Figures & data
Table 1. Localities and voucher specimens of Allium taxa examined in the present study.
Figure 1 Somatic chromosomes in the studied taxa. (a) Allium arlgirdense; (b) A. anacoleum; (c) A. microspathum; (d) A. rhetoreanum; (e) A. shirnakense; (f) A. oreophilum. Scale bars 10 μm.
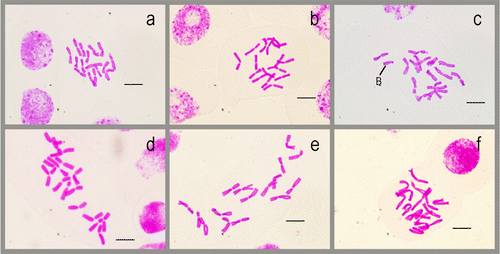
Figure 2 Haploid ideograms in the studied taxa. (a) Allium arlgirdense; (b) A. anacoleum; (c) A. microspathum; (d) A. rhetoreanum; (e) A. shirnakense; (f) A. oreophilum.
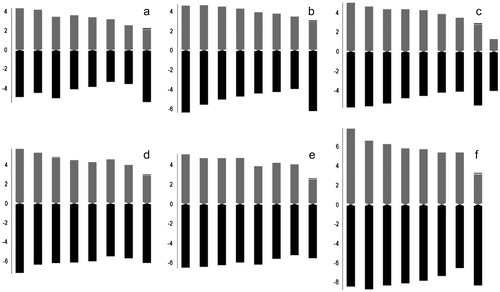
Table 2. Karyotype formula according to Levan et al. (Citation1964) and characteristics of the investigated taxa.
Table 3. Karyo-morphometric parameters and symmetry indices for the investigated taxa.
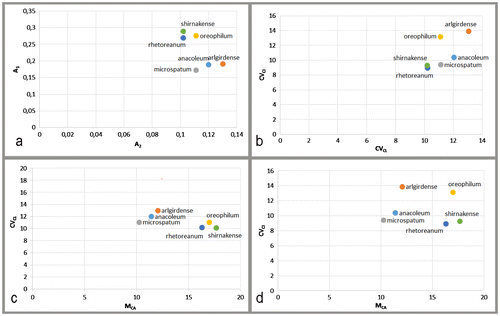
Figure 3 Scatter plot for the investigated taxa reported in Table : (a) against A2 (x axis) and A1 (y axis); (b) against CVCL (x axis) and CVCI (y axis); (c) against MCA (x axis) and CVCL (y axis); (d) against MCA (x axis) and CVCI (y axis).