Figures & data
Figure 1. A cluster dendrogram of 150 barley (Hordeum vulgare L.) accessions based on the results obtained by the TOPSIS method.
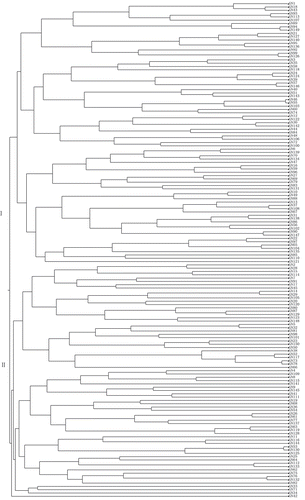
Figure 2. Economical characters of two barley (Hordeum vulgare L.) genotypes under normal and low phosphorus (P) levels after harvests (A: plant height, B: stem diameter, C: spike length, D: grain number of spike, E: total spike weight, F: thousand-grain weight, G: PUE). +P: normal P level; −P: P deficiency stress. Values are mean±standard deviation from three independent biological replicates. Lower case letters represent significant differences at the levels of P = 0.05.
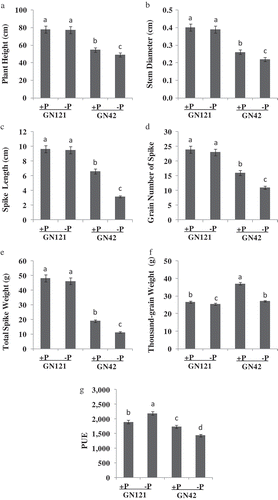
Table 1. Dry weight and root morphological parameters of two barley (Hordeum vulgare L.) genotypes under normal and low phosphorus (P) levels. +P: normal P level; −P: P deficiency stress. Values are mean ± standard deviation from three independent biological replicates. Lower case letters represent significant differences at the levels of P = 0.05.
Table 2. Chlorophyll and malondialdehyde (MDA) content of two barley (Hordeum vulgare L.) genotypes under normal and low phosphorus (P) levels. +P: normal P level; −P: P deficiency stress. Values are mean ± standard deviation from three independent biological replicates. Lower case letters represent significant differences at the levels of P = 0.05. FW: fresh weight.
Table 3. Protective enzymes activities of two barley (Hordeum vulgare L.) genotypes under normal and low phosphorus (P) levels. U (unit of enzyme activity): the amount of enzyme completely reacted with 1 μmol substrate during 1 min under the standard condition. +P: normal P level; −P: P deficiency stress; SOD: superoxide dismutase; POD: peroxidase; CAT: catalase. Values are mean ± standard deviation from three independent biological replicates. Lower case letters represent significant differences at the levels of P = 0.05.
Figure 3. Inorganic phosphate (Pi) concentration of two barley (Hordeum vulgare L.) genotypes under normal and low P levels. +P: normal P level; −P: P deficiency stress; PC: inorganic phosphate content; DW: dry weight. Values are mean ± standard deviation from three independent biological replicates. Lower case letters represent significant differences at the levels of P = 0.05.
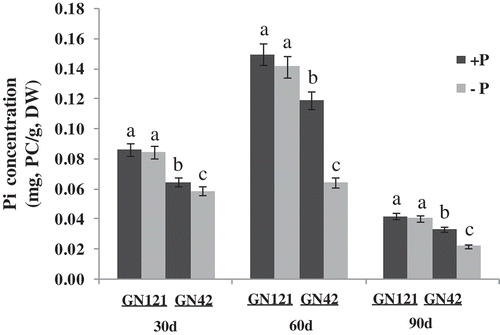
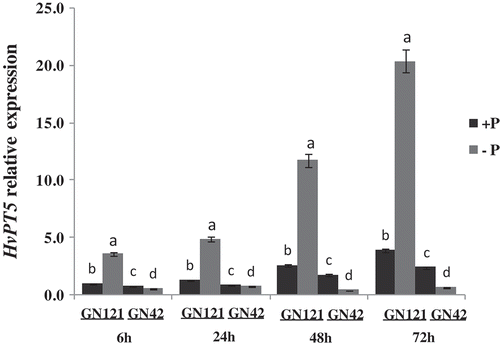
Figure 4. Relative expression analysis of HvPT5 in roots by qRT-PCR (quantitative reverse transcriptase polymerase chain reaction). +P: normal P level; −P: P deficiency stress. The y-axis indicates the relative expression level of HvPT5; values are mean±standard deviation from three independent biological replicates. Lower case letters represent significant differences at the levels of P = 0.05.