Abstract
Choroideremia is a rare X-linked recessive inherited disorder that causes chorioretinal dystrophy leading to visual impairment in its early stages which finally causes total blindness in the affected person. It is caused due to mutations in the CHM gene. In this study, we have recruited a pedigree with choroideremia and detected a nonsense variant (c.C799T:p.R267X) in CHM of the proband (I:1). Different primer sets for amplification refractory mutation system (ARMS) were designed and PCR conditions were optimized. Then, we evaluated the sequence variant in the patient, carrier, and a fetus by using ARMS technique to identify if they inherited the pathogenic gene from parental generation; we used amniotic fluid DNA for the diagnosis of the gene in the fetus. The primer pairs, WT2+C and MT+C, amplified high specific products in different DNAs which were verified by Sanger sequencing. Based on our results, ARMS technique is fast, accurate, and reliable prenatal gene diagnostic tool to assess CHM variants. Taken together, our study indicates that ARMS technique can be used as a potential molecular tool in the diagnosis of prenatal mutation for choroideremia as well as other genetic diseases in undeveloped and developing countries, where there might be shortage of medical resources and supplies.
Introduction
Choroideremia (CHM, OMIM: 303100) is a rare X-linked recessive inherited disorder that causes chorioretinal dystrophy with typical clinical features such as progressive degeneration of the choriocapillaris, retinal pigment epithelium (RPE), and photoreceptors.Citation1 Its prevalence ranges from 1:50,000 to 1:100,000.Citation2 Male patients develop night blindness in their childhood and gradually progress to loss of peripheral vision, and finally, some of them will reach complete peripheral blindness at their middle age with only tunnel vision left behind. Female carriers are usually asymptomatic, as to the heterozygous gene, some of them occasionally show mild symptoms as they reach middle age.Citation3,Citation4 Thus, with the progression of the disease, central vision is typically maintained in men as the fovea is spared.
CHM is caused by mutations in the CHM gene, which is mapped on chromosome Xq21.2-21.3 in 186,382 bp.Citation5 CHM mRNA comprises 15 exons, and it is expressed in many types of tissues including choroid, retinal photoreceptor, RPE, and lymphocyte tissues.Citation6 Ran et al reported that there are more than 100 different pathogenic CHM variants in the RetinoGenetics.Citation7 These variants demonstrate deletions, translocations, missense mutations, nonsense mutations, and splice-site mutationsCitation6,Citation8,Citation9 and have been identified in the 15-exon gene. The CHM gene encodes for Rab escort protein-1 (REP1) which comprises 653 amino acids.Citation10 REP1 plays a significant role in the intracellular vesicular trafficking process as well as in the post-translational isoprenyl modification of Rab proteins.Citation11,Citation12
Nowadays, varieties of molecular techniques have been invented to detect genetic mutations, for example, Sanger sequencing, but this technique cannot rapidly screen large numbers of mutations in samples. Fortunately, this problem can be solved by massive next-generation sequencing (NGS),Citation13 but both Sanger sequencing and NGS are expensive. The amplification refractory mutation system (ARMS), also termed allele-specific PCR, is a very simple method for detecting any known mutations involving single base changes or small deletions, which is based on the use of sequence-specific PCR primers that allow amplification of test DNA only when the target allele is contained within the sample. ARMS technique demands only PCR amplification and gel electrophoresis of the amplicons.
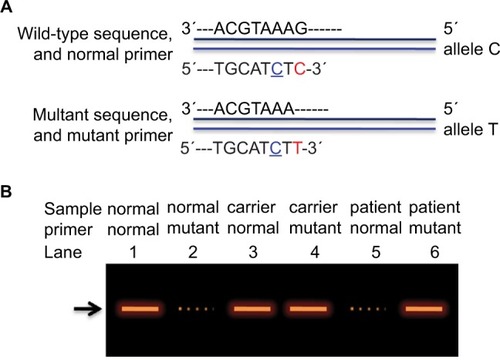
In ARMS technique, one PCR comprises one allele-specific oligonucleotide primer at 5′-end and a common primer at 3′-end. If the presence of an amplified mutant is detected by agarose gel electrophoresis, it suggests that the target sequence contains the mutant allele. Similarly, if the result displays an absence of the amplified mutant, it indicates the presence of the normal DNA sequence on that specific point.
