Abstract
Purpose
LncRNA MAFG-AS1 plays critical roles in several types of cancer, while its role in glioblastoma (GBM) is unknown. By analyzing the TCGA dataset, we observed the upregulation of MAFG-AS1 in GBM. This study aimed to investigate the involvement of MAFG-AS1 in GBM cancer.
Methods
The expression levels of MAFG-AS1, mature miR-34a, and miR-34a precursor in GBM and paired non-tumor tissues of 56 GBM patients were determined by RT-qPCR. Correlations are analyzed using linear regression. Overexpression of MAFG-AS1 was achieved in GBM cells, followed by measurement of the expression levels of mature miR-34a, miR-34a precursor, DICER and Drosha by RT-qPCR. The roles of MAFG-AS1 and miR-34a in regulating GBM cell proliferation were evaluated by CCK-8 assay. Flow cytometry was performed to explore the role of MAFG-AS1 and miR-34a in regulating the apoptosis and cell cycle of GBM cells. Cell scratch experiment was performed to determine the role of MAFG-AS1 and miR-34a in regulating the migration of GBM cells. Subcutaneous tumor animal model was used for in vivo study. The expressions levels of BCL-2 and caspase-3 were detected by Western blot analysis.
Results
MAFG-AS1 was upregulated in GBM, while mature miR-34a was downregulated in GBM. Interestingly, MAFG-AS1 was inversely correlated with mature miR-34a but not miR-34a precursor across GBM tissues. In GBM tissues, the overexpression of MAFG-AS1 did not affect the expression levels of miR-34a precursor but reduced the expression levels of mature miR-34a. MAFG-AS1 promoted the expression of DICER and Drosha in GBM cells. Moreover, the overexpression of MAFG-AS1 promoted the proliferation of GBM cells and reduced the inhibitory effects of miR-34a on cell proliferation but did not affect cell cycle, apoptosis and migration. The overexpression of MAFG-AS1 promoted the progression of GBM in vivo by promoting the proliferation of GBM cells while miR-34a reversed the effect of overexpression of MAFG-AS1.
Conclusions
MAFG-AS1 may suppress the maturation of miR-34a to promote GBM cell proliferation.
Keywords:
Introduction
Despite the low incidence rate (about 3.19 per 100,000),Citation1 glioblastoma (GBM), stage IV glioma, is considered a major cause of cancer-related deaths, mainly due to its extremely aggressive nature.Citation2 The median survival time of GBM patients after diagnosis is only about 16 months, and only about 10% of patients could survive for 5 years even after appropriate treatment.Citation3 At present, GBM patients are usually treated with surgery, radiation therapy, chemotherapy, or tumor treating field therapy.Citation4–Citation6 However, radical treatment for GBM remains lacking. Moreover, the molecular pathogenesis of GBM is still unclear, which limits the development of novel treatment approaches.Citation7
Previous studies on the pathogenesis of GBM have revealed multiple molecular players involved in this disease.Citation8,Citation9 A better understanding of these molecular factors facilitates the development of novel therapeutic approaches, such as targeted therapy that can be performed to suppress cancer progression by regulating the expression of cancer-related gene networks.Citation10 Even without protein-coding capacity, non-coding RNAs (ncRNAs), such as long ncRNAs (lncRNAs) and microRNAs (miRNAs), can regulate cancer development by affecting gene expression at multiple levels.Citation11,Citation12 Therefore, ncRNAs are promising targets for targeted therapy.Citation12 LncRNA MAFG-AS1 plays oncogenic roles in several types of cancer, such as colorectal cancer and lung cancer.Citation13,Citation14 However, the role of MAFG-AS1 in GBM remains unknown. We analyzed the TCGA dataset and observed the upregulation of MAFG-AS1 in GBM. In addition, our preliminary deep sequencing data analysis results showed inverse correlation between MAFG-AS1 and miR-34a, which is a well-characterized tumor suppressive miRNA.Citation15 This study was therefore carried out to investigate the interaction between MAFG-AS1 and miR-34a in GBM.
