Figures & data
Fig. 1 The distribution of the 72 isolates of Rhizoctonia solani AG-1 IA collected from Guangdong, Guangxi and Hainan provinces.
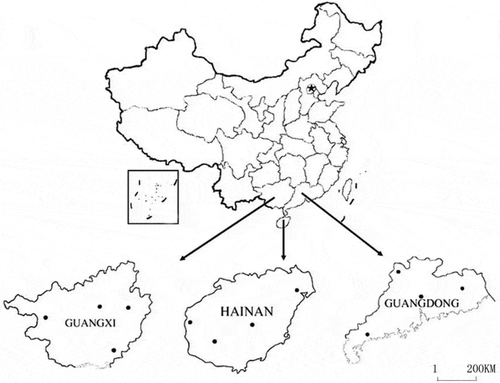
Table 1. Details of the 72 isolates of Rhizoctonia solani AG-1 IA from Guangdong (GD), Guangxi (GX) and Hainan (HN).
Table 2. Sequences of arbitrary primers, annealing temperature, numbers of amplified and polymorphic bands resulting from ISSR analysis.
Table 3. Genotype diversity of Rhizoctonia solani AG-1 IA from the 12 subpopulations.
Fig. 2 An example of the ISSR polymorphism of Rhizoctonia solani AG-1 IA isolates revealed by using ISSR primer EZ27. (Left to right: lane M, molecular weight marker DL5000 DNA ladder; lanes 1–16 represent the isolates).
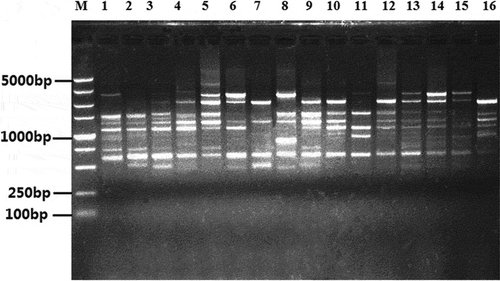
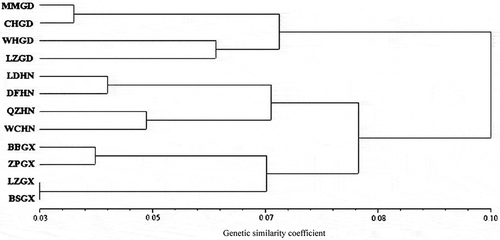
Fig. 3 Dendrogram of the 72 isolates of Rhizoctonia solani AG-1 IA based on ISSR data using Nei and Li’s similarity coefficient and UPGMA clustering method. The scale bar at the bottom represents the Li’s genetic similarity coefficient.
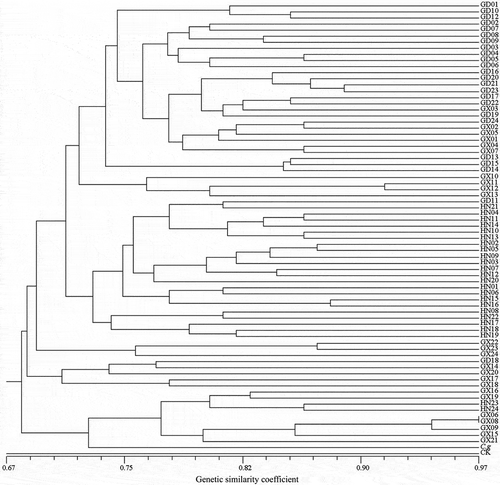
Table 4. Comparisons of genetic diversity and gene differentiation in three populations of Rhizoctonia solani AG-1 IA from south China.
Table 5. Analysis of molecular variance (AMOVA).