In the same way, a normal primer at 5′-end together with a common primer at 3′-end was used in another PCR. If normal amplified product is present, it reveals the existence of a natural DNA sequence, whereas if normal amplified product is absent, then it reveals the presence of a mutant allele.Citation14–Citation16 If there are both normal and mutant amplified products, then it reveals heterozygous gene variant by the existence of both normal DNA sequence and mutant DNA sequence within an individual.
In this study, evaluation of the ARMS technique for prenatal gene diagnosis of CHM with CHM gene point mutation was performed. This is a cheap, fast, accurate, and reliable pre- and postnatal diagnostic technique which provides the possibility to use it in the identification of various X-linked recessive gene mutations in western China and other undeveloped or developing countries.
Materials and methods
Clinical diagnosis and sample collection
A Chinese proband with CHM was diagnosed previously ().Citation17 DNA templates from the proband and his family of peripheral leukocytes and from the amniotic fluid of fetus at 17 weeks of pregnancy, were isolated using a previously described method.Citation18,Citation19 The quality of DNA was measured by a NanoDrop-2000-spectrophotometer. High quality intact genomic DNA with an optical density ratio of 260/280~1.8 and 260/230>1.5 were used for further analysis. The study was approved by the ethical committee of Southwest Medical University according to the Helsinki Declaration (1983 Revision). Pregnant women signed informed consent forms for approval of the prenatal diagnosis.
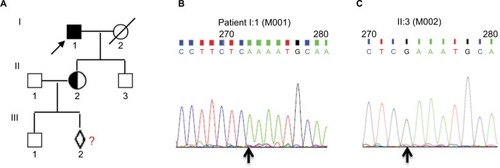
Figure 1 The pedigree of choroideremia and CHM gene mutation.
Notes: (A) Pedigree of choroideremia. (B) The proband (I:1) with CHM nonsense variant (CHM:NM_000390:c.C799T:p.R267X) is indicated by an arrow as a filled square, exhibiting a 799 C→T mutation at the first nucleotide position of codon 267 (R267X) in the exon 6 of the CHM gene locus (cga-tga). (C) The proband’s son (II:3) harbors normal DNA sequences. The carrier (II:2) is a female of proband’s daughter and is pregnant with a fetus (III:2) who underwent prenatal diagnosis to demonstrate whether the baby inherited a normal X-chromosome. All of the Sanger sequencing results in this study used reverse complementary sequencing of the gene. The black arrow shows the position of the point mutation. The primer pair of CHM-L and CHM-R was used for PCR amplification and primer CHM-R for Sanger sequencing.
Design of ARMS’ primers
We designed the 5′ ARMS’ primers with wild and mutant types with or without one base mismatched at the 3′-end following the nonsense mutation (c.C799T:p.R267X) of the CHM gene, respectively.Citation5 In addition, a common primer located at the 3′ terminus of CHM gene was designed. presents the information of primers.
Table 1 Amplification refractory mutation system and their combination
ARMS-PCR amplification
PCR was performed in a 10 µL reaction volume for DNA samples M001, M002, M003, M004, M050, and M051 that comprised 1 µL DNA template (15 ng), 1 µL 3.3 µM primers, 5 µL 2X PCR Taq Master Mix (TianGen Biotech Co. Ltd., Beijing, People’s Republic of China), and 3 µL double-distilled water. PCR cycles were performed in a programmable thermal cycler (Applied Biosystems Veriti® 96-Well Thermal Cycler, Life Technologies, Foster City, CA, USA).Citation20 PCR regimen was as follows: initial denaturation at 95°C for 60 s, followed by 33 cycles for 30 s at 94°C, 30 s at 61°C, 40 s at 72°C, and then a final extension for 5 min at 72°C, and finally the PCR products were maintained at 4°C in the end. In case of M151, which was amniotic fluid DNA sample with a lower concentration, we increased the total cycles to 35, and the rest of the conditions were same as the others.
