Figures & data
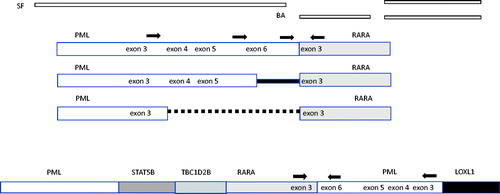
Figure 1. Molecular analysis of white blood cell DNA performed at RUMC. The karyotype did not show the expected t(15;17) rearrangement (a), but interphase and metaphase FISH revealed separation of the green and orange RARA break apart (BA) probe signals from their location on chromosome 17 (white arrows, b and c). Interphase and metaphase FISH with single fusion (SF) PML and RARA probes revealed the movement of RARA to chromosome 15 at the location of PML as a small yellow fusion signal (white arrows, d and e). The PML-RARA fusion gene was detected by reverse transcriptase PCR using primers to exon 3 and exon 6 of the PML gene and reverse primer to exon 3 of RARA (f: Lane 1, molecular weight standard, lane 2 PML exon 6 forward primer, lane 3 PML exon 3 forward primer lane 4, RARA exon 2 forward primer, lane 5 reagent blank). The major PCR product detected at about 500 bp (starred, lane 3) contained exons 3–6 of the PML gene.
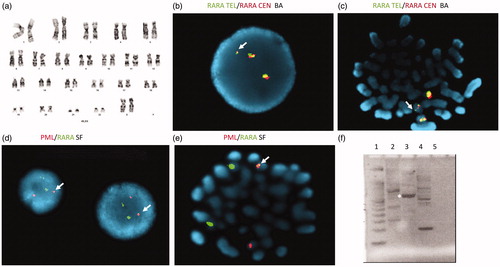
Figure 2. Structure and suggested structure of classical and cryptic translocations. Binding location of the single fusion (SF), break apart (BA) probes and RT-qPCR primers (arrows) are shown. In the classical APL t(15;17) translocations, the PML gene breaks at three breakpoint cluster regions (bcrs): intron 6 (bcr1), exon 6 (bcr2) and intron 3 (bcr3). RARA breakpoints occur in intron 2. A proposed structure based molecular analysis includes intra-chromosomal rearrangement of PML in addition to the insertion of RARA.