Abstract
Noncoding RNAs (ncRNAs) can be divided into microRNAs (miRNAs), long noncoding RNAs (lncRNAs), circular RNAs (circRNAs), pRNAs, and tRNAs. Traditionally, miRNAs exert their biological function mainly through the inhibition of translation via the induction of target RNA transcript degradation. lncRNAs and circRNAs were once considered to have no potential to code proteins. Here, we will review the current knowledge on ncRNAs in relation to their origins, characteristics, and functions. We will also review how ncRNAs work as competitive endogenous RNA, gene transcription and expression regulators, and RNA-binding protein sponges in colorectal cancer (CRC). Notably, except for the abovementioned mechanisms, recent advances revealed that lncRNAs can also act as the precursor of miRNAs, and a small portion of lncRNAs and circRNAs was verified to have the potential to code proteins, providing new evidence for the significance of ncRNAs in CRC tumorigenesis and development.
Introduction
Noncoding RNAs (ncRNAs) are major components of the human transcriptome.Citation1 Recently, ncRNAs were demonstrated to play important roles in multiple biological processes by directly or indirectly interfering with gene expression in various cancers. The regulatory role of ncRNAs in multiple cancers has been summarized, such as lncRNAs and microRNAs (miRNAs) in endocrine-related cancers,Citation2 lncRNAs in hepatocarcinogenesis,Citation3 and circRNAs in multiple types of cancer.Citation4 However, a comprehensive and in-depth analysis of ncRNAs in CRC has not been reported to date.
ncRNAs account for the majority of RNA transcribed by human genes, including miRNA, long noncoding RNA (lncRNA), circular RNA (circRNA), pRNA, and tRNA. With the development of RNA sequencing technologies and bioinformatics, numerous ncRNAs have been discovered that influence gene expression levels via chromatin modification, transcription, and posttranscriptional processing.Citation5 Moreover, the abnormal expression of ncRNAs is associated with invasion, metastasis, chemoresistance, and radioresistance of colorectal cancer (CRC).Citation6 For instance, TUG1 regulates the expression of growth-related genes, activates the expression of epithelial-to-mesenchymal transition (EMT)-associated genes, and plays important roles in signal transduction, cell morphology, migration, proliferation, and apoptosis in CRC. Overexpression of TUG1 is thought to be an independent poor prognostic factor for CRC patients.Citation7,Citation8 The circRNA circ_001569 is upregulated in CRC tissues and promotes CRC proliferation and invasion. This circRNA acts as a sponge to directly inhibit miR-145 transcription, which subsequently affects the functions of miR-145 targets E2F5, BAG4, and FMNL2 in CRC cells.Citation9
Herein, we performed a systematic literature review analysis of ncRNAs in CRC and the dysregulation of ncRNAs in CRC tissues or cells. Then, we discussed how these ncRNAs work as miRNA sponges, gene transcription and expression regulators, and RNA-binding protein (RBP) sponges in CRC, providing evidence for the significance of ncRNAs in CRC.
Classification of ncRNAs
Generally, according to their product size, ncRNAs can be divided into two groups: small ncRNAs and long ncRNAs.Citation10 The size of small ncRNAs, such as miRNA, is typically less than 200 nucleotides (nt).Citation11,Citation12 By contrast, long ncRNAs are typically greater than 200 nt, including long intergenic ncRNAs, long intronic ncRNAs, and pseudogene RNAs.Citation13 Except for these linear ncRNAs, circRNAs, which are formed through the ligation of the 5′ and 3′ ends of linear RNA, have been investigated recently.Citation14,Citation15 Accumulating evidence has revealed that aberrant ncRNA expression is correlated with various cancers, especially CRC.
Mechanisms of ncRNAs in regulating CRC progression
ncRNAs control individual genes and gene expression programs through changing the fundamental transcriptional mechanism or via epigenetic regulation at multiple levels, such as transcription, translation, and protein function ().