Agarose gel electrophoresis
PCR products were resolved on a 1% agarose gel with 1× Tris–acetate-ethylenediaminetetraacetic acid (TAE) buffer, and then subjected to electrophoresis at 120 V for 30 min. The agarose gel was stained with 0.5 µg/mL ethidium bromide and photographed by ChemiDocXR (Bio-Rad, Hercules, CA, USA).Citation19,Citation20 DNA Marker (DL2000) for electrophoresis was purchased from TaKaRa Biotechnology Co. Ltd., (Dalian, People’s Republic of China).
Mutation validation
To validate the ARMS-PCR results, the variant was sequenced by Sanger method on an ABI 3500 DX Genetic Analyzer (ABI, Thermo Fisher Scientific, Waltham, MA, USA) with primer CHM-R by amplification using the primer pair CHM-L and CHM-R (length of the product is 637 bp) ().
Table 2 Sequences of PCR primers and PCR product size (bp)
Results
Designing of ARMS amplification primers
In this study, we recruited a Chinese pedigree with CHM to perform the analysis using ARMS technique(). The results by Sanger sequencing with a variant (c.C799T:p. R267X) in the exon 6 of the CHM gene (GenBank accession number: NM_000390) on the X chromosome in the proband (I:1) and his son (II:3) with normal phenotype are shown in , respectively. As seen in , I:1 (M001) is the proband of pedigree (indicated by an arrow) with CHM gene nonsense mutation from C→T (), whereas proband’s son (II:3) was normal both in phenotype and genotype with no mutation (). His daughter was a carrier with an unborn fetus which was tested for inheritance of the mutant gene from the mother.
In order to establish and optimize the ARMS condition, two wild-type primer sets were designed: wild-type primer (CHM-WT) and point mutation wild-type primer (CHM-WT2) which matched with wild-type sequence of CHM, sharing the 3′ common primer (CHM-C3) as primer pairs (WT+C) and (WT2+C); two mutant primer sets were designed: mutant-type primer (CHM-MT) and point mutation for mutant-type primer (CHM-MT2) which matched with mutant-type sequence of CHM c.C799T, sharing the 3′ common primer (CHM-C3) as primer pairs (MT+C) and (MT2+C) (). The length of the PCR product was 490 bp. The artificial mismatch at the 3rd position of 3′-end in both CHM-WT2 and CHM-MT2 were designed to enhance PCR amplification.
The combination of primer pair (MT+C), which matched with the mutant CHM gene c.C799T, was used to detect the mutant, and the combination of primer pair (MT2+C) with one base mismatch was used to enhance the specificity for detecting the mutant CHM gene c.C799T.
Optimization of ARMS amplification
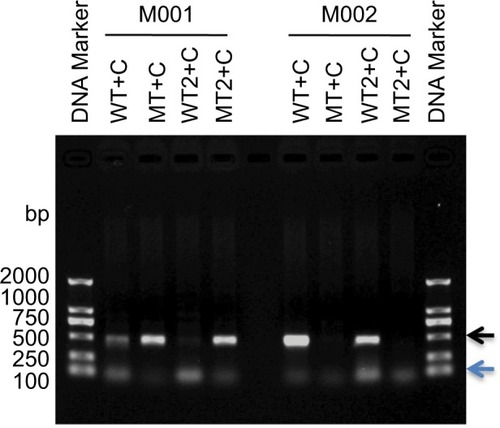
PCR amplification was performed to optimize ARMS condition by the aforementioned four primer pairs using DNA templates M001 (I:1) and M002 (II:3). As seen in , all four primer pairs amplified specific bands with right size, except the primer pair (WT+C), with no mismatch in wild-type sequence and amplified nonspecific band in M001 (, lane labeled “WT+C”). As we know, M001 is a male patient with T mutant allelic CHM gene (). The primer pair (WT2+C), with one base mismatch in wild-type sequence, amplified specific band only in normal person (M002) but not in patient (M001) (, lane labeled “WT2+C”). Both primer pairs (MT+C and MT2+C) amplified specific bands in the proband patient M001 with the mutant allele T (, lanes labeled “MT+C” and “MT2+C”) but amplified nonspecific band in M002 (, lanes labeled “MT+C” and “MT2+C”), indicating that there is no difference between these two primer pairs. Thus, we optimized the ARMS condition by selecting primer pair (WT2+C) of wild-type and primer pair (MT+C) of mutant-type.
