Figures & data
Table 1. Fifteen wild-type isolates and five tebuconazole-resistant mutants of Villosiclava virens used in this study.
Fig. 1 (Colour online) Symptoms of rice false smut caused by Villosiclava virens. (a) typical symptoms observed on rice panicles in the greenhouse. (b) close-up of galls.

Table 2. Primers used in this study.
Fig. 2 (Colour online) Sequence alignment of the polymorphic cyp51 coding region from tebuconazole-sensitive isolates and resistant mutants of V. virens.

Fig. 3 Sensitivity test of allele-specific PCR for detection of T409C mutation in tebuconazole-resistant mutant F10-338D of V. virens. Lanes 1–7, genomic DNA templates of F10-338D at concentrations of 600, 60, 6, 0.6, 0.06, 0.006 and 0.0006 ng µL−1, respectively. M, 2-kb DNA ladder (Takara Bio).
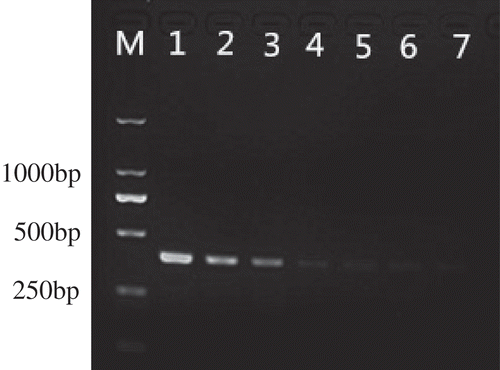
Fig. 4 Allele-specific PCR amplification products from genomic DNA extracted from eight tebuconazole-sensitive isolates and five tebuconazole-resistant mutants of V. virens. M, 2-kb DNA ladder (Takara Bio); lane 1, negative control; lanes 2–9, isolates F10-338, JY13107, JY1601, NH13018, NH13047, SX13023, SH13002 and SH13003; Lanes 10–14, isolates F10-338B, F10-338C, F10-338D, JY13107C and JY13107D.

Fig. 5 Allele-specific PCR amplification products from genomic DNA extracted from false smut balls. M, 2-kb DNA ladder (Takara Bio); lane 1, negative control; lanes 2–6, false smut balls derived from tebuconazole-resistant mutants; lanes 7–18, false smut balls derived from tebuconazole-sensitive isolates.