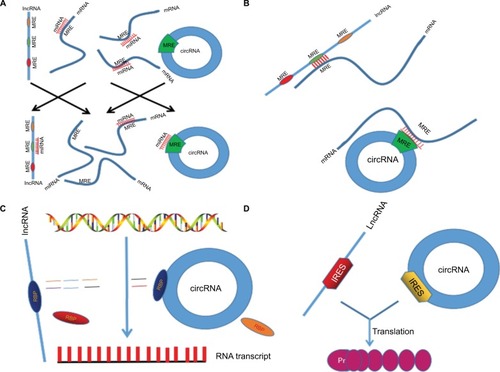
Figure 1 Mechanism of lncRNAs/circRNAs regulating CRC biological activities. (A) lncRNAs and circRNAs act as miRNA sponge or ceRNA. (B) Directly targeting mRNA by partial base pairing. (C) Binding RBP to regulate protein expression. (D) A small portion of lncRNAs/circRNAs can be translated to proteins.
Abbreviations: lncRNAs, long noncoding RNAs; circRNAs, circular RNAs; CRC, colorectal cancer; miRNA, microRNA; ceRNA, competitive endogenous RNA; IRES, internal ribosome entry site; RBP, RNA-binding protein.
As a competitive endogenous RNA (ceRNA) or miRNA sponge
miRNA is a small ncRNA that is 19–24 nt in length. miRNA binds to miRNA response elements (MREs) in RNA sequences and negatively regulates gene expression through the inhibition of translation via the induction of target RNA transcript degradation.Citation16,Citation17 ceRNA contains an MRE, which competitively binds to miRNA. Therefore, ceRNA affects the regulatory functions of miRNAs in gene expression and reduces the inhibitory effect of miRNAs on target moleculesCitation18–Citation21 (). For instance, UCC may act as an endogenous sponge by competing for miR-143, thereby regulating the targets of this miRNA. UCC and miR-143 may be promising molecular targets for CRC therapy.Citation22 The lncRNA CRNDE regulates the progression and chemoresistance of CRC via modulating the expression levels of miR-181a-5p and the activity of Wnt/β-catenin signaling.Citation23 HOXA11-AS promotes liver metastasis in CRC by functioning as a miR-125a-5p sponge, and the novel HOXA11-AS-miR-125a-5p-PADI2 regulatory network is involved in CRC liver metastasis.Citation24 At present, the main function of some lncRNAs and circRNAs involves acting as a miRNA sponge.
Regulating gene transcription
In addition to the abovementioned mechanisms, ncRNAs, as the product of transcription, are major regulators of the transcriptional process.Citation25 Researchers reported that ncRNAs can function as positive regulators of their parental gene transcription, targeting mRNA by partially base pairingCitation26 (). Accumulating evidence indicates that ncRNAs play a pivotal role in posttranscriptional and gene expression regulation. For example, the circRNAs circ-EIF3J and circ-PAIP2 combine with the U1 snRNP to further interact with RNA Pol II and enhance the expression of their parental genes in HeLa and HEK293 cells.Citation27
Regulating RBPs
miRNA biogenesis is modulated by a variety of RBPsCitation28 (), and the adsorption of protein factors by linear lncRNA has been reported.Citation29–Citation31 For example, sno-lncRNAs regulate alternative splicing of downstream genes by adsorbing the alternative splicing factor Fox2.Citation32 NEAT1 also has a profound effect on global pri-miRNA processing. Mechanistic dissection reveals that NEAT1 broadly interacts with the NONO-PSF heterodimer and numerous other RBPs. In addition, multiple RNA segments in NEAT1, including a pseudo pri-miRNA near its 3′ end, help attract the microprocessor.Citation33
Protein translation
A small portion of ncRNAs can also be translated to proteins similar to mRNAs. circRNA is efficiently translated in living human cells to produce abundant protein product via the RCA mechanism.Citation34 Long-repeating polypeptide chains were synthesized from RNA circles with continuous open reading frames, and an internal ribosome entry site and the initiation codon ATG in a specific circRNA allow the circRNA translation template to function as mRNA ().Citation35
Biological functions of miRNAs in CRC
miRNAs are 19–24 nt in length that regulate gene expression at the posttranscriptional level by binding to the 3′-untranslated regions or the open reading frames of target genes, leading to the degradation of target mRNAs or repression of mRNA translation.Citation36 Numerous investigations have been performed to analyze the biological functions of miRNAs in CRC.Citation37,Citation38