Figure 2 Optimization of ARMS condition.
Abbreviations: ARMS, amplification refractory mutation system; WT, wild-type primer; MT, mutant primer; C, common primer.
Evaluation of ARMS for gene diagnosis of CHM
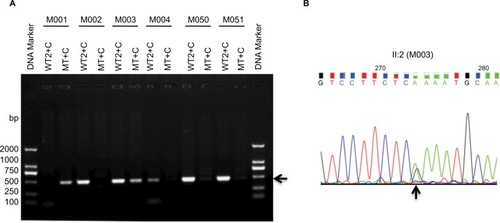
To complete the diagnosis of CHM, we used the primer pairs WT2+C and MT+C to amplify different DNA samples from all the family members and a normal male control (sample M051). As shown in , the patient’s DNA (M001) amplified the mutant allele but did not amplify the wild-type allele, which is consistent with our previous results (shown in ); however, carrier daughter’s DNA (M003) amplified both alleles. Sanger sequencing demonstrated that the daughter is heterozygous with one allele point mutation of CHM c.C799T (). All the normal individuals’ DNA (M002, M004, M050, including control M051) amplified only wild-type allele. Validation of the putative mutation was performed by Sanger sequencing ( and data not shown). The carrier M003 had a heterozygous variant from C→T and the mutant gene was same as her father, indicating that the mutant CHM gene is inherited from her father ( and ).
Figure 3 ARMS amplification by primers CHM-WT2 (WT2) and CHM-Common, and primers CHM-MT and CHM-Common.
Abbreviations: ARMS, amplification refractory mutation system; CHM, choroideremia; WT, wild-type primer; MT, mutant primer; C, common primer.
Evaluation of ARMS for quick and accurate prenatal gene diagnosis of CHM
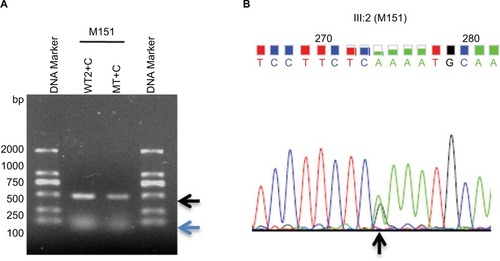
In this pedigree, M151 was the fetal baby of the carrier M003, and we amplified genetic material of M151 from amniotic fluid DNA using the same primers (WT2+C and MT+C) as for other family members used above. To evaluate ARMS for quick and accurate prenatal gene diagnosis of CHM, ARMS-PCR was performed and results are shown in , which indicates that the fetus (M151) amplified both mutant as well as the wild-type allele, similar to his/her mother (M003). Sanger sequencing result of M151 further revealed a heterozygous mutation from C→T (), confirming our ARMS results. PCR for SRY gene demonstrated that there was no Y chromosome, indicating that M151 was a female fetus which was demonstrated after birth. Thus, ARMS was performed very quickly, and its results were found to be very reliable and accurate for CHM gene prenatal diagnosis.
Figure 4 ARMS amplification from fetal amniotic fluid DNA (III:2) by primers CHM-WT2 (WT2) and Common primer CHM-C3, and primers CHM-MT and CHM-Common.
Abbreviations: ARMS, amplification refractory mutation system; CHM, choroideremia; WT, wild-type primer; MT, mutant primer; C, common primer.
