaPointe DA, Hasty J. 2018. Seasonal surveillance confirms the range expansion of Aedes japonicus japonicus (Theobald) (Diptera: Culicidae) to the Hawaiian Islands of Oahu and Kauai. J Asia-Pacific Entomol. 21(4):1366–1372.
- Ho CM, Chang HP, Liu YM. 2021. Aedes albopictus mitochondrion, complete genome [Internet] [accessed 2022 Dec 13]. http://www.ncbi.nlm.nih.gov/nuccore/NC_006817.1.
- Kaufman MG, Fonseca DM. 2014. Invasion biology of Aedes japonicus japonicus (Diptera: Culicidae). Annu Rev Entomol. 59:31–49.
- Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, et al. 2012. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics. 28(12):1647–1649.
- Kelly ET, Mack LK, Campos M, Grippin C, Chen TY, Romero-Weaver AL, Kosinski KJ, Brisco KK, Collier TC, Buckner EA, et al. 2021. Evidence of local extinction and reintroduction of Aedes aegypti in Exeter, California. Front Trop Dis. 2:8.
- Larish LB, Savage HM. 2005. Introduction and establishment of Aedes (Finlaya) japonicus japonicus (Theobald) on the island of Hawaii: implications for arbovirus transmission. J Am Mosq Control Assoc. 21(3):318–321.2.0.CO;2]
- Larish LB, Yang P, Asuncion BA. 2010. Distribution and Abundance of Aedes (Finlaya) japonicus japonicus (Theobald) in five districts on the Island of Hawaii. Proc Hawaiian Entomol Soc. 42:9–14.
- Matsunaga JN, Howarth FG, Kumashiro BR. 2019. New state records and additions to the alien terrestrial arthropod fauna in the Hawaiian Islands. Proc Hawaiian Entomol Soc. 51(1):1–71.
- Riles MT, Smith JP, Burkett-Cadena N, Connelly CR, Morse GW, Byrd BD. 2017. First record of Aedes japonicus in Florida. J Am Mosq Control Assoc. 33(4):340–344.
- Schmidt H, Hanemaaijer MJ, Cornel AJ, Lanzaro GC, Braack L, Lee Y. 2018. Complete mitogenome sequence of Aedes (Stegomyia) aegypti derived from field isolates from California and South Africa. Mitochondrial DNA B Resour. 3(2):994–995.
- Shin J, Jung J. 2020. First record of the complete mitochondrial genome of Aedes koreicus (Edwards, 1917) (Diptera: Culicidae) in South Korea. Mitochondrial DNA B Resour. 5(2):2000–2001.
- Silaghi C, Beck R, Capelli G, Montarsi F, Mathis A. 2017. Development of Dirofilaria immitis and Dirofilaria repens in Aedes japonicus and Aedes geniculatus. Parasit Vectors. 10(1):94.
- Tanaka K, Mizusawa K, Saugstad ES. 1979. A revision of the adult and larval mosquitoes of Japan (including the Ryukyu Archipelago and the Ogasawara Islands) and Korea (Diptera: Culicidae). Contrib Am Entomol Inst. 16:1–987.
- Theobald FV. 1901. A monograph of the Culicidae, or mosquitoes. London: British museum (Nat. hist.) Department of Zoology; p. 256.
- Turell MJ, Byrd BD, Harrison BA. 2013. Potential for populations of Aedes j. japonicus to transmit Rift Valley fever virus in the USA. J Am Mosq Control Assoc. 29(2):133–137.
- Wilkerson RC, Somboon P, Harbach RE. 2022. Reconsideration of the status of subspecies in the Japonicus Group of the subgenus Hulecoeteomyia Theobald of Aedes Meigen (Diptera: Culicidae). Zootaxa. 5162(2):198–200.
- Yang F, Chan K, Marek PE, Armstrong PM, Liu P, Bova JE, Bernick JN, McMillan BE, Weidlich BG, Paulson SL, et al. 2018. Cache valley virus in Aedes japonicus japonicus mosquitoes, Appalachian Region, United States. Emerg Infect Dis. 24(3):553–557.