miRNAs promote CRC cell proliferation and invasion
miRNAs exhibit a close relationship with the initiation and development of various human malignancies.Citation39 Abundant miRNAs play oncogenic roles in CRC tumorigenesis via diverse mechanisms.Citation40,Citation41 On one hand, miRNAs exert their tumor oncogenic functions primarily by binding to the 3′-untranslated region of the mRNA of target genes. For instance, miR19b-3p is overexpressed in CRC tissues compared with normal tissues. Furthermore, miR19b-3p plays an oncogenic role in CRC via directly targeting ITGB8.Citation42 Similarly, TIA1 is an important tumor suppressor in CRC. In addition, miR-19a is highly expressed in tumor tissues compared with normal tissues and exerts its oncomiR function by targeting TIA1.Citation43 On the other hand, miRNAs promote CRC cell proliferation and invasion via other multiple mechanisms. For example, the EMT plays important roles in tumor metastasis. For example, miR-19a is upregulated in CRC tissues, and high expression of miR-19a is signifi-cantly associated with lymph node metastasis. Interestingly, miR-19a is upregulated by TNF-α and is required for TNF-α-induced EMT and metastasis in CRC cells.Citation44
miRNAs suppress CRC cell proliferation and invasion
By contrast, miRNA may function as a tumor-repressive gene to inhibit cell proliferation in CRC. Some miRNAs suppress CRC cell proliferation and invasion by regulating tumor angiogenesis, tumor metabolism, and cancer stemness features.Citation45,Citation46 For instance, miR-590-5p overexpression inhibits CRC angiogenesis and metastasis by targeting nuclear factor 90, which acts as a positive regulator of vascular endothelial growth factor mRNA stability and protein synthesis. Notably, knockdown of miR-590-5p promotes the progression of CRC in vitro.Citation47 AEG-1 is a key oncogenic factor in various tumors.Citation48 In addition, miR-217 suppresses CRC cell proliferation and invasiveness by inhibiting AEG-1 expression with modulation of MMP2 or AMPK signaling.Citation49,Citation50
Biological functions of lncRNAs in CRC
lncRNA is defined as an RNA transcript of greater than 200 nt located in nuclear or cytosolic fractions.Citation51 The overexpression, deficiency, or mutation of lncRNAs could be involved in CRC progression via a variety of mechanisms.Citation52,Citation53 Abundantly expressed lncRNAs play crucial roles in CRC, such as HOTAIR, CCAT1, and MALAT1 ().
Table 1 lncRNAs reported in CRC
lncRNAs interact with miRNAs in CRC
The ceRNA hypothesis demonstrated that lncRNAs with shared miRNA binding sites compete for posttranscriptional control.Citation54 In early studies, it was found that PTENP1 shares conserved miRNA seed target sites with PTEN for the miR-17, miR-21, miR-214, miR-19, and miR-26 miRNA families. The role of lncRNAs as a miRNA sponge is the main mechanism of lncRNAs in cancer. For example, H19 is overexpressed in colon cancer tissues and cell lines, whereas miR-138 expression in tumor tissues is reduced compared with normal tissues. Interestingly, the silencing of H19 strongly increases the expression of miR-138 and suppresses the expression of high-mobility group A (HMGA1) protein, indicating that H19 promotes the migration and invasion of colon cancer by sponging miR-138 to upregulate HMGA1 expression.Citation55
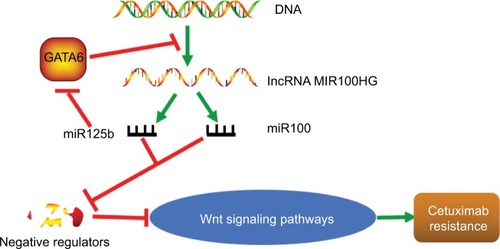
Notably, lncRNAs act as the precursor of miRNAs. For example, the lncRNA MIR100HG is the precursor of miR-100 and miR-125b, which coordinately represses five Wnt/β-catenin negative regulators, leading to cetuximab resistance. Furthermore, the transcription factor GATA6 represses MIR100HG. By contrast, the repression is relieved by miR-125b targeting of GATA6, revealing a double-negative feedback loop between MIR100HG and the transcription factor GATA6 (). In addition, miR-17-5p expression is increased in tumors compared with paired non-tumorous tissues, and high miR-17-5p expression is significantly associated with TNM staging and lymph node metastasis. Furthermore, miR-17-5p expression is upregu-lated by CCAT2 through TCF7 L2-mediated transcriptional regulation, enhancing WNT activity.Citation56