Therefore, the female fetus M151 has the same genotype as her mother by inheriting the pathogenic mutant gene from her grandfather passing on to her mother and then to herself. With genetic counseling, the mother decided to give birth to the baby, showing no symptoms of CHM with more than 3 years’ of follow-up information.
Discussion
As a rare X-linked recessive inherited disorder, CHM causes chorioretinal dystrophy with typical clinical features such as progressive degeneration of the choriocapillaris, RPE, and photoreceptors.Citation2,Citation6,Citation21 Different kinds of variants/mutations have been identified in CHM gene,Citation6,Citation8,Citation9 and also some methods have been applied to detect these mutations.Citation4,Citation21 Gene therapy and retinal transplantation of CHM are considered possible treatments and are in clinical trials from last few years. Thirteen different human retinal gene therapy clinical trials have been performed. The majority of these studies used adeno-associated virus type 2 (AAV2) as the gene transfer vector.Citation22,Citation23 To date, the safety assessment of gene therapy mediated by AAV2 in CHM patients has been finished already.Citation21 Another clinical study demonstrated significant progress and suggests that AAV8 might be a candidate vector for CHM gene therapy.Citation24 Therefore, individuals affected with CHM might be cured in the near future. However, gene therapy has still a long way to go in order to become a reality. Thus, prenatal diagnosis is the current used strategy that can prevent genetic disorders.
Mutations in CHM gene are known to lead CHM.Citation25,Citation26 Li et al reported that at least 147 kinds of mutations have been identified in patients with CHM, containing 39 nonsense mutations, 2 missense mutations, 25 splicing mutations, 32 small deletions, 5 small indels, 9 small insertions, 1 gross insertions/duplication, 31 gross deletions, and 3 complex rearrangements, based on HGMD® Professional 2012.4. All the aforementioned kinds of mutations are loss of function mutations except two types.Citation8,Citation27 In another study, they found 6 mutations including 1 nonsense, 2 splicings, and 3 small deletions,Citation28 and all of them are loss of function mutations. Cai et al added two novel mutations to expand the spectrum of CHM-related mutations in two unrelated Chinese families.Citation29 Guo et al discovered one frameshift mutation in a Chinese family.Citation30 In our studies, the mutation was identified by NGS and noninvasive prenatal testing (NIPT) of SRY gene.Citation17
ARMS is a very technique that can detect known mutations involving single base changes or small deletions, by using sequence-specific PCR primers. In this study, we successfully applied this technique to evaluate the value of prenatal gene diagnosis for CHM. To the best of our knowledge, this is the first report on gene and prenatal gene diagnosis of CHM by ARMS. shows the diagram of ARMS strategy for CHM gene diagnosis by introducing one base mismatch T→C at 3rd position from 3′ end of both normal and mutant primers in our study.
Figure 5 A diagram for ARMS strategy to diagnose CHM gene.
Abbreviations: ARMS, amplification refractory mutation system; CHM, choroideremia.
Previously, Fu et al and Moghadam et al successfully used ARMS to diagnose β-thalassemia prenatally in China and Iran, respectively;Citation18,Citation31 Chiu et al applied this method to quantify given mutant mtDNA heteroplasmies,Citation32 whereas Newton et al used ARMS for antenatal diagnosis of cystic fibrosis.Citation33 Recently, Yang et al performed rapid gene diagnosis for retinitis pigmentosa,Citation34 and Singh et al performed prenatal gene diagnosis for sickle cell disease by ARMS.Citation35 More recently, Aquino et al performed ARMS-PCR to detect the most common mutations of the CFTR gene in Peruvian patients with cystic fibrosis.Citation36 Shah et al combined ARMS and DNA sequencing to diagnose β-thalassemia in East-Western Indian population for better management.Citation37 From the above instances, it is noteworthy that ARMS can be used in the pre- and postnatal diagnosis of various genetic diseases.Citation38 Although there are many sophisticated tools applied to clinical diagnosis, it is very expensive for people in western China and some other undeveloped countries. As for known mutations in genes, ARMS provides an optional way to solve this problem with its additional characteristics such as time-efficient, unbiased, sensitive, accurate, rapid, and reliable. In our study, we successfully developed the ARMS technique by using wild-type or mutant-type primers with matched or one-base mismatched to examine the known mutation in the CHM gene. Furthermore, we tried to obtain samples to test how early ARMS method can be applied in clinical settings, and currently, we are isolating cff DNAs from different stages to test whether ARMS work.
