Figures & data
Figs 1–6. Images of Sirenicapillaria glauca (reference strain BLCC-M125).

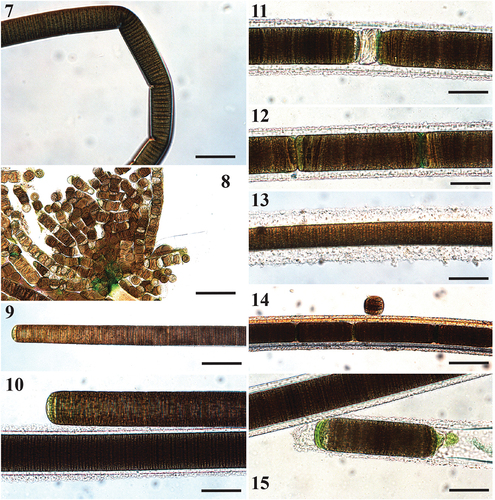
Figs 7–15. Microscope images of Sirenicapillaria rigida (reference strain BLCC-M116).

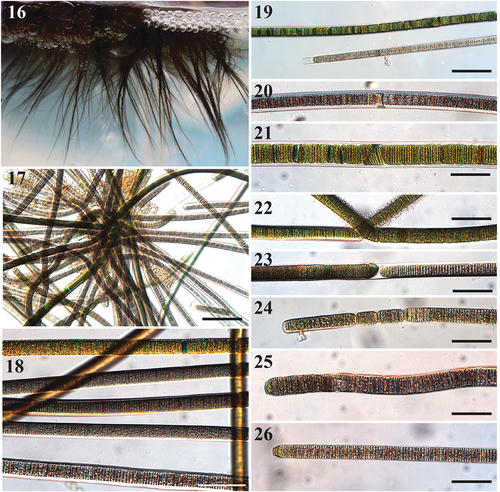
Figs 16–26. Images of Sirenicapillaria stauglerae (reference strain BLCC-M121).

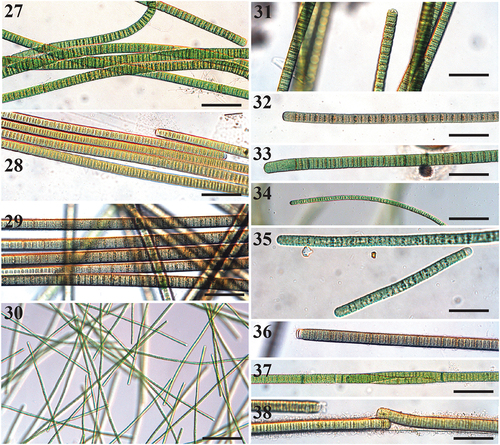
Figs 27–38. Microscope images of species of Tigrinifilum.

