Abstract
With its nearly 200 species, the Mammillaria genus is the most species-rich within the Cactaceae family, yet surprisingly, few of its chloroplast genomes have been studied. We focused on the species Mammillaria elongata DC. 1828, a petite cactus native to Mexico and favored by horticulturists, yet whose phylogenetic relationships remain uncertain due to a lack of genomic data. We extracted the DNA from a sample obtained in China, sequenced it using the NovaSeq 6000 platform, and assembled the chloroplast genome using GetOrganelle software. Our assembly resulted in a chloroplast genome of 110,981 base pairs with an overall GC content of 36.28%, which included 100 genes (95 unique). Notably, several protein-coding genes were absent. Phylogenetic analysis using 59 shared genes across nine Mammillaria species and one Obregonia species revealed that M. elongata and M. gracilis are closely related, suggesting a recent common ancestor and possible shared evolutionary pressures or ecological niches. This study provides crucial genomic data for M. elongata and hints at intriguing phylogenetic relationships within the Mammillaria genus.
Introduction
Situated within the Caryophyllales order, the cactus family (Cactaceae) graces the horticultural world with its captivating ornamental plants. This extensive family encompasses about 174 genera and nearly 2000 species (Novoa et al. Citation2014; Abouseadaa et al. Citation2020). Historically, the taxonomic classification of Mammillaria (Cactaceae) has been challenged by extensive morphological variation and species sympatry. Through the deployment of chloroplast markers, Cristian R. Cervantes et al. have advanced our understanding by constructing a detailed phylogenetic tree, thereby refining the infrageneric classifications, encompassing subgeneric, sectional, and series categorizations (Cervantes et al. Citation2021). Among this remarkable biodiversity, the Mammillaria genus stands out with almost 200 universally accepted taxa, marking it as the genus with the greatest number of species (https://cactiguide.com/cactus/?genus=Mammillaria). One of the most cherished species within this genus is M. elongata. This petite cactus is easily propagated and favored among plant enthusiasts. Native to Mexico, it prospers in Guanajuato, Hidalgo, and Querétaro states. It is adapted to altitudes that range from 1300 to 2300 m above sea level, demonstrating its ability to flourish under such conditions (Breslin et al. Citation2021). Despite the extensive use of chloroplast genomes for phylogenetic analysis, there is a surprising scarcity of publicly available data for Mammillaria species on the NCBI database. Despite the genus’s rich diversity, only a handful of Mammillaria chloroplast genomes have been unveiled. Consequently, the phylogenetic relationships of M. elongata remain shrouded in uncertainty.
Materials and methods
We selected the sample from China to explore genetic and evolutionary differences in the chloroplast genome between this sample and the native Mammillaria populations in Mexico, further studying the adaptive evolution of M. elongata. Our sampling site was at the College of Horticulture and Landscape Architecture, Southwest University, No. 2 Tiansheng Road, Chongqing, China, 400716. The site’s geospatial coordinates are N29.842889, E106.394527 (). We employed a Magnetic Plant Genomic DNA Kit (DP342, Tiangen, China) for DNA extraction. The subsequent library construction was carried out using the NEB Next Ultra DNA Library Prep Kit for Illumina (NEB, Ipswich, MA), and an insert size of 350 bp was achieved. Sequencing procedures were conducted on the NovaSeq 6000 platform. The plant samples stored at the herbarium of the Institute of Medicinal Plant Development with the accession number Implad ME01 (Institute website: http://www.implad.ac.cn/, Contact person: Yang Ni, E-mail: [email protected]).
Figure 1. Photographs of M. elongata (the photograph were taken by prof. Haimei Chen). the image was captured at the Horticulture and Landscape Architecture College of Southwest University, Chongqing, China. The geospatial reference for this location is N29.842889, E106.394527. This visual representation demonstrates the distinctive features of the compact cactus, which is native to Mexico and highly appreciated for its ease of propagation in horticultural practice.

