Figures & data
Table 1. List of samples used for phylogenetic analysis.
Table 2. Primers used to sequence the ND2 gene in T. troglodytes subspecies.
Table 3. Polymorphic nucleotide sites defining the 12 ND2 haplotypes resolved from the 33 T. troglodytes sequences examined. Sequences are given in relation to haplotype 1, where a dot indicates identity. The subspecies that each haplotype was found in is listed.
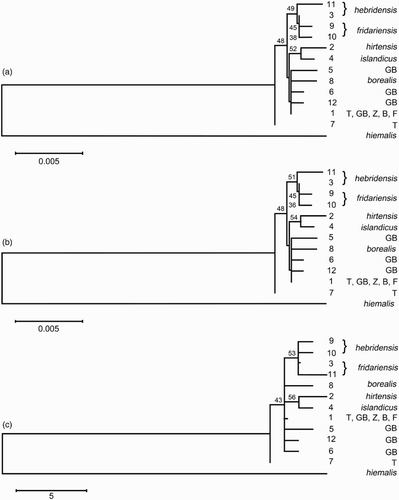
Figure 2. Minimum evolution (a) and neighbour joining (b) phylogenies of 13 Troglodytes ND2 haplotypes based on the Tamura–Nei distance with a gamma shape parameter of 1.31. Maximum parsimony (c) phylogeny shows most parsimonious tree with length = 58. Numbers at nodes denote bootstrap confidence indexes > 30%. Haplotype numbers (as per ) are given to the right of branch termini. T = nominate troglodytes, GB = birds from mainland Great Britain (troglodytes/indigenus), Z = zetlandicus, B = borealis and F = fridariensis.