Figures & data
Table 1. Adaptors and primers used in MSAP analysis.
Table 2. Primers for real-time quantitative PCR.
Table 3. MSAP-based cytosine methylation levels at different time-points under salt stress.
Table 4. Analysis of DNA methylation patterns under salt stress with respect to control conditions in maize seedlings.
Table 5. The BLAST results of differentially methylated sequences.
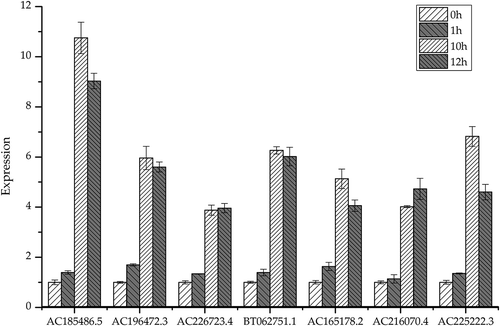
Figure 1. The expression of selected genes detected by RT-qPCR in maize inbred line LH196 at different time points. The transcript levels were normalized to that of GAPDH, and the level of each gene in the control samples was set at 1.0. Error bars represent the SEM for three independent experiments. The small letters a, b, c and d represent differences at level of P < 0.05 between different processing times.