lncRNAs function through other mechanisms in CRC
Tumor metabolism is responsible for rapid recurrence and poor survival of CRC. The relationship between ncRNAs and tumor metabolism was investigated recently. For instance, lncRNA NONHSAT062994 expression was negatively correlated with the Akt downstream targets c-Myc and cyclin D1 in clinical CRC samples, indicating that NONHSAT062994 functioned as a tumor suppressor to inhibit CRC cell growth by inactivating Akt signaling.Citation57 Autophagy is a crucial intracellular process associated with CRC tumorigenesis and progression. lncRNAs play crucial roles in CRC by regulating tumor autophagy by targeting various proteins, such as CPS1-IT1, AC023115.3, and MALAT1.Citation58,Citation59 For example, the lncRNA CPS1-IT suppresses metastasis and EMT by inhibiting hypoxia-induced autophagy through inactivation of HIF-1α in CRC.Citation60
Biological functions of circRNAs in CRC
circRNAs are a special type of endogenous ncRNA molecule. We have witnessed an explosion in published studies on all aspects of circRNA biology leading to the common understanding that these molecules are important players in cancers, especially in CRC. As previously mentioned, circRNAs differ structurally from other lncRNAs given that their 3′ and 5′ ends are covalently joined. Certain circRNAs act as highly stable sponges for specific miRNAs, such as CiRS-7 and SRY for miR-7 and miR-138, respectively, and are involved in competing endogenous RNA networks.Citation61 Thousands of circRNAs have been identified and annotated in the circRNA repository (circBase). However, the biogenesis of circRNAs remains elusive.Citation62
miRNAs are extremely well studied as tumor suppressors or oncogenes. It has been hypothesized that circRNAs form a class of posttranscriptional regulators, acting as epigenetic, highly stable miRNA sponges to compete with the endogenous RNA network and directly affecting the expression of any related gene.Citation63 circRNAs sequester specific miRNA complexes and release them after cleavage.Citation64 Circ_001569 was upregulated in CRC tissues compared with adjacent normal tissues, and high circ_001569 expression was closely correlated with differentiation and TNM classification. Interestingly, circ_001569 is negatively correlated with miR-145, and miR-145 is negatively correlated with E2F5, BAG4, and FMNL2 expression. These data indicate that circ_001569 promotes CRC cell proliferation and invasion by regulating miR-145 and its targets E2F5, BAG4, and FMNL2.Citation65 In addition, circ_0020397 promotes CRC cell viability and invasion and inhibits their apoptosis by promoting the expression of the miR-138 target genes TERT and PD-L1.Citation66 circCCDC66 possesses an oncogenic capacity through protecting multiple oncogenes from being attacked by a group of miRNAs, and overexpression of circCCDC66 potentiates multiple tumor characteristics, including proliferation, migration, and invasion ().Citation67
Figure 3 circRNAs investigated in CRC.
Note: Red lines represent inhibition effect; green arrows indicate promotion effect.
Abbreviations: circRNAs, circular RNAs; CRC, colorectal cancer.
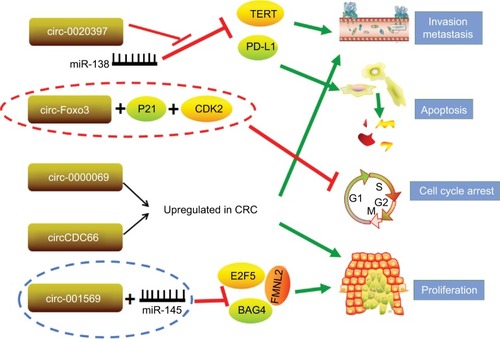
In addition to the abovementioned circRNAs, numerous other circRNAs govern fundamental biological process and cancer progression through multiple mechanisms instead of acting as a special class of endogenous RNAs. circ-Foxo3 functions as a tumor suppressor in CRC by forming circ-Foxo3-p21-CDK2 ternary complexes.Citation68 CDK2 interacts with cyclin A and cyclin E to facilitate cell cycle entry, whereas p21 inhibits these interactions and arrests cell cycle progression.Citation69
Therefore, circRNAs can also exert biological function in CRC by binding to partners.
