The mother of the yellow offspring ate normal mouse chow during pregnancy, whereas that of the brown offspring ate mouse chow supplemented with methyl donors. Coat color and size differences are due to variations in the epigenome; an example of nature via nurture.

A colleague of mine once took a small vial out of his pocket, and placed it on the table in front of him before he sat down for a conference. As the conference drew to a close, he carefully picked up the vial, and announced, “This is a vial of DNA, and in the past two days, it hasn‘t done a damn thing!”. Like a computer, DNA is impotent without its software, the epigenome, telling it when, where and how to work.
‘Epigenome‘ simply means ‘above the genome‘, and it refers to a heritable chemical marking system, comprised of DNA methylation, histone marks and noncoding RNA species, that alters gene expression without relying on a disturbance in the underlying genetic code. In contrast to the genome, the epigenome is different in every cell type in the body. The ability to modify epigenetic programming is indeed critical for cellular differentiation and the development of complex living organisms from a single set of genomic instructions. The epigenome is also more malleable than the genome, producing a conduit by which the environment can readily and rapidly affect change in gene function. Such variation can be acted upon by natural selection, ultimately resulting in adaptive responses; however, it can also increase susceptibility to diseases.
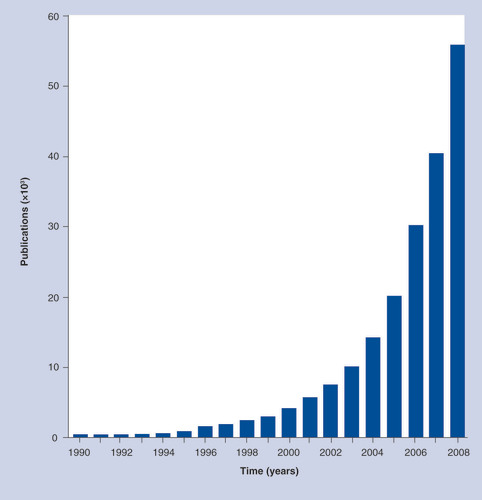
The developmental biologist Conrad Waddington coined the term ‘epigenetics‘ in the 1940s Citation[1]. Over the next 50 years, the number of papers published on this subject was limited, but the accumulation of epigenetic information is now growing exponentially, doubling almost every 2 years . We are presently in the rapidly accelerating part of the exponential curve, making it difficult to predict with confidence the future of epigenetics research. Nevertheless, it is safe to say that with the creation of faster massive paralleled sequencing platforms, the line-by-line decoding of epigenomic programs will continue at an ever increasing pace in the quest to understand cell differentiation, organismal development, species evolution and how epigenetic deregulation leads to the pathogenesis of human diseases and neurological disorders. To assist in the dissemination of this burgeoning amount of information, Future Medicine, Ltd (London, UK) is publishing the inaugural issue of the journal Epigenomics this month.
It has been known for years that in utero exposure to famine is adversely associated with an offspring‘s risk of disease later in life Citation[2–5]. To determine if alterations in the epigenome are mechanistically involved in this phenomenon, we used the viable yellow agouti (Avy) mouse Citation[3,6]. Metastable epialleles at the Avy locus are epigenetically modified in a variable and reversible manner, such that a distribution of coat colors, ranging from yellow to brown, occurs in genetically identical offspring .
We demonstrated with the use of the Avy mouse that maternal dietary supplementation during pregnancy with methyl donors (i.e., choline, betaine, folic acid and vitamin B12) shifts offspring coat color distribution, decreasing the incidence of a yellow coat color Citation[3,7]. This is associated concomitantly with a reduction in their risk of developing obesity, diabetes and cancer. Furthermore, these phenotypic changes were shown to result from increased DNA methylation of a transposable element upstream of the Agouti gene, rather than from mutation of the gene. Interestingly, genistein, a weak phytoestrogen found in soy products, elicits a similar epigenetic effect at the Avy locus, even though it is not a methyl donor Citation[8]. Methyl donors and genistein can also counteract the CpG hypomethylation caused by bisphenol A (BPA), an endocrine-disrupting agent used to make hard clear plastic and epoxy resins Citation[9]. As Hippocrates asserted over two millennia ago – food is medicine.
Thus, we provided the first experimental evidence that the memory system linking these two disparate time points in life involves epigenetic modifications established soon after fertilization Citation[7]. In so doing, we opened the mechanistic ‘black box‘ of the developmental origins of adult disease susceptibility, and firmly placed the word, epigenetics, in the vernacular of this research field.