Taken together, ARMS could be useful for quick and accurate prenatal gene diagnosis of CHM for CHM gene as well as other genetic diseases in undeveloped and developing countries which are in shortage of medical resources and supplies.
Acknowledgments
This work was supported by the Research Foundation of the Education Department of Sichuan Province (17ZA0427,17ZB0467), the Research Foundation of the Science and Technology Department of Luzhou City (2013LZLY-J10, 2016-S-65(9/9), 2015-S-42(3/4)), and in part by the National Natural Science Foundation of China (Nos. 30371493, 31701087, 81672887, and 81172049).
Disclosure
The authors report no conflicts of interest in this work.
References
- RobertsMFFishmanGARobertsDKRetrospective, longitudinal, and cross sectional study of visual acuity impairment in choroideraemiaBr J Ophthalmol200286665866212034689
- CoussaRGTraboulsiEIChoroideremia: a review of general findings and pathogenesisOphthalmic Genet2012332576522017263
- RudolphGPreisingMKalpadakisPHaritoglouCLangGELorenzBPhenotypic variability in three carriers from a family with choroideremia and a frameshift mutation 1388delCCinsG in the REP-1 geneOphthalmic Genet200324420321414566650
- DimopoulosISRadziwonALaurentCDStMacDonaldIMChoroideremiaCurr Opin Ophthalmol201728541041528520608
- CremersFPvan de PolDJvan KerkhoffLPWieringaBRopersHHCloning of a gene that is rearranged in patients with choroideremiaNature199034762946746772215697
- Van BokhovenHvan den HurkJABogerdLCloning and characterization of the human choroideremia geneHum Mol Genet199437104110467981670
- RanXCaiWJHuangXF‘RetinoGenetics’: a comprehensive mutation database for genes related to inherited retinal degenerationDatabase (Oxford)20142014bau04724939193
- SergeevYVSmaouiNSuiRThe functional effect of pathogenic mutations in Rab escort protein 1Mutat Res20096651445019427510
- ItabashiTWadaYKawamuraMSatoHTamaiMClinical features of Japanese families with a 402delT or a 555–556d elAG mutation in choroideremia geneRetina200424694094515579993
- SeabraMCBrownMSSlaughterCASüdhofTCGoldsteinJLPurification of component A of Rab geranylgeranyl transferase: possible identity with the choroideremia gene productCell1992706104910571525821
- PereiralealJBHumeANSeabraMCPrenylation of Rab GTPases: molecular mechanisms and involvement in genetic diseaseFEBS Lett200149823197200
- GoodyRSRakAAlexandrovKThe structural and mechanistic basis for recycling of Rab proteins between membrane compartmentsCell Mol Life Sci200562151657167015924270
- FuQWangFWangHNext-generation sequencing–based molecular diagnosis of a Chinese patient cohort with autosomal recessive retinitis pigmentosaInvest Ophthalmol Vis Sci20135464158416623661369
- NewtonCRGrahamAHeptinstallLEAnalysis of any point mutation in DNA. The amplification refractory mutation system (ARMS)Nucleic Acids Res1989177250325162785681
- HassanSAhmadRZakariaZZulkafliZWanZADetection of β-globin gene mutations among β-thalassaemia carriers and patients in Malaysia: application of multiplex amplification refractory mutation system–polymerase chain reactionMalays J Med Sci20132011320
- HanafiSHassanRBaharRMultiplex amplification refractory mutation system (MARMS) for the detection of β-globin gene mutations among the transfusion-dependent β-thalassemia Malay patients in Kelantan, Northeast of Peninsular MalaysiaAm J Blood Res201441334025232503
- ZhuLChengJZhouBDiagnosis for choroideremia in a large Chinese pedigree by next-generation sequencing (NGS) and non-invasive prenatal testing (NIPT)Mol Med Rep20171531157116428098911