ncRNAs as new potential tumor biomarkers and targets
An increasing number of ncRNAs are being reported to be aberrantly expressed in CRC, and differential expression of ncRNAs can be detected in the circulation of CRC patients. Importantly, ncRNAs are abundant and their expression is both step and location specific. The characteristics of ncRNAs afford them the potential to become novel diagnostic and prognostic markers for cancer. For instance, lncRNA SPRY4-IT1 is upregulated in CRC tissues and promotes proliferation and invasion by targeting miR-101-3p. These data indicate that knockdown of SPRY4-IT1 represents a rational therapeutic strategy for colorectal carcinoma.Citation70 CircHIPK3 was significantly highly expressed in CRC tissues and positively correlated with metastasis and TNM stage, revealing that circHIPK3 may be a potential prognostic biomarker in CRC.Citation71 On the basis of the results acquired from latest papers, circulating ncRNA profiles are emerging as potential biomarkers of diagnosis, prognosis, and therapeutic response in CRC.
Conclusion
With the development of high-throughput sequencing technologies and bioinformatics methods, ncRNAs are increasingly investigated.Citation72 Traditionally, ncRNA has been viewed as an unstable molecule because ribonucleases are ubiquitous and extremely stable biomarkers for CRC. Emerging evidence have revealed the biological roles and relevant mechanisms of diverse ncRNAs in CRC tumorigenesis.Citation73,Citation74 For example, miR-17 acts as an oncogenic miRNA that promotes CRC development by activating the Wnt/β-catenin pathway and by targeting P130.Citation75 We also found that miR-149 methylation contributes to CRC growth and invasion by targeting the transcription factor Sp1.Citation76
Notably, the molecular mechanisms of these dysregulated ncRNAs have been investigated recent years. EMT is an important step in cancer development that involves the cooperation of a variety of signaling pathways, including the transformation growth factor-β, Sonic Hedgehog, and WNT pathways.Citation77,Citation78 HOTAIR, H19, AFAP1-AS1, TUG1, BANCR, lncRNA-ATB, SPRY4-IT1, SLC25A25-AS1, LINC01133, PANDAR, lncRNA-ATB, and Sox2ot are involved in the EMT pathways.
circRNAs as a new class of ncRNAs contributing to the regulatory network governing protein coding gene expression by acting as miRNA target decoys, RBP sponges and transcriptional regulators exhibit great biological potential for circRNA-related functionalities. Uncovering these additional functions (if any) and understanding these functionalities represent key research topics in the circRNA field in the future.
Via a systematic literature review, we have discussed the origins, characteristics, and main functions of ncRNAs in CRC. Continuous exploration and research in this field will provide an important molecular basis for understanding the complex regulation of CRC.
Acknowledgments
This research was supported by the National Natural Science Foundation of China (81560385); the Medical Scientific and Technological Research Project of Henan Province (201702027); Youth Innovation Fund Project of The First Affiliated Hospital of Zhengzhou University (YNQN2017035); and the China Postdoctoral Science Foundation (2017M610462). Shuaixi Yang, Zhenqiang Sun, and Quanbo Zhou are co-first authors.
Disclosure
The authors report no conflicts of interest in this work.
