Figures & data
Cetacean morbillivirus detection in cetacean samples from the North Sea and the Mediterranean Sea
a Genome coverage of wild-type CeMVs. The scale on the left indicates sequencing depth and genome regions are color-coded according to CeMV gene positions. b Sequencing chromatograms of leader and trailer regions of wild-type CeMV strains DMV-16A and PMV-2990 using RACE
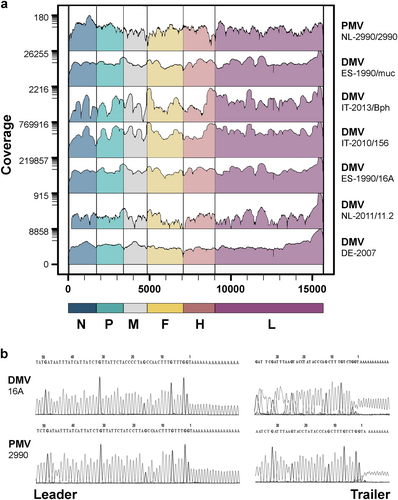
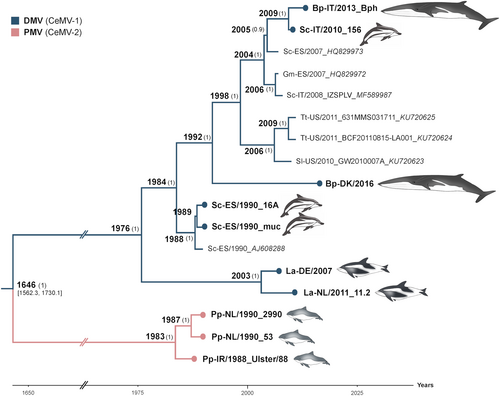
Fig. 2 Bayesian phylogenetic analysis of full-length CeMVs excluding L sequences. Most common recent ancestor ages are presented at the nodes with posterior values > 0.7 shown in parenthesis. New CeMV genomes generated during this study are presented with circles at the tips. Branches were truncated for graphical reasons. The taxon names are presented as host-country/year of collection_variant_GenBank accession number. Branch colors represent virus strains. New CeMV variants and GenBank accession numbers Bph (MH430938), 156 (MH430937), DK/16 (MH430939), 16A (MH430934), muc (MH430935), DE/2007 (MH430940), NL/11.2 (MH430941), 2990 (MH430945), 53 (MH430943), and Ulster/88 (MH430942). Bp Balaenoptera physalus, La Lagenorhynchus albirostris, Pp Phocoena phocoena, Sc Stenella coeruleoalba, Sl Stenella longirostris, and Tt Tursiops truncatus
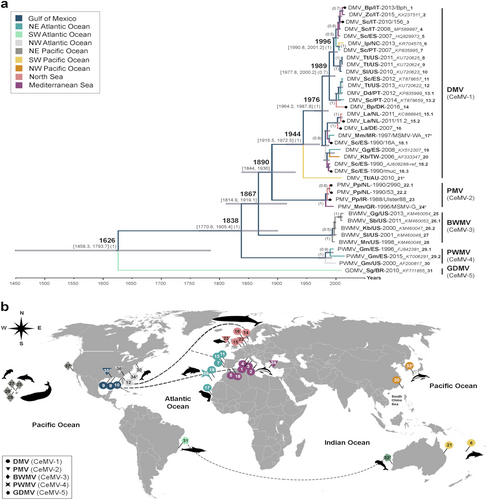
a Bayesian phylogenetic analysis of partial P genes (400 bp). Most common recent ancestor ages are presented at the nodes in cursive with 95% highest posterior density interval values in brackets and as gray horizontal bars. Posterior values > 0.5 are shown in parenthesis. CeMV genomes in this study are presented with black circles at the tips. Each branch is color-coded according to the ocean/sea in which the cetaceans were found. The taxon names are presented as virus_host/Country-year of collection/variant_GenBank accession number_ID for b. Sequences 17* and 24* were extracted from a published paperCitation22. Sequence 21* was kindly provided by Dr. Stone and Jianning Wang. Host abbreviations: Bp Balaenoptera physalus, Dd Delphinus delphis, Gg Grampus griseus, Gm Globicephala melas, Ip Indopacetus pacificus, Kb Kogia breviceps, La Lagenorhynchus albirostris, Mm Monachus monachus, Mn Megaptera novaeangliae, Pp Phocoena phocoena, Sb Steno bredanensis, Sc Stenella coeruleoalba, Sg Sotalia guianensis, Sl Stenella longirostris, Tt Tursiops truncatus, and Zc Ziphius cavirostris. b