Imprinted genes are also regulated epigenetically. Genomic imprinting results in autosomal genes with parent-of-origin dependent, monoallelic expression because of gamete-specific epigenetic marks that silence one allele Citation[10,11]. This unique epigenetic form of gene regulation evolved approximately 180 million years ago in an ancestor common to the Therian subclass of mammals Citation[12–14]. The imposition of a functional haploid state at imprinted loci greatly increases their prominence in disease formation, since a single genetic mutation or lone epigenetic event can alter their function Citation[10,11].
Several syndromes involving uniparental disomy (UPD), such as Beckwith–Wiedemann, Prader–Willi and Angelman syndromes Citation[10,15], as well as myoclonus-dystonia Citation[16], transient neonatal diabetes mellitus Citation[17] and a variety of cancers Citation[18] result from genetic and epigenetic modifications at imprinted loci. Epigenetic dysregulation of imprinting may also explain the variability and parental effects observed in the pathogenesis of autism and schizophrenia Citation[19]; its role in these disorders, however, still remains largely unexplored. Children exposed prenatally to the Dutch famine at the end of World War II have an increased incidence of metabolic disorders, cardiovascular disease and mental disorders in adulthood Citation[2,4,5]. They also have significant alterations in DNA methylation at nonimprinted and imprinted loci decades after birth Citation[20,21].
There are presently approximately 50 genes known to be imprinted in humans, and 80 in mice Citation[22,101]. We recently developed and employed computer-learning algorithms for genome-wide prediction of genes with a high probability of being imprinted in the mouse Citation[23] and human Citation[24]. Not only are humans predicted to have fewer imprinted genes than mice, there is also only a mere 30% overlap between their imprinted gene repertoires. Swine appear to have a greater concordance in imprinted genes with humans Citation[25], even though they reside in a different superordinal clade than mouse and human Citation[26]. Plainly stated, mice are not humans.
Recently, there have been reports questioning the usefulness of the mouse as a model for human diseases Citation[27–30]. In 2004, Clifton Leaf came to the provocative conclusion that although mouse tumors can often be cured, this has not led to therapies as effective in eradicating human cancers Citation[27]. This disturbing species difference in therapeutic efficacy is postulated to have occurred because scientists either forgot or ignored the fact that although a mouse gene may be very similar to a human gene, the rest of the mouse is very different. This is true, in part, because a mouse epigenome differs markedly from that of a human.
Thus, despite the immense popularity and ease of using the mouse to ‘model‘ human diseases, it may not be a suitable choice for studying diseases resulting primarily from the dysregulation of the epigenome, rather than mutation of the genome. This means it is critically important to identify the environmentally labile cis-acting imprint regulatory elements in the human genome – the ‘Imprintome‘. Without this genome-wide epigenetic information, the ability to diagnose, prevent and treat human pathological conditions including autism, cancer and schizophrenia will remain compromised.
We are clearly entering a new era of biological research – one where it is becoming increasingly apparent that humans are indeed the best model for understanding diseases afflicting mankind, as stated so prophetically by the English poet Alexander Pope in the early 18th century: “The proper study of mankind is man”. This important realization, and the change in the research approach to which it points, will require scientists from numerous disciplines to collaborate. This will allow for the enrolling of patients and the collection of biological samples and exposure information needed to tease out the epigenetic alterations that link environmental exposures during susceptible stages of life to disease formation years later. The field of medicine will be revolutionized by this epigenetic perspective of disease formation – potentially shifting our healthcare emphasis from therapy to prevention.
Financial & competing interest disclosure
This work was supported by the NIH grant 5R01-ES008823, DOE DE-FG02–05ER64101, and an Esther B. O‘Keeffe Charitable Foundation Award. The author has no other relevant affiliations or financial involvement with any organization or entity with a financial interest in or financial conflict with the subject matter or materials discussed in the manuscript apart from those disclosed.
No writing assistance was utilized in the production of this manuscript.
Additional information
Funding
Bibliography
- Waddington CH : Organisers and Genes. Cambridge University Press, Cambridge, UK (1940).
- Susser E , NeugebauerR, HoekHW et al.: Schizophrenia after prenatal famine. Further evidence.Arch. Gen. Psychiatry53 , 25–31 (1996).
- Jirtle RL , SkinnerMK: Environmental epigenomics and disease susceptibility.Nat. Rev. Genet.8 , 253–262 (2007).
- Barker DJ : The origins of the developmental origins theory.J. Intern. Med.261 , 412–417 (2007).