References
- LuoJQuJWuDKLuZLSunYSQuQLong non-coding RNAs: a rising biotarget in colorectal cancerOncotarget2017813221872220228108736
- KlingeCMNon-coding RNAs: long non-coding RNAs and microRNAs in endocrine-related cancersEndocr Relat Cancer2018254R259R28229440232
- LanzafameMBiancoGTerraccianoLMNgCKYPiscuoglioSThe role of long non-coding RNAs in hepatocarcinogenesisInt J Mol Sci2018193 pii: E682
- JiQZhangCSunXLiQCircular RNAs function as competing endogenous RNAs in multiple types of cancerOncol Lett2018151233029387208
- GuttmanMRinnJLModular regulatory principles of large non-coding RNAsNature2012482783533934622337053
- MengSZhouHFengZcircRNA: functions and properties of a novel potential biomarker for cancerMol Cancer20171619428535767
- WangLZhaoZFengWLong non-coding RNA TUG1 promotes colorectal cancer metastasis via EMT pathwayOncotarget2016732517135171927421138
- ZhaiHYSuiMHYuXOverexpression of long non-coding RNA TUG1 promotes colon cancer progressionMed Sci Monit2016223281328727634385
- XieHRenXXinSEmerging roles of circRNA_001569 targeting miR-145 in the proliferation and invasion of colorectal cancerOncotarget2016718266802669127058418
- WeiZBatagovAOSchinelliSCoding and noncoding landscape of extracellular RNA released by human glioma stem cellsNat Commun201781114529074968
- GongQSuGRoles of miRNAs and long noncoding RNAs in the progression of diabetic retinopathyBiosci Rep2017376BSR2017115729074557
- LongMZhanMXuSmiR-92b-3p acts as a tumor suppressor by targeting Gabra3 in pancreatic cancerMol Cancer201716116729078789
- HuXSoodAKDangCVZhangLThe role of long noncoding RNAs in cancer: the dark matter mattersCurr Opin Genet Dev20184881529054012
- XiaWQiuMChenRCircular RNA has_circ_0067934 is upregulated in esophageal squamous cell carcinoma and promoted proliferationSci Rep201663557627752108
- LiuWMaWYuanYZhangYSunSCircular RNA hsa_cir-cRNA_103809 promotes lung cancer progression via facilitating ZNF121-dependent MYC expression by sequestering miR-4302Biochem Biophys Res Commun2018500484685129698681
- FabianMRSonenbergNFilipowiczWRegulation of mRNA translation and stability by microRNAsAnnu Rev Biochem20107935137920533884
- OuCSunZLiXMiR-590-5p, a density-sensitive microRNA, inhibits tumorigenesis by targeting YAP1 in colorectal cancerCancer Lett2017399536328433598
- LiDYangMLiaoAZengBLinc00483 as ceRNA regulates proliferation and apoptosis through activating MAPKs in gastric cancerJ Cell Mol Med2018 Epub ahead of print
- LiuWHTsaiZTTsaiHKComparative genomic analyses highlight the contribution of pseudogenized protein-coding genes to human lincRNAsBMC Genomics201718178629037146
- WangQJiangSSongAHOXD-AS1 functions as an oncogenic ceRNA to promote NSCLC cell progression by sequestering miR-147aOnco Targets Ther2017104753476329033588
- LiuTChiHChenJCurcumin suppresses proliferation and in vitro invasion of human prostate cancer stem cells by ceRNA effect of miR-145 and lncRNA-RORGene2017631293828843521
- HuangFTChenWYGuZQThe novel long intergenic noncoding RNA UCC promotes colorectal cancer progression by sponging miR-143Cell Death Dis201785e277828492554
- HanPLiJWZhangBMThe lncRNA CRNDE promotes colorectal cancer cell proliferation and chemoresistance via miR-181a-5p-mediated regulation of Wnt/β-catenin signalingMol Cancer2017161928086904
- ChenDSunQZhangLThe lncRNA HOXA11-AS functions as a competing endogenous RNA to regulate PADI2 expression by sponging miR-125a-5p in liver metastasis of colorectal cancerOncotarget2017841706427065229050308
- LongYWangXYoumansDTCechTRHow do lncRNAs regulate transcription?Sci Adv201739eaao211028959731
- ZhangYZhangXOChenTCircular intronic long noncoding RNAsMol Cell201351679280624035497
- LiZHuangCBaoCExon-intron circular RNAs regulate transcription in the nucleusNat Struct Mol Biol201522325626425664725
- G HendricksonDKelleyDRTenenDBernsteinBRinnJLWidespread RNA binding by chromatin-associated proteinsGenome Biol2016172826883116