Materials and Methods
GBM Patients
This study enrolled a total of 56 GBM patients () including 35 males and 21 females who were admitted to Zibo Central Hospital from May 2018 to January 2020. The age of the patients ranged from 50 to 69 years old with a mean age of 61.8 ± 5.5 years old. All GBM patients were newly diagnosed cases and no recurrent GBM patients were included. All the 56 patients were excluded from other clinical disorders and initiated therapies. This study was approved by the Ethics Committee of Zibo Central Hospital and was complied with the Declaration of Helsinki. All patients signed the written informed consent.
Table 1 The Characteristics of GBM Patients
GBM Tissues and Cells
Fine needle aspiration was performed to collect GBM and paired non-tumor tissues from the 56 GBM patients. Following histopathological confirmation, RNA isolation was performed on all tissue samples.
Two human GBM cell lines U87 and U-118 (ATCC, USA) were used. An incubator with 5% CO2 and 95% humidity was used to cultivate the cells at 37 °C. DMEM supplemented with 10% FBS was used as the culture medium.
Cell Transfections
Vector expressing MAFG-AS1 was constructed with pcDNA3.1 (Invitrogen) as the backbone. Mimic of miR-34a and negative control (NC) miRNA were synthesized by Invitrogen. U87 and U-118 cells were counted and 5×107 cells were transfected with either 1 µg vector or 50 nM miRNA using lipofectamine 2000 (Invitrogen)-mediated transient transfections. To perform NC experiments, cells were transfected with empty vector or NC miRNA. Control (C) cells are cells without transfections. After transfection, cells were cultivated in a fresh medium for another 48 h before use.
RNA Preparations
RNAzol reagent was used to isolate total RNAs from paired tissues and U87 and U-118 cells. All RNA samples were digested with DNase I (Invitrogen) at 37 °C for 100 min to remove genomic DNAs. The OD 260/280 ratio was determined to reflect RNA purity. RNA samples were separated by 6% Urine-PAGE gels to check RNA integrity.
RT-qPCR Assay
RNA samples with a ratio of OD 260/280 close to 2.0 were subjected to reverse transcriptions to synthesize cDNA samples. The cDNA samples were subjected to qPCRs to measure the expression of MAFG-AS1, DICER and Drosha with 18S rRNA as the internal control. All qPCRs reactions were prepared using SYBR Green Master Mix (Bio-Rad).
The expression levels of mature miR-34a and miR-34a precursor were determined using Genecopoeia All-in-One™ miRNA qRT-PCR Reagent Kit. Sequence-specific primers were used to determine the expression of miR-34a precursor. To determine the expression of mature miR-34a, all miRNAs were added with poly (A), followed by reverse transcriptions and qPCRs, which were performed using poly (T) reverse primer and sequence-specific forward primer. U6 was used as the internal control for the expression levels of miR-34a. Primer sequences were as follows: 5ʹ-CGAAGATCTCCTCACCTCCC-3ʹ (forward) and 5ʹ-TTAAAGCCGGTCGTGGAGAT-3ʹ (reverse) for MAFG-AS1; 5ʹ-CTACCACATCCAAGGAAGCA-3ʹ (forward) and 5ʹ-TTTTTCGTCACTACCTCCCCG-3ʹ (reverse) for human 18S rRNA; 5ʹ-GGCCAGCTGTGAGTGTTTC-3ʹ (forward) and 5ʹ-GGGCCCCACAACGTGCAGC-3ʹ (reverse) for miR-34a precursor; 5ʹ-TGGCAGTGTCTTAGCTGG-3ʹ (forward) and poly (A) (reverse) for mature miR-34a. U6 primers were from the kit.
5ʹ-GGTCTCAGTTTGGTGGTTC-3ʹ (forward) and 5ʹ-GTAATGCACATTCACCAAAGTCA-3ʹ (reverse) for DICER; 5ʹ-CTAGTTTTCCTGCAGACAATGCA-3ʹ (forward) and 5ʹ-GTAATGCACATTCACCAAAGTCA-3ʹ (reverse) for Drosha. Each experiment was performed in 3 technical replicates, and Ct values of the targeted genes were normalized to internal controls using the 2−ΔΔCq method.
