Figures & data
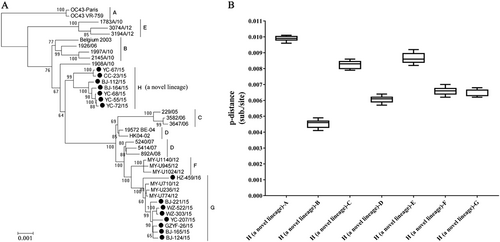
a Phylogenetic tree of the HCoV-OC43 strains based on whole-genome sequences. The tree was constructed by using the maximum likelihood (ML) method with the best-fit general time reversible model with gamma-distributed rate variation across sites and 1000 bootstrap replicates implemented in MEGA 5.03. Bootstrap values over 70% are shown in the nodes. b Estimation of pairwise genetic distances between genotype H and genotypes A–G of HCoV-OC43 strains based on the whole-genome sequences
Demographic and clinical manifestation of HCoV-OC43 positive cases