- WangXSchwartzJCCechTRNucleic acid-binding specificity of human FUS proteinNucleic Acids Res201543157535754326150427
- DavidovichCZhengLGoodrichKJCechTRPromiscuous RNA binding by Polycomb repressive complex 2Nat Struct Mol Biol201320111250125724077223
- HudsonWHOrtlundEAThe structure, function and evolution of proteins that bind DNA and RNANat Rev Mol Cell Biol2014151174976025269475
- YinQFYangLZhangYLong noncoding RNAs with snoRNA endsMol Cell201248221923022959273
- JiangLShaoCWuQJNEAT1 scaffolds RNA-binding proteins and the microprocessor to globally enhance pri-miRNA processingNat Struct Mol Biol2017241081682428846091
- AbeNMatsumotoKNishiharaMRolling circle translation of circular RNA in living human cellsSci Rep201551643526553571
- ChenCYSarnowPInitiation of protein synthesis by the eukaryotic translational apparatus on circular RNAsScience199526852094154177536344
- XiXPZhuangJTengMJMicroRNA-17 induces epithelial-mesenchymal transition consistent with the cancer stem cell phenotype by regulating CYP7B1 expression in colon cancerInt J Mol Med201638249950627278684
- BaSXuanYLongZWChenHYZhengSSMicroRNA-27a promotes the proliferation and invasiveness of colon cancer cells by targeting SFRP1 through the Wnt/β-catenin signaling pathwayCell Physiol Biochem20174251920193328772260
- BasatiGRazaviAEPakzadIMalayeriFACirculating levels of the miRNAs, miR-194, and miR-29b, as clinically useful biomarkers for colorectal cancerTumour Biol20163721781178826318304
- SvoronosAAEngelmanDMSlackFJOncomiR or tumor suppressor? The duplicity of microRNAs in cancerCancer Res201676133666367027325641
- LiuSQuDLiWMiR-647 and miR-1914 promote cancer progression equivalently by downregulating nuclear factor IX in colorectal cancerMol Med Rep20171668189819928990086
- ZhangLFengGZhangXDingYWangXMicroRNA-630 promotes cell proliferation and inhibits apoptosis in the HCT116 human colorectal cancer cell lineMol Med Rep20171644843484828791386
- HuangLCaiJLHuangPZMiR19b-3p promotes the growth and metastasis of colorectal cancer via directly targeting ITGB8Am J Cancer Res201771019962008 eCollection 201729119049
- LiuYLiuRYangFmiR-19a promotes colorectal cancer proliferation and migration by targeting TIA1Mol Cancer20171615328257633
- HuangLWangXWenCHsa-miR-19a is associated with lymph metastasis and mediates the TNF-α induced epithelial-to-mesenchymal transition in colorectal cancerSci Rep201551335026302825
- ZhangRLiuRLiuCA novel role for MiR-520a-3p in regulating EGFR expression in colorectal cancerCell Physiol Biochem20174241559157428738328
- ZhaiHSongBXuXZhuWJuJInhibition of autophagy and tumor growth in colon cancer by miR-502Oncogene201332121570157922580605
- ZhouQZhuYWeiXMiR-590-5p inhibits colorectal cancer angiogenesis and metastasis by regulating nuclear factor 90/vascular endothelial growth factor A axisCell Death Dis2016710e241327735951
- LiuXWangDLiuHKnockdown of astrocyte elevated gene-1 (AEG-1) in cervical cancer cells decreases their invasiveness, epithelial to mesenchymal transition, and chemoresistanceCell Cycle201413111702170724675891
- SongHTianZQinYYaoGFuSGengJAstrocyte elevated gene-1 activates MMP9 to increase invasiveness of colorectal cancerTumour Biol20143576679668524705862
- SongHTQinYYaoGDTianZNFuSBGengJSAstrocyte elevated gene-1 mediates glycolysis and tumorigenesis in colorectal carcinoma cells via AMPK signalingMediators Inflamm2014201428738124829520
- HanDWangMMaNXuYJiangYGaoXLong noncoding RNAs: novel players in colorectal cancerCancer Lett20153611132125754818
- ChenNGuoDXuQLong non-coding RNA FEZF1-AS1 facilitates cell proliferation and migration in colorectal carcinomaOncotarget2016710112711128326848625
- ZhangZZhouCChangYLong non-coding RNA CASC11 interacts with hnRNP-K and activates the WNT/β-catenin pathway to promote growth and metastasis in colorectal cancerCancer Lett20163761627327012187
