Figures & data
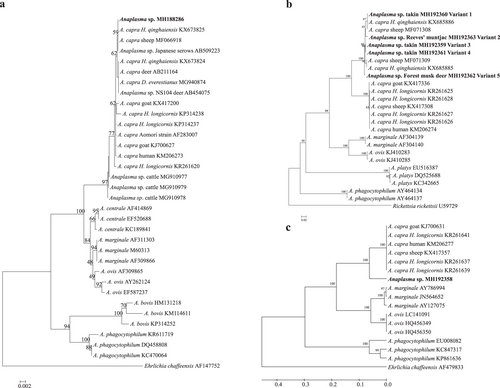
Fig. 1 Phylogenetic analysis of “Anaplasmacapra” based on partial sequences of the 16S rRNA (1261 bp, a), gltA (594 bp, b), and msp4 genes (656 bp, c). Phylogenetic trees were constructed based on the sequence distance method using the neighbor-joining (NJ) algorithm with the Kimura two-parameter model in MEGA 4.0 software. Bootstrap analysis was performed with 1000 replicates. Ehrlichia chaffeensis or Rickettsia rickettsii was used as the outgroup. Boldface indicates the sequences obtained in this study