Figures & data
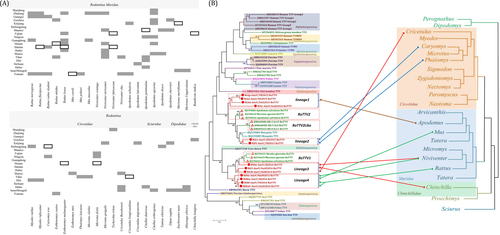
Fig. 1 a Anellovirus-positive pool samples of rodents in China. Blank spaces represent absent samples. The black frames represent anellovirus negativity, and dark shading represents anellovirus positivity in each species and province. b Combined phylogenetic trees for anelloviruses and rodent hosts. The anellovirus phylogenetic tree based on the complete ORF1 protein sequences was constructed using MEGA6 with the mtREV + F + G + I model after alignment with the MUSCLE package, with 1000 bootstrap replicates. The rodent phylogenetic tree based on mitochondrial-cyt b was drawn from genus to family according to previous reportsCitation12. The viruses found in the current study are indicated by red circles (![]()