Figures & data
Table 1. Primer sequences used in this study.
Table 2. Predicted open reading frames (ORFs) for the begomovirus and associated DNA-satellites identified from Malva parviflora.
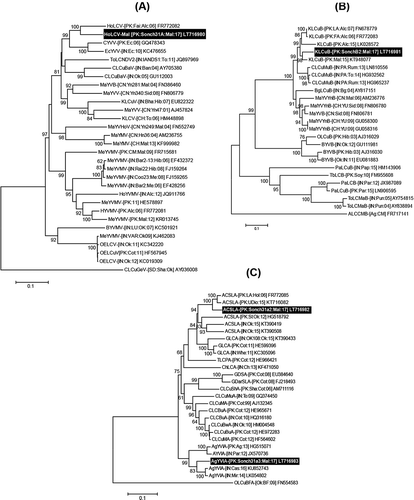
Fig. 2 Phylogenetic dendrograms based on complete nt sequences of (a) begomovirus, (b) betasatellite, and (c) alphasatellite isolates, using Neighbour-Joining (NJ) algorithm in MEGA7. All the isolates identified from Pakistan in this study are shown in bold white text on black background. Horizontal lines represent nt substitutions per site. Numeric values at branch nodes are representing per cent bootstrap values higher than 60 (1000 replicates). All isolates used for comparison are represented by their respective accession numbers in the trees. All abbreviations for begomovirus, alpha- and betasatellite isolates are according to Brown et al. (Citation2012) and Briddon et al. (2012), respectively.