Figures & data
Figure 1. Uricosis. Chalk-like urate crystals are deposited on the epicardium of the heart, under the pleura and in the subcutaneous connective tissue. Swollen, pale, enlarged kidneys show a prominent tubular pattern. The ureters on both sides are dilated and fully packed with urate salts.

Figure 2. Degenerated tubules and mild interstitial nephritis among normal kidney parenchyma. The affected epithelial cells are necrotized and desquamated in the lumen of the tubule. Some of the epithelial cells contain cytoplasmic inclusion bodies (arrows).
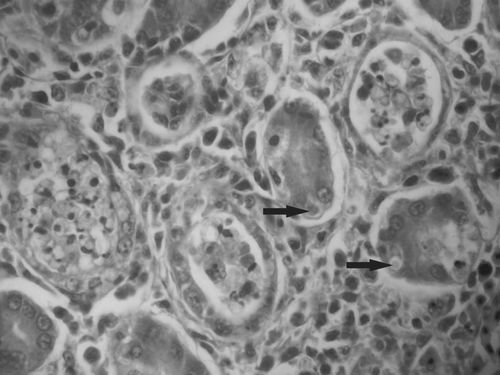
Figure 3. AVN-specific amplicons visualized by agarose gel electrophoresis. Lane M, molecular weight marker; lanes 1 to 10, samples; lane 11, positive control; lane 12, negative control. Lanes 5, 8, 9 and 10 show the ANV-specific 607 base pairs (bp) product, while lanes 1, 2, 3, 4, 6 and 7 show no amplified product and were considered negative.
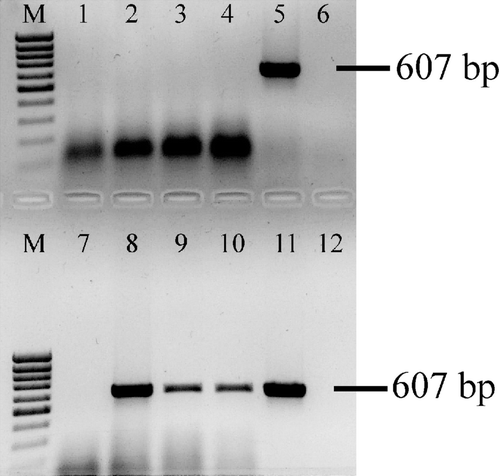
Figure 4. Phylogenetic relationship of the investigated strains based on the examined region of the Hungarian ANV strains and the reference strain. Three-letter codes indicate the farm: first number is the serial number of the sample, followed by the year of collection (20 …). The reference ANV strain (NC_003790; Imada et al., Citation2000) was used as the outgroup. Bar on the left demonstrates the genetic distance. Internal labels represent the bootstrap values of 1000 replicates.
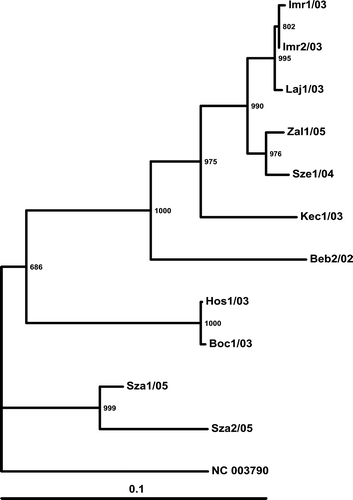
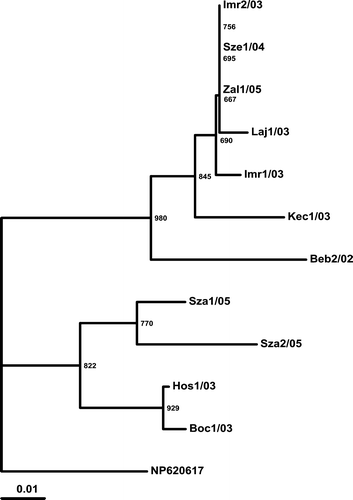
Figure 5. Phylogenetic relationship of the putative amino acid sequence of the investigated strains based on the examined region of the Hungarian ANV strains and the reference strain. Codes are indicated at . The reference ANV strain (NP_620617; Imada et al., Citation2000) was used as the outgroup. Bar on the left demonstrates the genetic distance. Internal labels represent the bootstrap values of 1000 replicates.