Figures & data
Table 1 Examples of molecular methods used for environmental monitoring of aquatic habitats. Advantages and limitations, and examples of use in New Zealand are given.
Figure 1 Schematic of fluorescent in situ hybridisation (FISH). A fluorescently labelled complementary DNA or RNA probe binds to a specific target. Fluorescent microscopy is used to visualise where the fluorescent probe binds. The photo shows a fluorescein-labelled species-specific oligonucleotide probe, targeted against rRNA in a New Zealand native starfish.
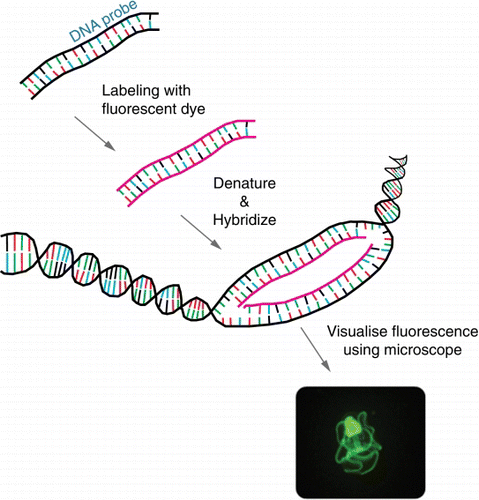
Figure 2 Schematic of sandwich hybridisation array (SHA). SHA uses an oligonucleotide capture probe targeting a specific DNA/RNA target and one or more signal probes, which attach to the bound region, forming a ‘sandwich’. The presence of target DNA/RNA in the sample is represented colorimetrically, fluorescently or chemiluminescently.
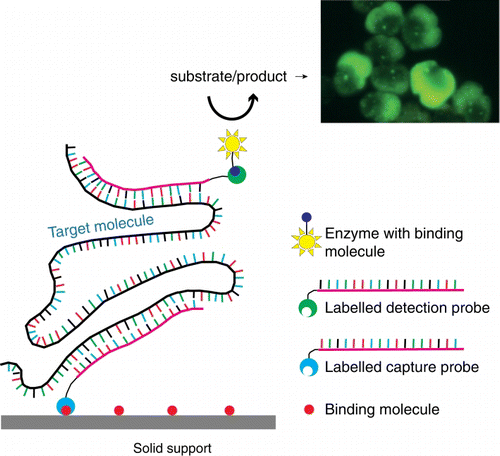
Figure 3 Schematic of end-point PCR and quantitative PCR (QPCR). Using a combination of deoxyribonucleotides (dNTP), oligonucleotides (primers) which are specific to the target, a polymerase enzyme and a series of heating and cooling steps, the DNA/RNA template is exponentially amplified. The resulting ‘amplicon’ products are visualised on an agarose gel for end-point PCR (bottom left) or by measuring florescence for QPCR (bottom right).
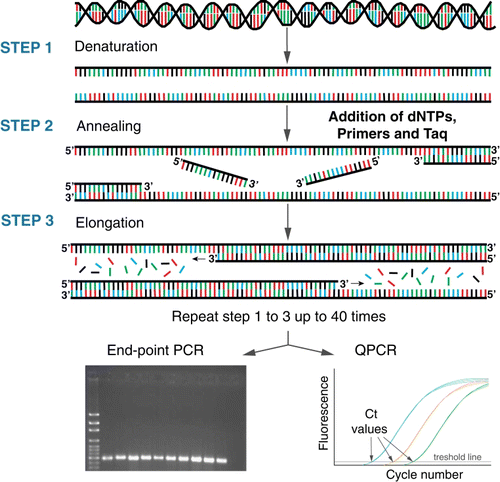
Figure 4 Schematic of automated ribosomal intergenic spacer analysis (ARISA). A PCR (see ) that targets the internal transcribed spacer (ITS) regions of the small and large subunit (SSU and LSU) rRNA operon is undertaken using a fluorescently labelled primer. The amplicon fragments are then separated by size on an electropherogram. The pattern of peaks provides information on the microbial community structure.
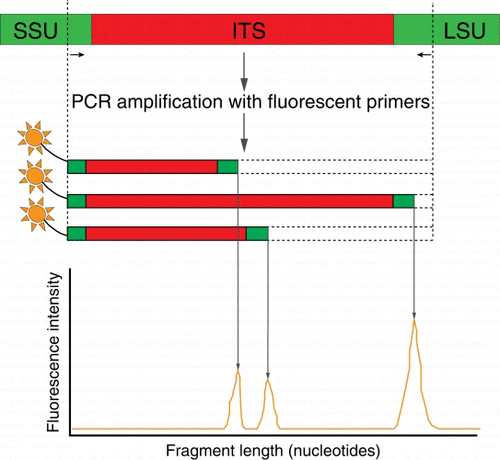
Figure 5 Schematic of Sanger sequencing. Each of the four dideoxyribonucleotides (ddNTPs) chain terminators are labelled with a fluorescent dye, emitting light at a specific wavelength, the actual sequence read is translated from peak trace chromatograms (generated by size separation, detection and recording of dye fluorescence) using capillary electrophoresis.
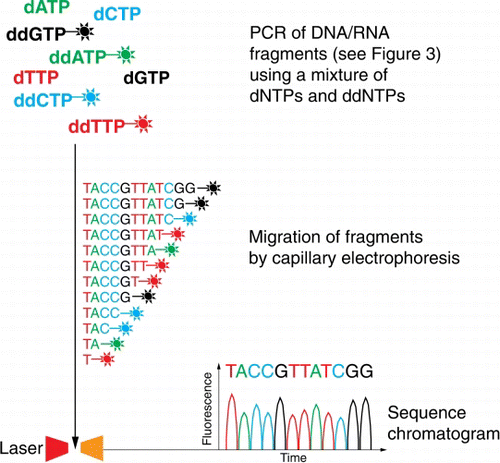
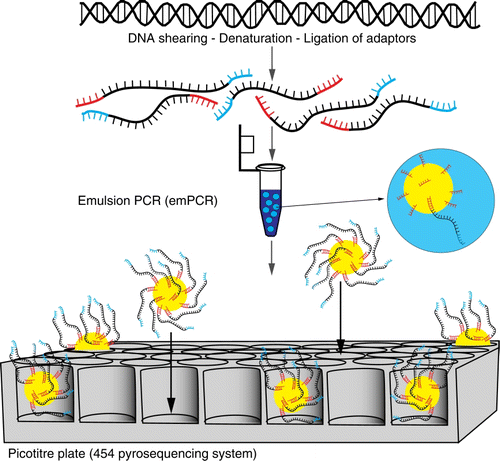
Figure 6 Schematic of Roche 454™ next-generation sequencing. A PCR reaction using primers with specifically designed adaptors is undertaken. These templates are bound to beads and amplified via emulsion PCR. Each bead is loaded into a picotitre plate and sequenced in parallel by flowing pyrosequencing reagents across the plate.