- FuJJLiLYLiXRLuGXRapid prenatal gene diagnosis for β-thalassemia by amplification refractory mutation system (ARMS)Chin J Obstet Gynecol200035359360
- FuJShort Protocols in Medical Molecular BiologyBeijingChina Medical Science Press2012
- FuJYangLKhanMAMeiZGenetic characterization and authentication of Lonicera japonica Thunb. by using improved RAPD analysisMol Biol Rep201340105993599924057241
- MaclarenREGroppeMBarnardARRetinal gene therapy in patients with choroideremia: initial findings from a phase 1/2 clinical trialLancet201438399231129113724439297
- AmadoDMingozziFHuiDSafety and efficacy of subretinal re-administration of an AAV2 vector in large animal models: implications for studies in humansSci Transl Med201022121ra16
- BennettJAshtariMWellmanJAAV2 gene therapy readministration in three adults with congenital blindnessSci Transl Med20124120120ra15
- BlackAVasireddyVChungDCAdeno-associated virus 8-mediated gene therapy for choroideremia: preclinical studies in in vitro and in vivo modelsJ Gene Med20141656122130
- CremersFPArmstrongSASeabraMCBrownMSGoldsteinJLREP-2, a Rab escort protein encoded by the choroideremia-like geneJ Biol Chem19942693211121178294464
- YingLXialinLLixiaLMolecular analysis of the choroideremia gene related clinical findings in two families with choroideremiaMol Vis2011172772564256922025891
- EspositoGDe FalcoFTintoNComprehensive mutation analysis (20 families) of the choroideremia gene reveals a missense variant that prevents the binding of REP1 with rab geranylgeranyl transferaseHum Mutat201132121460146921905166
- LiSGuanLFangSExome sequencing reveals CHM mutations in six families with atypical choroideremia initially diagnosed as retinitis pigmentosaInt J Mol Med201434257357724913019
- CaiXBHuangXFTongYLuQKJinZBNovel CHM mutations identified in Chinese families with ChoroideremiaSci Rep201663536027739455
- GuoHLiJGaoFLiJWuXLiuQWhole-exome sequencing reveals a novel CHM gene mutation in a family with choroideremia initially diagnosed as retinitis pigmentosaBMC Ophthalmol2015158526216097
- MoghadamMKarimiMDehghaniSJEffectiveness of β-thalassemia prenatal diagnosis in Southern Iran: a cohort studyPrenat Diagn201535121238124226296249
- ChiuRWMurphyMFFidlerCZeeBCWainscoatJSLoYMDetermination of RhD zygosity: comparison of a double amplification refractory mutation system approach and a multiplex real-time quantitative PCR approachClin Chem200147466767211274016
- NewtonCRHeptinstallLESummersCAmplification refractory mutation system for prenatal diagnosis and carrier assessment in cystic fibrosisLancet198928678867914811483
- YangWCZhuLZhouBXEstablishment and rapid detection of a heterozygous missense mutation in the CACNA1F gene by ARMS technique with double-base mismatched primersGenet Mol Res2015143114801148726436388
- SinghPJShrivastavaACShrikhandeAVPrenatal diagnosis of sickle cell disease by the technique of PCRIndian J Hematol Blood Transfus201531223324125825564
- AquinoRProtzelARiveraJFrequency of the most common mutations of the CFTR gene in peruvian patients with cystic fibrosis using the ARMS-PCR techniqueRev Peru Med Exp Salud Pública20173416269 Spanish28538847
- ShahPSShahNDRayHSPMutation analysis of β-thalassemia in East-Western Indian population: a recent molecular approachAppl Clin Genet201710273528546763
- ImaniSChengJShasaltanehMDGenetic identification and molecular modeling characterization reveal a novel PROM1 mutation in Stargardt4-like Macular DystrophyOncotargetIn press2017
