Figures & data
Figure 2. The eZ-value of S. marcescens (ATCC 6911). Each value is the average of three measured values and error bar represents the standard deviation.
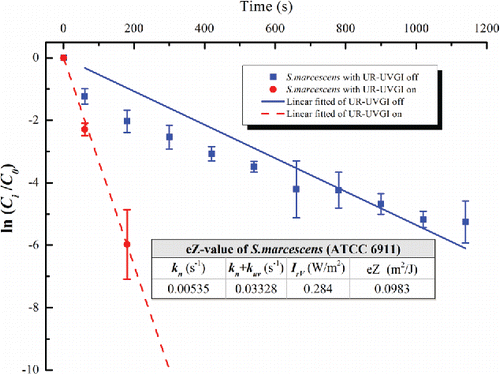
Figure 3. The eZ-value of S. epidermidis (ATCC 12228). Each value for UR-UVGI off and on is the average of four and three measured values, respectively. Error bar represents the standard deviation.
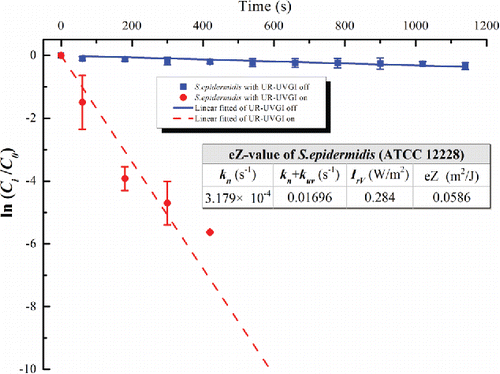
Figure 4. The eZ-value of P. alcaligenes (ATCC 14909). Each value for UR-UVGI off and on is the average of two and three measured values, respectively. Error bar represents the standard deviation.
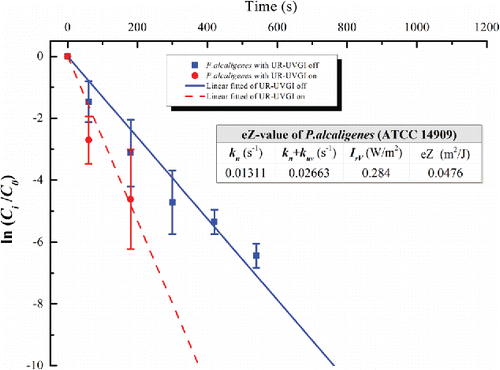
Figure 5. The eZ-value of M. luteus (ATCC 4698). Each value is the average of four measured values and error bar represents the standard deviation.
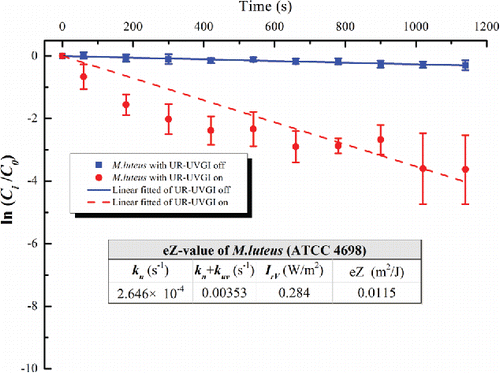
Table 1. The UV susceptibility constants (Z) of test bacteria reported in references Beggs et al. (Citation2006), Kowalski (Citation2009), Noakes et al. Citation(2004a), and Yang et al. (Citation2012).
Figure 6. Comparisons of the present simulation and the experiment of Yang et al. (Citation2012) and the simulation of Yang et al. (Citation2016). The inactivation rate (kuv) of experiment is the difference of the slopes derived from “Ventilation” and “Ventilation + UR-UVGI.” Each point value is the average values of location A, B, and C at w = 2.05 m (Figures 5(d) and 6(d) in Yang et al. Citation2012). Error bar represents the standard deviation.
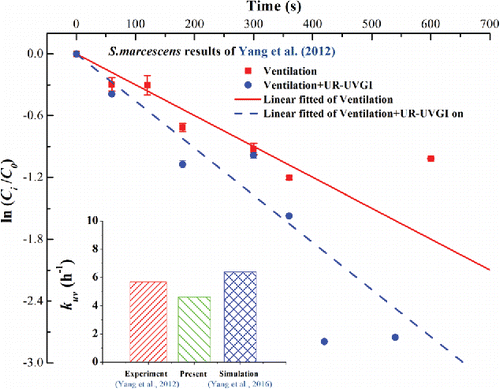
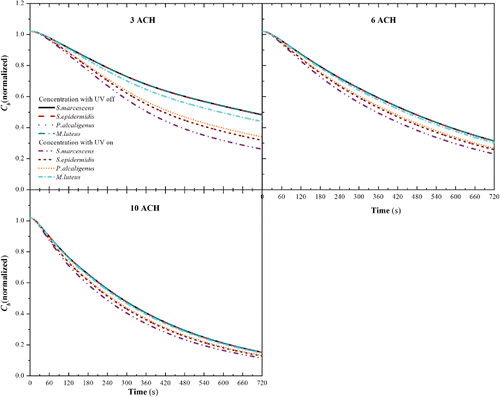
Figure 7. The average concentration of the tested bacteria in breathing zone (w ≤ 1.68 m) (Cb) with IrV = 0.284 W/m2 for different ACHs.