- LiYLvMSongZLouZWangRZhuangMLong non-coding RNA NNT-AS1 affects progression of breast cancer through miR-142-3p/ZEB1 axisBiomed Pharmacother201810393994629710510
- YangQWangXTangCChenXHeJH19 promotes the migration and invasion of colon cancer by sponging miR-138 to upregulate the expression of HMGA1Int J Oncol20175051801180928358427
- PetrovNZhidkovaOSerikovVZeninVPopovBInduction of Wnt/β-catenin signaling in mouse mesenchymal stem cells is associated with activation of the p130 and E2f4 and formation of the p130/Gsk3β/β-catenin complexStem Cells Dev201221458959721631154
- HeXSGuoLCDuMZThe long non-coding RNA NON-HSAT062994 inhibits colorectal cancer by inactivating Akt signalingOncotarget2017840686966870628978149
- MaBYuanZZhangLLong non-coding RNA AC023115.3 suppresses chemoresistance of glioblastoma by reducing autophagyBiochim Biophys Acta2017186481393140428499919
- YuanPCaoWZangQLiGGuoXFanJThe HIF-2α-MALAT1-miR-216b axis regulates multi-drug resistance of hepatocellular carcinoma cells via modulating autophagyBiochem Biophys Res Commun201647831067107327524242
- ZhangWYuanWSongJWangSGuXlncRNA CPS1-IT1 suppresses EMT and metastasis of colorectal cancer by inhibiting hypoxia-induced autophagy through inactivation of HIF-1αBiochimie2018144212729017924
- ZhaoZJShenJCircular RNA participates in the carcinogenesis and the malignant behavior of cancerRNA Biol201714551452126649774
- GlažarPPapavasileiouPRajewskyNcircBase: a database for circular RNAsRNA201420111666167025234927
- Bachmayr-HeydaAReinerATAuerKCorrelation of circular RNA abundance with proliferation – exemplified with colorectal and ovarian cancer, idiopathic lung fibrosis, and normal human tissuesSci Rep20155805725624062
- HansenTBJensenTIClausenBHNatural RNA circles function as efficient microRNA spongesNature2013495744138438823446346
- CristóbalISanz-AlvarezMTorrejónBPotential therapeutic impact of miR-145 deregulation in colorectal cancerMol Ther2018 S1525-0016(18)302119
- ZhangXLXuLLWangFHsa_circ_0020397 regulates colorectal cancer cell viability, apoptosis and invasion by promoting the expression of the miR-138 targets TERT and PD-L1Cell Biol Int20174191056106428707774
- HsiaoKYLinYCGuptaSKNoncoding effects of circular RNA CCDC66 promote colon cancer growth and metastasisCancer Res20177792339235028249903
- BealeGHaagensenEJThomasHDCombined PI3K and CDK2 inhibition induces cell death and enhances in vivo antitumour activity in colorectal cancerBr J Cancer2016115668269027529512
- DuWWYangWLiuEYangZDhaliwalPYangBBFoxo3 circular RNA retards cell cycle progression via forming ternary complexes with p21 and CDK2Nucleic Acids Res20164462846285826861625
- JinJChuZMaPMengYYangYLong non-coding RNA SPRY4-IT1 promotes proliferation and invasion by acting as a ceRNA of miR-101-3p in colorectal cancer cellsTumour Biol2017397 1010428317716250
- ZengKChenXXuMCircHIPK3 promotes colorectal cancer growth and metastasis by sponging miR-7Cell Death Dis20189441729549306
- WengMWuDYangCNoncoding RNAs in the development, diagnosis, and prognosis of colorectal cancerTransl Res201718110812027810413
- FangZTangJBaiYPlasma levels of microRNA-24, microRNA-320a, and microRNA-423-5p are potential biomarkers for colorectal carcinomaJ Exp Clin Cancer Res2015348626297223
- SunZOuCRenWXieXLiXLiGDownregulation of long non-coding RNA ANRIL suppresses lymphangiogenesis and lymphatic metastasis in colorectal cancerOncotarget2016730475364755527286457
- MaYZhangPWangFElevated oncofoetal miR-17-5p expression regulates colorectal cancer progression by repressing its target gene P130Nat Commun20123129123250421
- WangFMaYLZhangPSP1 mediates the link between methylation of the tumour suppressor miR-149 and outcome in colorectal cancerJ Pathol20132291122422821729
- ZhangJTianXJXingJSignal transduction pathways of EMT induced by TGF-β, SHH, and WNT and their crosstalksJ Clin Med201654 pii: E41
- LamouilleSXuJDerynckRMolecular mechanisms of epithelial-mesenchymal transitionNat Rev Mol Cell Biol201415317819624556840