Cell Counting Kit 8 (CCK-8) Assay
U87 and U-118 cells were subjected to cell proliferation assay using a CCK-8 kit (Sigma-Aldrich). A 96-well cell culture plate was filled with a fresh medium containing 4000 cells per well. Cells were cultivated at 37 °C. To determine the cell proliferation, the OD value at 450 nm of each well was determined every 24 h for 4 times. At 2 h before the measurement, CCK-8 solution was added to 10%.
Colony Formation Assay
U87 and U-118 cells were harvested at 48 h post-transfections placed on a cell culture plate. Three replicate wells were set for each experiment. Cell culture was performed for 6 d, followed by washing with PBS. After staining with crystal violet, positive colonies were observed under a light microscope.
Scratch Test
Cell scratch experiment was conducted to determine the effect of MAFG-AS1 on the wound healing process. Briefly, cells were inoculated into a 6-well plate with 1×106 cells per well and cultured for 24 h. Cells were gently scratched with a pipette tip across the center of the well, followed by washing with PBS for 3 times. Then, the cell wound healing process was observed with an inverted fluorescence microscope (Nikon, Japan).
Flow Cytometry
Flow cytometry was performed to determine cell apoptosis using Annexin V-FITC/PI Apoptosis Detection Kit (Invitrogen). Transfected cells were harvested and resuspended in Binding Buffer (1×), followed by an incubation with Annexin VPITC and PI at 37 °C for 30 min in the dark. Cells were inoculated into a 6-well plate with a cell density of 1×106 cells per well. After adherent growth for 24 h, cells were digested by trypsin and the fine cells in the supernatant were collected, washed with PBS for 3 times, fixed with 70% ethanol, and each cell sample was resuspended and added with 500 L pre-mixed PI dye. The cells were resuspended and incubated in the dark for 30 min. The apoptotic rate and G0/G1 cell were analyzed by FACScan flow cytometer (Becton Dickinson) using the CELL QUEST software.
Establishment of Subcutaneous Tumor Animal Model
Nude mice were raised in an SPF environment for 1 week and inoculated with cells. U87 and U-118 cells were harvested and the concentration was adjusted to 1.0 × 107/mL. The cell suspension with the adjusted concentration was injected into the subcutaneous tissue of nude mice with 0.2 mL cell suspension. The animals’ procedures of this study were approved by the Ethics Committee of Zibo Central Hospital.
Western Blot
Total proteins of the cells and tissues were extracted and electrophoretic separated by SDS-PAGE gel, followed by transferring to PVDF membrane. After blocking with BSA at room temperature for 2 h, the membrane was incubated with GAPDH, BCL-1 and caspase-3 primary antibodies at 4 °C with shaking overnight. Secondary antibodies were then added and incubated at room temperature for 2 h, followed by cleaning with TBST for 3 times. Luminescent solution was used for exposure. GAPDH was used as an internal reference.
Statistical Analysis
Gene expression levels in paired tissues were expressed as the mean values of 3 technical replicates and were compared by paired t test. Mean± SD values were used to express data of 3 independent replicates and were compared by ANOVA Tukey’s test. Correlations were analyzed using linear regression. P < 0.05 was considered to be statistically significant.