The chloroplast genome was directly assembled de novo from this raw data, utilizing the GetOrganelle software with its default settings (Jin et al. Citation2020). The assembled genome was assessed rigorously: original sequencing results were aligned with the reference genome using BWA software (Li and Durbin Citation2009), genome coverage depth was computed using Samtools (Li et al. 2009), and a line graph was constructed using a customized Python script (https://www.protocols.io/view/generating-sequencing-depth-and-coverage-map-for-o-4r3l27jkxg1y/v1). The Gepard software aided in verifying the assembly’s start point and direction accuracy (Krumsiek et al. Citation2007). Upon verification, we performed genome annotation using the CPGAVAS2 software and data set 2 (Shi et al. Citation2019), with the annotation results subsequently inspected and visualized using CPGView (Liu et al. Citation2023). Any detected annotation inaccuracies were manually corrected with the Apollo (Lewis et al. Citation2002) software. The final annotated results were submitted to the NCBI database through the Bankit software.
Phylogenetic analysis was initiated by downloading eight chloroplast genomes from the NCBI database. Shared genes were extracted using PhyloSuite software (Zhang et al. Citation2020), and the identified gene clusters underwent multiple sequence alignments using MAFFT (Katoh and Standley Citation2013). A Maximum Likelihood (ML) tree was subsequently built using the IQTREE2 software (Nguyen et al. Citation2015), adopting the best-fit model as per the Bayesian Information Criterion (BIC), which was GTR + F + I + G4. Branch supports were evaluated using the ultrafast bootstrap method with 1000 replicates (UFBoot) (Hoang et al. Citation2018).
Results
The M. elongata chloroplast genome spans 110,981 base pairs (bp), featuring an overall GC content of 36.28%. The genome assembly quality was assessed. The average sequencing coverage depth is 2396.22x, with a maximum depth of 3586x and a minimum of 921x (Figure S1). The genome is structurally organized into four distinct regions: two inverted repeats (IR) regions, each 1711 bp long with a GC content of 38.05%; a small single copy (SSC) region spanning 28,985 bp and boasting a GC content of 38.06%; and a large single copy (LSC) region, which is 78,574 bp in length and has a GC content of 35.55% (; Table S1).
Figure 2. The chloroplast genome of Mammillaria elongata visualized by CPGView. The outer circle depicts the locations of protein-coding genes, tRNA genes, and rRNA genes along the 110,981 bp circular chloroplast chromosome. Genes are color-coded by function: photosystem I (orange), photosystem II (red), cytochrome b/f complex (purple), ATP synthase (blue), rubisco large subunit (green), RNA polymerase (teal), ribosomal proteins (small subunit in pink, large subunit in grey), other (light blue). The inner circle indicates the genome position in kilobase pairs.
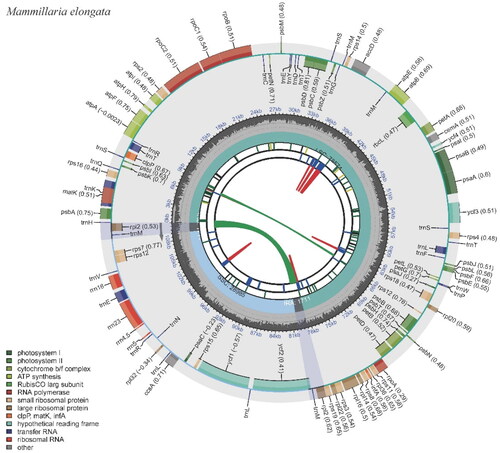
This chloroplast genome encompasses 100 genes, of which 95 are unique. The genes comprise 67 protein-coding genes (with 66 of them being unique), 29 tRNA genes (of which 25 are unique), and four rRNA genes, all unique (Table S2). Notably, several anticipated protein-coding genes are absent. The absent genes include the complete suite of subunit genes for the NADH-dehydrogenase complex (namely ndhA, ndhB, ndhC, ndhD, ndhE, ndhF, ndhG, ndhH, ndhI, ndhJ, and ndhK), as well as two large ribosomal subunit proteins, rpl23 and rpl33. The genome also lacks the conserved open reading frame, ycf15. Embedded within the genome are 13 genes that carry introns (Figure S2). Among these, 11 genes harbor a single intron and two genes with two introns (Figure S3; Table S2). Interestingly, the rpl2 gene has lost its intron. To summarize, most intron-carrying genes in this chloroplast genome possess one intron, two genes contain two introns, and the intron typically present in the two copies of the rpl2 gene is absent (Table S2).