- Gluckman PD , HansonMA, BeedleAS: Early life events and their consequences for later disease: a life history and evolutionary perspective.Am. J. Hum. Biol.19 , 1–19 (2007).
- Duhl DM , VrielingH, MillerKA, WolffGL, BarshGS: Neomorphic agouti mutations in obese yellow mice.Nat. Genet.8 , 59–65 (1994).
- Waterland RA , JirtleRL: Transposable elements: targets for early nutritional effects on epigenetic gene regulation.Mol. Cell. Biol.23 , 5293–5300 (2003).
- Dolinoy DC , WeidmanJR, WaterlandRA, JirtleRL: Maternal genistein alters coat color and protects Avy mouse offspring from obesity by modifying the fetal epigenome.Environ. Health Perspect.114 , 567–572 (2006).
- Dolinoy DC , HuangD, JirtleRL: Maternal nutrient supplementation counteracts bisphenol A-induced DNA hypomethylation in early development.Proc. Natl Acad. Sci. USA104 , 13056–13061 (2007).
- Murphy SK , JirtleRL: Imprinting evolution and the price of silence.Bioessays25 , 577–588 (2003).
- Jirtle RL , WeidmanJR: Imprinted and more equal.Am. Sci.95 , 143–149 (2007).
- Killian JK , ByrdJC, JirtleJV et al.: M6P/igf2r imprinting evolution in mammals.Mol. Cell5 , 707–716 (2000).
- Killian JK , NolanCM, StewartN et al.: Monotreme IGF2 expression and ancestral origin of genomic imprinting.J. Exp. Zool.291 , 205–212 (2001).
- Nolan CM , KillianJK, PetitteJN, JirtleRL: Imprint status of M6P/IGF2R and IGF2 in chickens.Dev. Genes Evol.211 , 179–183 (2001).
- Falls JG , PulfordDJ, WylieAA, JirtleRL: Genomic imprinting: implications for human disease.Am. J. Pathol.154 , 635–647 (1999).
- Kinugawa K , VidailhetM, ClotF, ApartisE, GrabliD, RozeE: Myoclonus-dystonia: an update.Mov. Disord.24 , 479–489 (2009).
- Temple IK , ShieldJP: Transient neonatal diabetes, a disorder of imprinting.J. Med. Genet.39 , 872–875 (2002).
- Feinberg AP : An epigenetic approach to cancer etiology.Cancer J.13 , 70–74 (2007).
- Crespi B : Genomic imprinting in the development and evolution of psychotic spectrum conditions.Biol. Rev. Camb. Philos. Soc.83 , 441–493 (2008).
- Heijmans BT , TobiEW, SteinAD et al.: Persistent epigenetic differences associated with prenatal exposure to famine in humans.Proc. Natl Acad. Sci. USA105 , 17046–17049 (2008).
- Tobi EW , LumeyLH, TalensRP et al.: DNA Methylation differences after exposure to prenatal famine are common and timing- and sex-specific.Hum. Mol. Genet. (2009) (Epub ahead of print).
- Morison IM , RamsayJP, SpencerHG: A census of mammalian imprinting.Trends Genet.21 , 457–465 (2005).
- Luedi PP , HarteminkAJ, JirtleRL: Genome-wide prediction of imprinted murine genes.Genome Res.15 , 875–884 (2005).
- Luedi PP , DietrichFS, WeidmanJR, BoskoJM, JirtleRL, HarteminkAJ: Computational and experimental identification of novel human imprinted genes.Genome Res.17 , 1723–1730 (2007).
- Bischoff SR , TsaiS, HardisonN et al.: Characterization of conserved and nonconserved imprinted genes in swine.Biol. Reprod. (2009).
- Murphy WJ , EizirikE, JohnsonWE, ZhangYP, RyderOA, O‘BrienSJ: Molecular phylogenetics and the origins of placental mammals.Nature409 , 614–618 (2001).
- Leaf C : Why are we losing the war on cancer (and how to win it).Fortune149(6) , 76–82, 84–86, 88 (2004).
- Davis MM : A prescription for human immunology.Immunity29 , 835–838 (2008).
- Schnabel J : Neuroscience: standard model.Nature454 , 682–685 (2008).
- Steinman RM , MellmanI: Immunotherapy: bewitched, bothered, and bewildered no more.Science305 , 197–200 (2004).
▪ Website
- Geneimprint website: list of experimentally determined and predicted imprinted genes in mammals www.geneimprint.com/site/genes-by-species