Results
The Expression of MAFG-AS1, miR-34a Precursor and Mature miR-34a Were Altered in GBM
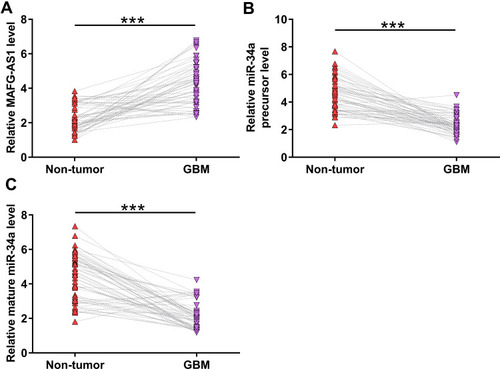
The analysis of TCGA dataset revealed that MAFG-AS1 was upregulated in GBM tissues compared to that in non-tumor tissues (3.24 vs, 1.49). The expression levels of MAFG-AS1 in paired tissue samples from 56 GBM patients were determined by RT-qPCR. Compared to non-tumor tissues, GBM tissues exhibited significantly higher expression levels of MAFG-AS1 (, p < 0.001). The expression levels of miR-34a and miR-34a precursor in paired tissues were also determined by RT-qPCR. Compared to non-tumor tissues, GBM tissues exhibited significantly downregulated expression of miR-34a precursor (, p < 0.001) and mature miR-34a (, p < 0.001).
Figure 1 The expression of MAFG-AS1, miR-34a precursor and mature miR-34a were altered in GBM. The expression levels of MAFG-AS1 (A), miR-34a precursor (B) and mature miR-34a (C) in paired tissue samples from 56 GBM patients were determined by RT-qPCR. Gene expression levels in paired tissues were expressed as average values of 3 technical replicates. ***p < 0.001.
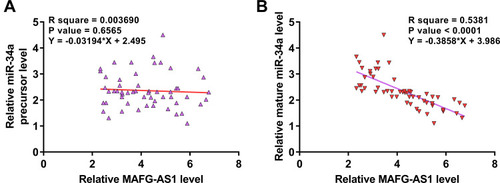
MAFG-AS1 Was Inversely Correlated with Mature miR-34a Across GBM Tissues
The correlations between MAFG-AS1 and miR-34a precursor or mature miR-34a across GBM tissues were analyzed by linear regression. It was observed that MAFG-AS1 and miR-34a precursor was not closely correlated with each other (). In contrast, MAFG-AS1 and mature miR-34a were inversely and significantly correlated across GBM tissues (). The correlation between key genes for miRNA maturation (Dicer and Drosha) and tumor diseases has been reported.Citation16 We also demonstrated that MAFG-AS1 promoted the expression levels of DICER and Drosha in U87 and U-188 cells (Supplemental Figure 1).
Overexpression of MAFG-AS1 Resulted in Downregulation of Mature miR-34a, but Not miR-34a Precursor in GBM Cells
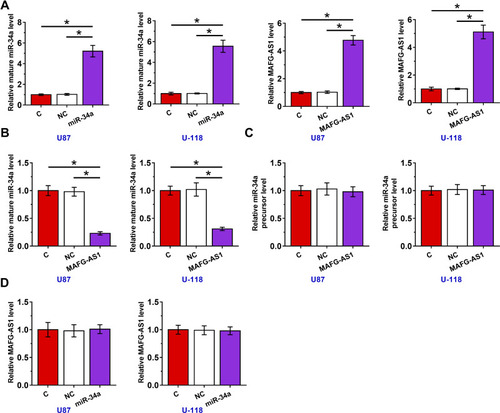
U87 and U-118 cells were transfected with MAFG-AS1 expression vector or miR-34a mimic to explore the interaction between MAFG-AS1 and miR-34a. Transfections were confirmed at 48 h post-transfection by RT-qPCR (, p < 0.05). It was observed that the overexpression of MAFG-AS1 resulted in the downregulation of mature miR-34a (, p < 0.05), but not miR-34a precursor (). Moreover, the overexpression of miR-34a did not affect the expression of MAFG-AS1 (). Therefore, MAFG-AS1 may affect the maturation of miR-34a.
Figure 3 Overexpression of MAFG-AS1 resulted in downregulation of mature miR-34a, but not miR-34a precursor in GBM cells. U87 and U-118 cells were transfected with MAFG-AS1 expression vector or miR-34a mimic. Transfections were confirmed at 48 h post-transfection by RT-qPCR (A), the effects of overexpression of MAFG-AS1 on mature miR-34a (B) and miR-34a precursor (C), and the effects of overexpression of miR-34a on the expression of MAFG-AS1 (D) were also explored by RT-qPCR. Mean± SD values were used to express data of 3 independent replicates. *p < 0.05.