A world map of CeMV migration using sequences from a. Locations of viruses in the map are relative and were obtained from the publications in which the sequences were published. The hashtag (#) indicates no sequence availability (No. 32: CeMV-5 from AU/2010 and No. 33: CeMV-1_JP/1999). Proposed virus migratory routes (dashed lines) were based on sequence clades from the phylogenetic tree in a
Sequence variation between wild-type and isolated CeMV strains
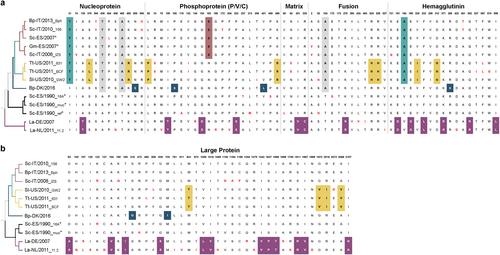
Fig. 4 Amino acid changes between DMV variants. Defined molecular signatures of each monophyletic group are color-colored. Single changes not representing the group are shown in red. The asterisk denotes sequences obtained from outbreaks. The taxon names are presented as host-Country/year of collection_variant (first three characters). a comparison of amino acid changes in the nucleoprotein, phosphoprotein, matrix, fusion and hemagglutinin in the complete set of DMV genomes. b Comparison of amino acid changes in the L protein. DMV variants and GenBank accession numbers: Bph (MH430938), 156 (MH430937), Sc-ES/2007 (HQ829973), Gm-ES/2007(HQ829972) IZSPLV (MF589987), 631MMS031711 (KU720625); BCF20110815-LA001 (KU720624), GW2010007A (KU720623), DK/2016 (MH430939), 16A (MH430934), muc (MH430935), ref (AJ608288), DE/2007 (MH430940), and 11.2 (MH430941). Host abbreviations: Bp Balaenoptera physalus, La Lagenorhynchus albirostris, Pp Phocoena phocoena, Sc Stenella coeruleoalba, Sl Stenella longirostris, and Tt Tursiops truncatus
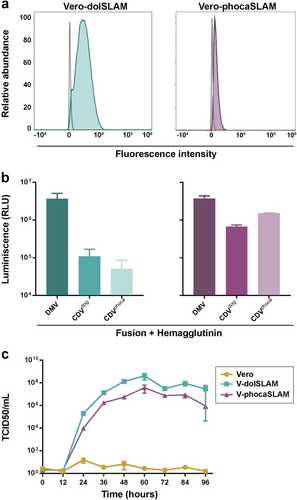
a Characterization of SLAM expression on Vero cells. Vero-dolSLAM cells (teal colored), Vero-phocaSLAM cells (purple colored) and untransfected Vero cells (gray line) were stained with a HA-tag antibody and analyzed by flow cytometry. b Cell-to-cell fusion activity of DMV, CDV-dog/2016, and CDV-phoca/1988 glycoproteins in Vero-dolSLAM cells (left) and Vero-phocaSLAM cells (right) as measured by β-galactosidase activity. Relative luminescence unit (RLU) values on the Y-axis were calculated by subtracting the RLU values obtained from Vero-dolSLAM or Vero-phocaSLAM cells with values generated from Vero cells (control). Experiments were performed in quadruplicates (n = 3). c Multistep growth curve analyses of DMV-16A inoculated at an moi of 0.001 in Vero cells (control), Vero-dolSLAM and Vero-phocaSLAM cells. Virus titers were determined in triplicates