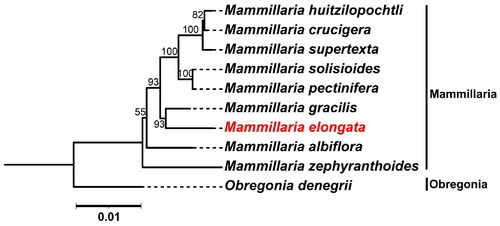
We conducted a phylogenetic analysis using 59 shared genes across nine Mammillaria species and one Obregonia species (serving as an outgroup) (Yu et al., Citation2023). In the resulting phylogenetic topology, M. elongata and M. gracilis are distinctly clustered. This grouping suggests a close genetic relationship between these two species, which may point to a relatively recent common ancestor. Furthermore, this close genetic relationship could suggest that these species have experienced similar evolutionary pressures or inhabited shared ecological niches. Future research focusing on these genetic connections and their ecological implications could further elucidate the biodiversity and conservation needs of this significant group of cacti ().
Figure 3. Maximum Likelihood (ML) phylogenetic tree Illustrating the relationship between M. elongata and eight other species within the Mammillaria genus based on chloroplast genome sequences. Obregonia species are utilized as outgroup references. M. elongata (MW553058.1) is distinctively marked in red. The sequences incorporated in the tree are as follows: M. huitzilopochtli (MN517612.1) (Solórzano et al., Citation2019), M. supertexta (MN508963.1) (Solórzano et al., Citation2019), M. solisioides (MN518341.1) (Solórzano et al., Citation2019), M. pectinifera (MN519716.1) (Solórzano et al., Citation2019), M. gracilis (MW553059.1) (Yu et al., Citation2023), M. albiflora (MN517610.1) (Solórzano et al., Citation2019), M. zephyranthoides (MN517611.1) (Solórzano et al., Citation2019), and O. denegrii (MW553062.1) (Yu et al., Citation2023). the scale bar represents a genetic distance of 0.1 substitutions per site.
Discussion and conclusion
Within the Caryophyllales order, the typical length of chloroplast genomes hovers around 150–160 kb, with the IR region generally spanning approximately 25,000 bp (Downie et al. Citation1997; Choi et al. Citation2018; Yang et al. Citation2019). However, within the cactus genus Mammillaria, the chloroplast genome of M. elongata stands at a mere 110,981 bp. The chloroplast genome size is notably shorter than the chloroplast genomes found in other species of the Caryophyllales order, suggesting multiple genomic contractions and expansions throughout its evolutionary history (Chincoya et al. Citation2020). The canonical structure of chloroplast genomes is quadripartite. Intriguingly, the IR region in the M. elongata chloroplast genome is just 1,711 bp, significantly less than the standard 25,000 bp. This phenomenon points to the contraction and expansion events within the IR region as a likely major contributor to the size variation of the M. elongata chloroplast genome. This observation aligns with previous reports on M. gracilis from the same genus, which also exhibited a significantly contracted IR region, measuring only 1693 bp (Solórzano et al., Citation2019; Yu et al., Citation2023).
The ndh genes are pivotal in encoding the NADH complex within the chloroplast, instrumental to photosynthesis and vital for plant growth and maturation (Krause Citation2008). Intriguingly, our genomic annotations spotlight a notable void of the ndh gene cluster in M. elongata. While prior studies underscore that this deficiency is not comprehensive, traces of some ndh gene fragments remain discernible (Yu et al., Citation2023). Furthermore, the scientific realm is embroiled in a discourse debating if the ndh genes faced an all-encompassing loss or gradual attrition throughout evolutionary periods (Ranade et al. Citation2016), emphasizing the need for more exhaustive research. Evidence from both the Orchidaceae family and the parasitic plants of the genus Cuscuta in the Convolvulaceae family points to the varying degrees of ndh gene loss (Kim et al. Citation2015; Ni et al. Citation2021). One hypothesis posits that the apparent loss of the ndh gene family may stem from the transfer of their expression to the nuclear genome. This transfer would then allow these genes to remain functional, albeit from a different genomic location (Stegemann et al. Citation2003). Supporting this hypothesis are observations from the nuclear genome of Phalaenopsis aphrodite, where complete sequences of ndhA, ndhF, and ndhH have been identified (Cai et al. Citation2015). This suggests the potential relocation of the original ndh genes to the nuclear genome. To summarize, the observed loss or diminution of ndh genes in certain plant species appears to be a multifaceted process, potentially entailing gene transfer events as well as ecological and evolutionary adaptations.