Overexpression of MAFG-AS1 Promoted the Proliferation of GBM Cells Through miR-34a
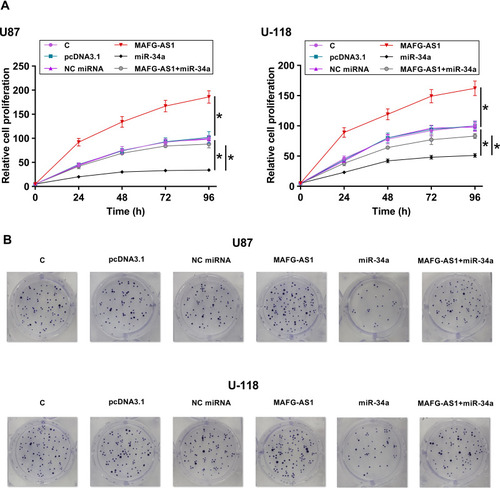
CCK-8 assay was performed to explore the roles of MAFG-AS1 and miR-34a in regulating the proliferation of U87 and U-118 cells. It was observed that overexpression of MAFG-AS1 resulted in increased cell proliferation, while overexpression of miR-34a reduced cell proliferation. Moreover, the overexpression of MAFG-AS1 reduced the inhibitory effects of miR-34a on cell proliferation (, p < 0.05). Colony formation assay was performed to confirm the role of MAFG-AS1 and miR-34a in regulating cell proliferation. As shown in , overexpression of MAFG-AS1 increased colony formation, while overexpression of miR-34a decreased colony formation. Moreover, the overexpression of MAFG-AS1 reduced the inhibitory effects of miR-34a on colony formation. Cell cycle, apoptosis, scratch test and colonies formation experiments were also conducted and it showed that overexpression of MAFG-AS1 just increased cell proliferation but did not affect cell cycle, apoptosis and migration (Supplemental Figures 2 and 3).
Figure 4 Overexpression of MAFG-AS1 promoted the proliferation of GBM cells through miR-34a. CCK-8 assay (A) and colony formation assay (B) were performed to analyze the roles of MAFG-AS1 and miR-34a in regulating the proliferation of U87 and U-118 cells. Mean ± SD values were used to express data of 3 independent replicates. *p < 0.05.
Overexpression of MAFG-AS1 Promoted the Progression of GBM in vivo by Promoting the Proliferation of GBM Cells Through miR-34a
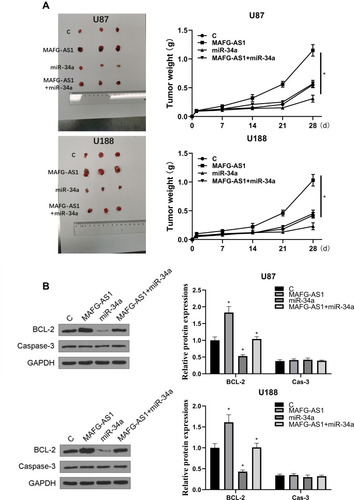
Finally, the role of MAFG-AS1 was explored in vivo, and it showed that overexpression of MAFG-AS1 promoted GBM progression (). In addition, the overexpression of MAFG-AS1 could significantly promote tumor weight (). We also found that overexpression of miR-34a could significantly reverse the effect of overexpression of MAFG-AS1 (p < 0.05). Compared with NC group, the expression levels of bcl-2 protein in the miR-34a group were significantly reduced (p < 0.05), and overexpression of MAFG-AS1 reversed miR-34a-induced inhibition on the expression levels of bcl-2 (). Moreover, we found that the expression of caspase-3 was not affected by MAFG-AS1. Therefore, we speculated that overexpression of MAFG-AS1 promoted the progression of GBM in vivo by promoting the proliferation of GBM cells through miR-34a.
