Figures & data
Table 1. Pathogenic variation of 39 Canadian isolates of Pyrenophora teres f. teres (net form net blotch of barley) on nine barley differentials based on a 1–10 scale. Regular and bold font numbers denote resistant (<5) and susceptible (≥5) phenotypes, respectively.
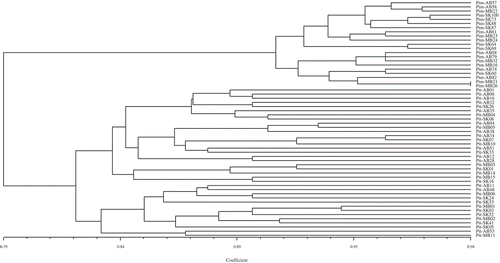
Fig. 1 Pathogenic similarity of 39 Pyrenophora teres f. teres (net form net blotch of barley) isolates recovered from barley crops in western Canada; two pathotype groups, highlighted in boxes, were found to be predominant. The dendrogram was produced using the unweighted pair-group method using the arithmetic means (UPGMA) procedure and simple similarity coefficient with NTSYSpc ver. 2.2.
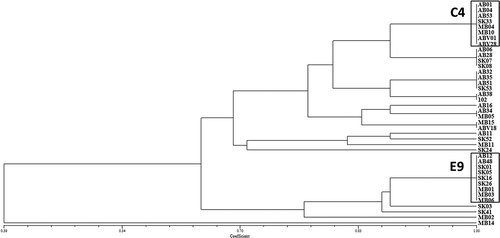
Fig. 2 a, Percentage of western Canadian Pyrenophora teres f. teres isolates virulent on nine barley differential genotypes; BT 201 (97.4%, 38 isolates), ‘OAC 21ʹ (84.6%, 33 isolates), ‘Norbert’ (35.9%, 14 isolates), TR 473 (33.3%, 13 isolates), ‘Steptoe’ (30.8%, 12 isolates), CI 9214 (28.2%, 11 isolates), ‘Heartland’ (12.8%, 5 isolates), CI 5791 (2.6%, 1 isolate), CI 9820 (2.6%, 1 isolate). b, Percentage of western Canadian Pyrenophora teres f. maculata isolates virulent on 11 barley differential genotypes; ‘Betzes’ (92.6%, 25 isolates), ‘Herta’ (88.9%, 24 isolates), TR 473 (85.2%, 23 isolates), ‘Norbert’ (85.2%, 23 isolates), ‘Bonanza’ (85.2%, 23 isolates), ‘Steptoe’ (81.5%, 22 isolates), CI 9820 (77.8%, 21 isolates), CI 5791 (70.4%, 19 isolates), ‘OAC 21ʹ (55.6%, 15 isolates), ‘Heartland’ (51.9%, 14 isolates), CI 9214 (7.4%, 2 isolates).
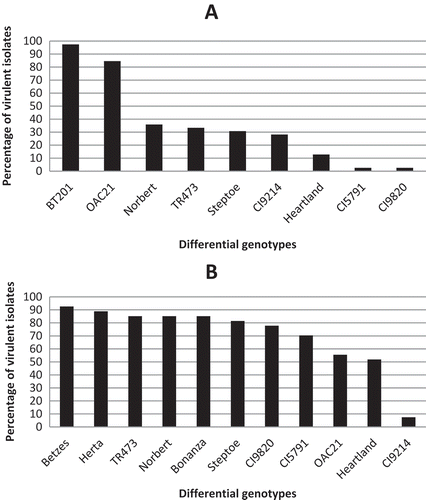
Fig. 3 a, Comparison of the main pathotype groups (A–J) found among Pyrenophora teres f. teres (net form net blotch of barley) populations from western Canada in 1985 (Tekauz Citation1990) versus 2009–2011 (current study). b, Comparison of the main pathotype groups (L–W) found among the Pyrenophora teres f. maculata (spot form net blotch of barley) populations from western Canada in 1985 (Tekauz Citation1990) versus 2009–2011 (current study).
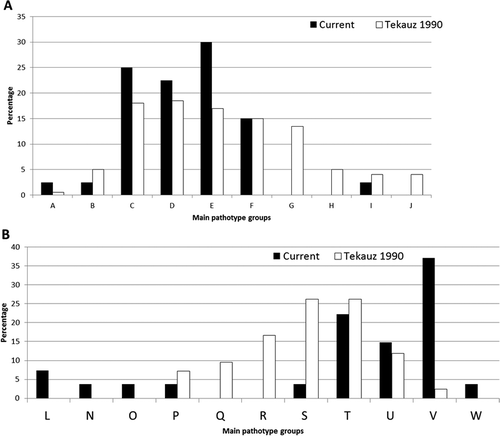
Table 2. Pathogenic variation of 27 Canadian isolates of Pyrenophora teres f. maculata (spot form net blotch of barley) on 11 barley differentials based on a 1–9 scale. Regular and bold font numbers denote resistant (1–3) and susceptible (>3) phenotypes, respectively.
Fig. 4 Pathogenic similarity of Pyrenophora teres f. maculata (spot form net blotch of barley) isolates recovered from barley crops in western Canada; two pathotype groups, highlighted in boxes, were found to be predominant. The dendrogram was produced using the unweighted pair-group method using the arithmetic means (UPGMA) procedure and simple similarity coefficient with NTSYSpc ver. 2.2.
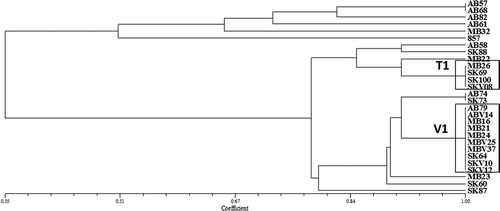
Fig. 5 Pathogenic similarity of 36 P. teres f. teres (Ptt) and 21 P. teres f. maculata (Ptm) isolates collected from western Canada. Isolates clustered in two distinct groups with 35 Ptt and 6 Ptm isolates in the first group and 1 Ptt and 15 Ptm isolates in the second group. The dendrogram was produced using the unweighted pair-group method using the arithmetic means (UPGMA) procedure and simple similarity coefficient with NTSYSpc ver. 2.2.
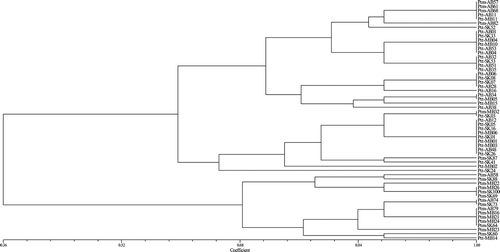
Fig. 6 Genetic similarity of 36 P. teres f. teres (Ptt) and 21 P. teres f. maculata (Ptm) isolates collected from western Canada. Cluster analysis revealed that all isolates clustered in two distinct divergent groups conforming to either Ptt or Ptm, with no intermediate cluster between the two forms. The dendrogram was produced using the unweighted pair-group method using the arithmetic means (UPGMA) procedure and simple similarity coefficient with NTSYSpc ver. 2.2.