The chloroplast genome has consistently emerged as a pivotal molecular marker in conservation-focused studies pertaining to cacti. For example, investigations centered on the Copiapoa genus, which is native to the Chilean Atacama Desert, harnessed data from chloroplast sequences coupled with microsatellite analyses (Fava et al. Citation2020). This dual-data approach facilitated a comprehensive assessment of the distribution and potential extinction vulnerabilities of particular taxa, unearthing pronounced extinction susceptibilities for certain members within the genus (Larridon et al. Citation2018; Fava et al. Citation2020). A parallel study, honing in on the Sclerocactus species, underscored the importance of integrating nuclear microsatellites with chloroplast DNA sequence data. This integrated method proved instrumental in discerning genetic variances and in pinpointing minimal hybridization occurrences between distinct species (Schwabe et al. Citation2015). Collectively, these endeavors reinforce the instrumental role of the chloroplast genome of cacti in advancing conservation biology. Building on this foundation, our exploration of M. elongata paves the way for broader applications of chloroplast genomics, particularly in elucidating and safeguarding the genetic fabric of cacti species under threat of extinction. However, it’s important to note that our samples come from China, whereas M. elongata is originally from Mexico, which might affect the reflection of its native genetic diversity.
We sequenced and analyzed the complete chloroplast genome of the M. elongata cactus for the first time. The genome is about 111,000 bp long and contains 100 genes, 95 of which are unique. Some genes were found missing in this chloroplast genome. Additionally, our research showed that M. elongata and another cactus species, M. gracilis, are closely related, suggesting they may share a recent common ancestor and similar evolutionary histories or habitats. This work provides valuable genetic data for further study of M. elongata and can help us understand cacti’s evolution and conservation needs in the Mammillaria genus and beyond.
Ethical approval
The manuscript does not encompass any research conducted on human subjects or animals by the authors. The flora explored in this study is not classified as endangered, protected, or privately owned. The plant samples were collected in strict adherence to the regulations put forth by the national authorities. Furthermore, guidelines established by our affiliated institution, the College of Horticulture and Landscape Architecture at Southwest University, were meticulously followed. The university is at No. 2 Tiansheng Road, Chongqing, China, 400716.
Author contributions
This study was conceptualized and structured by Haimei Chen and Jingling Li. Yang Ni and Jingling Li assemble and annotate the chloroplast genome. Qiqian Lu undertook the task of conducting the phylogenetic analysis. Yang Ni wrote the manuscript. Haimei Chen and Yang Ni reviewed the manuscript critically. All authors have perused and consented to the published iteration of the manuscript.
Supplemental Material
Download MS Word (205.6 KB)Disclosure statement
The authors reported no potential conflict of interest.\
Data availability statement
The genome sequence data supporting this study’s findings have been deposited in the NCBI GenBank and can be accessed at https://www.ncbi.nlm.nih.gov/ with the accession number MW553058.1. Corresponding BioProject, Sequence Read Archive (SRA), and BioSample entries are available under the identifiers PRJNA995908, SRR25317666, and SAMN36509928, respectively.
Additional information
Funding
References
- Abouseadaa HH, Atia MAM, Younis IY, Issa MY, Ashour HA, Saleh I, Osman GH, Arif IA, Mohsen E. 2020. Gene-targeted molecular phylogeny, phytochemical profiling, and antioxidant activity of nine species belonging to family Cactaceae. Saudi J Biol Sci. 27(6):1649–1658. doi: 10.1016/j.sjbs.2020.03.007.