Figure 5 Overexpression of MAFG-AS1 promoted the progression of GBM in vivo. Transfected GBM cells were injected subcutaneously into 8-weeks old nude mice. (A) Representative images of three groups of subcutaneous tumors and comparison of tumor weight in nude mice. (B) The expression levels of bcl-2 and cas-3 proteins were detected by Western blot. N = 5, *p < 0.05.
Discussion
This study mainly investigated the interaction between MAFG-AS1 and miR-34a in GBM. It was observed that MAFG-AS1 was upregulated in GBM and it might affect the maturation of miR-34a to affect cell proliferation.
The oncogenic functions of MAFG-AS1 have been characterized in several types of cancer.Citation13,Citation14 It was reported that MAFG-AS1 is upregulated in colorectal cancer and can sponge miR-147b to upregulate NDUFA4, thereby promoting the progression of cancer.Citation13 In another study, MAFG-AS1 was reported to be upregulated in lung cancer and might sponge miR-744-5p to increase the expression levels of MAFG, thereby promoting the proliferation of cancer cells.Citation14 This study is the first to report the upregulation of MAFG-AS1 in GBM. Moreover, increased proliferation of two GBM cell lines was observed after overexpression of MAFG-AS1. Therefore, MAFG-AS1 may play an oncogenic role in GBM by promoting cancer cell proliferation. We also investigated the effect of overexpression of MAFG-AS1 on cell cycle, apoptosis and migration and verified that MAFG-AS1 just increased cell proliferation.
MiR-34a plays tumor suppressive role in several types of cancer, including glioma.Citation17,Citation18 MiR-34a is downregulated in glioma and targets PD-L1 to promote cancer cell proliferation and inhibit cell apoptosis.Citation17,Citation18 Moreover, the overexpression of miR-34a suppresses the development of chemoresistance in glioma cells.Citation17 In the present study, we observed the downregulation of both miR-34a precursor and mature miR-34a in GBM, as well as the inhibitory effects of miR-34a on cancer cell proliferation.
To form mature miRNAs, pri-miRNA and pre-miRNA (precursor miRNA) are sequentially formed.Citation19 In this study, we observed that overexpression of MAFG-AS1 resulted in downregulation of mature miR-34a, but not miR-34a precursor in GBM cells. Therefore, MAFG-AS1 may regulate the maturation of miR-34a from miRNA precursor to mature miRNAs. Although we did not include in vivo animal model experiment, our results in GBM tissues showed that MAFG-AS1 was inversely correlated with mature miR-34a, but not miR-34a precursor. Therefore, MAFG-AS1 may affect the maturation of miR-34a in GBM cells. It is known that RNase III enzyme Dicer is known to process miRNA precursors to miRNAs.Citation19 However, Dicer affects general miRNAs but not specific miRNAs. We observed that MAFG-AS1 only affected the maturation of miR-34a, but not other miRNAs. Therefore, MAFG-AS1 is unlikely to affect Dicer. The transportation of premature miRNA out of the nucleus to the cytoplasm is required for the formation of mature miRNAs. Therefore, MAFG-AS1 may affect the movement of premature miR-34a. Our future studies will explore this possibility. In addition, our in vivo experience further demonstrated that overexpression of MAFG-AS1 promoted the progression of GBM by promoting the proliferation of GBM cells through miR-34a.
Conclusion
In conclusion, MAFG-AS1 is overexpressed in GBM and may suppress the maturation of miR-34a to promote GBM cell proliferation.
Disclosure
The authors report no conflicts of interest in this work.