Figs 39–48. Microscope images of Capilliphycus. Scale bars = 50 µm.
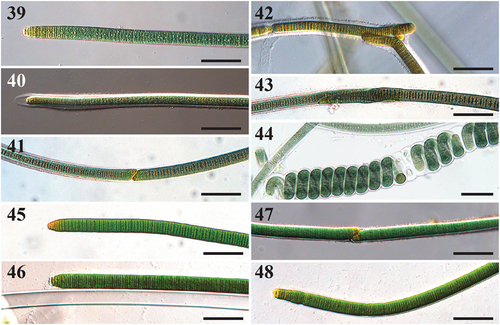
Table 1. Morphological characteristics of known species of Sirenicapillaria gen. nov., Tigrinifilum gen. nov. and Capilliphycus. Measurements (μm) are represented as min–max and mean ± st. dev
Fig. 49. Maximum likelihood and Bayesian inference, showing Bayesian topology of the phylogenetic relationship of the 16S rRNA gene sequence of the 30 cyanobacterial strains presented in this work and 141 cyanobacterial taxa including Gloeobacter violaceus PCC 7421 (NR074282) as outgroup. Bootstrap support and posterior probabilities ≥50 and 0.5 are shown at consensus nodes of both inferential analyses, respectively. (*) represents the reference strains for the species; triangle indicates the branch where the family Sirenicapillariaceae is delimited; stars denote the type species of the genus.
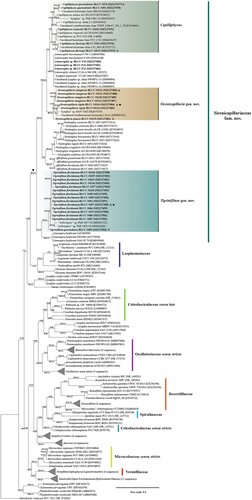
Table 2. Mean p-distance (percentage of genetic similarity) between and within (last column) established cyanobacterial families, based on 16S rRNA gene sequences. Values are based on an alignment of 91 taxa and 7 established cyanobacterial families including the novel family Sirenicapillariaceae with eight genera: Affixifilum, Capilliphycus, Lyngbya, Limnospira, Limnoraphis, Neolyngbya, Sirenicapillaria and Tigrinifilum. Taxonomic family classifications were based on Guiry & Guiry (Citation2021) and Hauer & Komárek (Citation2021)
Table 3. Morphological characteristics of the genera Affixifilum, Neolyngbya, Limnoraphis, Capilliphycus, Sirenicapillaria gen. nov. and Tigrinifilum gen. nov. within the family Sirenicapillariaceae fam. nov.
Table 4. Lengths of identifiable domains in the 16S–23S rRNA Internal Transcribed Spacer (ITS) regions of strains of Capilliphycus, Limnoraphis, Sirenicapillaria gen. nov. and Tigrinifilum gen. nov. Duplicated strain numbers indicate different ITS patterns (with or without tRNAs). Multiple values indicate variable operon length within species and/or strains. A dash (-) denotes not observed
Fig. 50. The 16S–23S rRNA ITS sequence secondary structures of the D1-D1ʹ of four cyanobacterial genera (Capilliphycus, Limnoraphis, Sirenicapillaria, Tigrinifilum) presented in this work. Nonbolded taxa represent previously published data. A: C. guerandensis BLCC-M76 & BLCC-M92. B: C. guerandensis BLCC-M79 & BLCC-M92 with tRNA. C: C. flaviceps BLCC-M53. D: C. flaviceps BLCC-M137 with and without tRNA. E: C. salinus. F: C. tropicalis. G: Limnoraphis sp. BLCC-F19 & BLCC-F23 with and without tRNA. H: S. glauca BLCC-M125. I: S. rigida BLCC-M116 & BLCC-M134 with and without tRNA. J: S. stauglerae BLCC-M121, BLCC-M122, BLCC-M123, & BLCC-M138. K: S. stauglerae BLCC-M121 with tRNA. L: T. floridanum BLCC-M48, BLCC-M50 clone A, BLCC-M57 clone C, BLCC-M66 clone B, BLCC-M77 clone C, BLCC-M87 clone C, BLCC-M107, BLCC-M118 clone B, BLCC-M119 clone B, & BLCC-M120 clones A & B, with tRNA. M: T floridanum BLCC-M50, BLCC-M57 clones A & B, BLCC-M66 clones A & C, BLCC-M73, BLCC-M87 clones A & B, BLCC-M118 clones A & C, BLCC-M119 clone C, & BLCC-M120 clone C, with tRNA. N: T. guerandensis BLCC-M99, with tRNA.
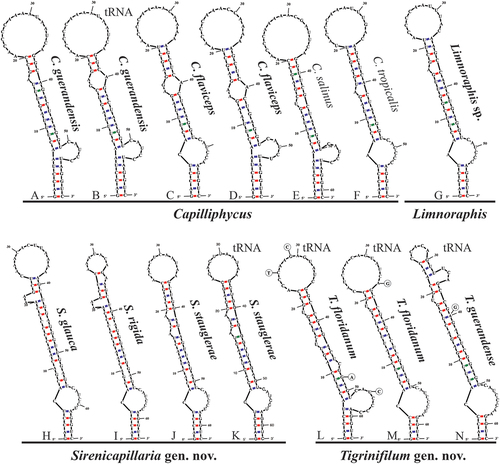
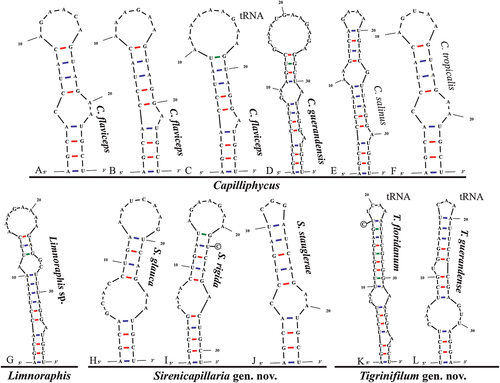
Fig. 51. The 16S–23S rRNA ITS sequence secondary structures of the Box-B of the four cyanobacterial genera (Capilliphycus, Limnoraphis, Sirenicapillaria, Tigrinifilum) presented in this work. Nonbolded taxa represent previously published data. A: C. flaviceps BLCC-M53. B: C. flaviceps BLCC-M137. C: C. flaviceps BLCC-M137 with tRNA. D: C. guerandensis BLCC-M76 & BLCC-M92 with and without tRNA. E: C. salinus. F: C. tropicalis. G: Limnoraphis sp. BLCC-F19 & BLCC-F23 with and without tRNA. H: S. glauca BLCC-M125. I: S. rigida BLCC-M116 & BLCC-M134 with and without tRNA. J: S. stauglerae BLCC-M121, BLCC-M122, BLCC-M123, & BLCC-M138 with and without tRNA. K: T. floridanum BLCC-M48, BLCC-M50, BLCC-M57, BLCC-M66, BLCC-M73, BLCC-M77, BLCC-M87, BLCC-M107, BLCC-M118, BLCC-M119, & BLCC-M120, with tRNA. L: T. guerandensis BLCC-M99, with tRNA.