- Breslin PB, Wojciechowski MF, Majure LC. 2021. Molecular phylogeny of the Mammilloid clade (Cactaceae) resolves the monophyly of Mammillaria. Taxon. 70(2):308–323. doi: 10.1002/tax.12451.
- Cai J, Liu X, Vanneste K, Proost S, Tsai WC, Liu KW, Chen LJ, He Y, Xu Q, Bian C, et al. 2015. The genome sequence of the orchid Phalaenopsis equestris. Nat Genet. 47(1):65–72. doi: 10.1038/ng.3149.
- Cervantes CR, Hinojosa-Alvarez S, Wegier A, Rosas U, Arias S. 2021. Evaluating the monophyly of Mammillaria series Supertextae (Cactaceae). PhytoKeys. 177:25–42. doi: 10.3897/phytokeys.177.62915.
- Chincoya DA, Sanchez-Flores A, Estrada K, Díaz-Velásquez CE, González-Rodríguez A, Vaca-Paniagua F, Dávila P, Arias S, Solórzano S. 2020. Identification of high molecular variation loci in complete chloroplast genomes of mammillaria (Cactaceae, Caryophyllales). Genes (Basel). 11(7):830. doi: 10.3390/genes11070830.
- Choi KS, Kwak M, Lee B, Park S. 2018. Complete chloroplast genome of Tetragonia tetragonioides: molecular phylogenetic relationships and evolution in Caryophyllales. PLoS One. 13(6):e0199626. doi: 10.1371/journal.pone.0199626.
- Downie SR, Katz-Downie DS, Cho K-J. 1997. Relationships in the Caryophyllales as suggested by phylogenetic analyses of partial chloroplast DNA ORF2280 homolog sequences. Am J Bot. 84(2):253–273. doi: 10.2307/2446087.
- Fava WS, Gomes VGN, Lorenz AP, Paggi GM. 2020. Cross-amplification of microsatellite loci in the cacti species from Brazilian Chaco. Mol Biol Rep. 47(2):1535–1542. doi: 10.1007/s11033-019-05064-3.
- Hoang DT, Chernomor O, von Haeseler A, Minh BQ, Vinh LS. 2018. UFBoot2: improving the ultrafast bootstrap approximation. Mol Biol Evol. 35(2):518–522. doi: 10.1093/molbev/msx281.
- Jin JJ, Yu WB, Yang JB, Song Y, dePamphilis CW, Yi TS, Li DZ. 2020. GetOrganelle: a fast and versatile toolkit for accurate de novo assembly of organelle genomes. Genome Biol. 21(1):241. doi: 10.1186/s13059-020-02154-5.
- Katoh K, Standley DM. 2013. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol. 30(4):772–780. doi: 10.1093/molbev/mst010.
- Kim HT, Kim JS, Moore MJ, Neubig KM, Williams NH, Whitten WM, Kim JH. 2015. Seven new complete plastome sequences reveal rampant independent loss of the NDH gene family across orchids and associated instability of the inverted repeat/small single-copy region boundaries. PLoS One. 10(11):e0142215. doi: 10.1371/journal.pone.0142215.
- Krause K. 2008. From chloroplasts to “cryptic” plastids: evolution of plastid genomes in parasitic plants. Curr Genet. 54(3):111–121. doi: 10.1007/s00294-008-0208-8.
- Krumsiek J, Arnold R, Rattei T. 2007. Gepard: a rapid and sensitive tool for creating dotplots on genome scale. Bioinformatics. 23(8):1026–1028. doi: 10.1093/bioinformatics/btm039.
- Larridon I, Veltjen E, Semmouri I, Asselman P, Guerrero PC, Duarte M, Walter HE, Cisternas MA, Samain MS. 2018. Investigating taxon boundaries and extinction risk in endemic Chilean cacti (Copiapoa subsection Cinerei, Cactaceae) using chloroplast DNA sequences, microsatellite data and 3D mapping. Kew Bull. 73(4):55. doi: 10.1007/s12225-018-9780-3.