References
- Shabihkhani M, Telesca D, Movassaghi M, et al. Incidence, survival, pathology, and genetics of adult Latino Americans with glioblastoma. J Neurooncol. 2017;132(2):351–358. doi:10.1007/s11060-017-2377-028161760
- Hibler E. The glioblastoma patient mortality rate and immunosuppression coincidently increase during advanced age. Front Pharmacol. 2019;10:10. doi:10.3389/fphar.2019.0020030733675
- Brown TJ, Brennan MC, Li M, et al. Association of the extent of resection with survival in glioblastoma: a systematic review and meta-analysis. JAMA Oncol. 2016;2(11):1460–1469. doi:10.1001/jamaoncol.2016.137327310651
- Davis ME. Glioblastoma: overview of disease and treatment. Clin J Oncol Nurs. 2016;20(5 Suppl):S2–8. doi:10.1188/16.CJON.S1.2-8
- Delgado-Lopez PD, Corrales-Garcia EM. Survival in glioblastoma: a review on the impact of treatment modalities. Clin Transl Oncol. 2016;18(11):1062–1071. doi:10.1007/s12094-016-1497-x26960561
- Batash R, Asna N, Schaffer P, Francis N, Schaffer M. Glioblastoma multiforme, diagnosis and treatment; recent literature review. Curr Med Chem. 2017;24(27):3002–3009. doi:10.2174/092986732466617051612320628521700
- Juratli TA, Qin N, Cahill DP, Filbin MG. Molecular pathogenesis and therapeutic implications in pediatric high-grade gliomas. Pharmacol Ther. 2018;182:70–79. doi:10.1016/j.pharmthera.2017.08.00628830841
- Balca-Silva J, Matias D, Carmo AD, Sarmento-Ribeiro AB, Lopes MC, Moura-Neto V. Cellular and molecular mechanisms of glioblastoma malignancy: implications in resistance and therapeutic strategies. Semin Cancer Biol. 2019;58:130–141. doi:10.1016/j.semcancer.2018.09.00730266571
- Lieberman F. Glioblastoma update: molecular biology, diagnosis, treatment, response assessment, and translational clinical trials. F1000Res. 2017;6:1892. doi:10.12688/f1000research.11493.129263783
- Touat M, Idbaih A, Sanson M, Ligon KL. Glioblastoma targeted therapy: updated approaches from recent biological insights. Ann Oncol. 2017;28(7):1457–1472. doi:10.1093/annonc/mdx10628863449
- Ferreira HJ, Esteller M. Non-coding RNAs, epigenetics, and cancer: tying it all together. Cancer Metastasis Rev. 2018;37(1):55–73. doi:10.1007/s10555-017-9715-829374363
- Arun G, Diermeier SD, Spector DL. Therapeutic targeting of long non-coding RNAs in cancer. Trends Mol Med. 2018;24(3):257–277. doi:10.1016/j.molmed.2018.01.00129449148
- Cui S, Yang X, Zhang L, Zhao Y, Yan W. LncRNA MAFG-AS1 promotes the progression of colorectal cancer by sponging miR-147b and activation of NDUFA4. Biochem Biophys Res Commun. 2018;506(1):251–258. doi:10.1016/j.bbrc.2018.10.11230348529
- Sui Y, Lin G, Zheng Y, Huang W. LncRNA MAFG-AS1 boosts the proliferation of lung adenocarcinoma cells via regulating miR-744-5p/MAFG axis. Eur J Pharmacol. 2019;859:172465. doi:10.1016/j.ejphar.2019.17246531211984
- Slabakova E, Culig Z, Remsik J, Soucek K. Alternative mechanisms of miR-34a regulation in cancer. Cell Death Dis. 2017;8(10):e3100. doi:10.1038/cddis.2017.49529022903
- Annese T, Tamma R, De Giorgis M, Ribatti D. microRNAs biogenesis, functions and role in tumor angiogenesis. Front Oncol. 2020;10:581007. doi:10.3389/fonc.2020.58100733330058
- Wang Y, Wang L. miR-34a attenuates glioma cells progression and chemoresistance via targeting PD-L1. Biotechnol Lett. 2017;39(10):1485–1492. doi:10.1007/s10529-017-2397-z28721584
- Li Q, Wang C, Cai L, et al. miR34a derived from mesenchymal stem cells stimulates senescence in glioma cells by inducing DNA damage. Mol Med Rep. 2019;19(3):1849–1857. doi:10.3892/mmr.2018.980030592284
- Achkar NP, Cambiagno DA, Manavella PA. miRNA biogenesis: a dynamic pathway. Trends Plant Sci. 2016;21(12):1034–1044. doi:10.1016/j.tplants.2016.09.00327793495