- Lewis SE, Searle SMJ, Harris N, Gibson M, Lyer V, Richter J, Wiel C, Bayraktaroglu L, Birney E, Crosby MA, et al. 2002. Apollo: a sequence annotation editor. Genome Biol. 3(12):RESEARCH0082. doi: 10.1186/gb-2002-3-12-research0082.
- Li H, Durbin R. 2009. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 25(14):1754–1760. doi: 10.1093/bioinformatics/btp324.
- Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R. The sequence alignment/map format and SAMtools. Bioinformatics. 25(16):2078–2079. doi: 10.1093/bioinformatics/btp352.
- Liu S, Ni Y, Li J, Zhang X, Yang H, Chen H, Liu C. 2023. CPGView: a package for visualizing detailed chloroplast genome structures. Mol Ecol Resour. 23(3):694–704. doi: 10.1111/1755-0998.13729.
- Nguyen LT, Schmidt HA, Von Haeseler A, Minh BQ. 2015. evolution: IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol. 32(1):268–274. doi: 10.1093/molbev/msu300.
- Ni Y, Jiang M, Chen H, Huang L, Chen P, Liu C. 2021. Adaptation of a parasitic lifestyle by Cuscuta gronovii Willd. ex Roem. & Schult.: large scale gene deletion, conserved gene orders, and low intraspecific divergence. Mitochondrial DNA B Resour. 6(4):1475–1482. doi: 10.1080/23802359.2021.1911702.
- Novoa A, Le Roux JJ, Robertson MP, Wilson JRU, Richardson DM. 2014. Introduced and invasive cactus species: a global review. AoB Plants. 7 doi: 10.1093/aobpla/plu078.
- Ranade SS, García-Gil MR, Rosselló JA. 2016. Non-functional plastid ndh gene fragments are present in the nuclear genome of Norway spruce (Picea abies L. Karsch): insights from in silico analysis of nuclear and organellar genomes. Mol Genet Genomics. 291(2):935–941. doi: 10.1007/s00438-015-1159-7.
- Schwabe AL, Neale JR, McGlaughlin ME. 2015. Examining the genetic integrity of a rare endemic Colorado cactus (Sclerocactus glaucus) in the face of hybridization threats from a close and widespread congener (Sclerocactus parviflorus). Conserv Genet. 16(2):443–457. doi: 10.1007/s10592-014-0671-3.
- Shi L, Chen H, Jiang M, Wang L, Wu X, Huang L, Liu C. 2019. CPGAVAS2, an integrated plastome sequence annotator and analyzer. Nucleic Acids Res. 47(W1):W65–W73. doi: 10.1093/nar/gkz345.
- Solórzano S, Chincoya DA, Sanchez-Flores A, Estrada K, Díaz-Velásquez CE, González-Rodríguez A, Vaca-Paniagua F, Dávila P, Arias S. De novo assembly discovered novel structures in genome of plastids and revealed divergent inverted repeats in Mammillaria (Cactaceae, Caryophyllales). Plants 2019;8(10):392. doi: 10.3390/plants8100392.
- Stegemann S, Hartmann S, Ruf S, Bock R. 2003. High-frequency gene transfer from the chloroplast genome to the nucleus. Proc Natl Acad Sci USA. 100(15):8828–8833. doi: 10.1073/pnas.1430924100.
- Yang L, Li Q, Zhao G. 2019. Characterization of the complete chloroplast genome of Chenopodium sp. (Caryophyllales: Chenopodiaceae). Mitochondrial DNA B Resour. 4(2):2574–2575. doi: 10.1080/23802359.2019.1640089.
- Yu J, Li J, Zuo Y, Qin Q, Zeng S, Rennenberg H, Deng H. 2023. Plastome variations reveal the distinct evolutionary scenarios of plastomes in the subfamily Cereoideae (Cactaceae). BMC Plant Biol. 23(1):132. doi: 10.1186/s12870-023-04148-4.
- Zhang D, Gao F, Jakovlić I, Zou H, Zhang J, Li WX, Wang GT. 2020. PhyloSuite: an integrated and scalable desktop platform for streamlined molecular sequence data management and evolutionary phylogenetics studies. Mol Ecol Resour. 20(1):348–355. doi: 10.1111/1755-0998.13096.
