Figures & data
Table 1. Characteristics of the studies in meta-analysis
Table 2. Distribution of miR-146a rs2910164 C > G genotypes and alleles
Table 3. Quality assessment of the meta-analysis
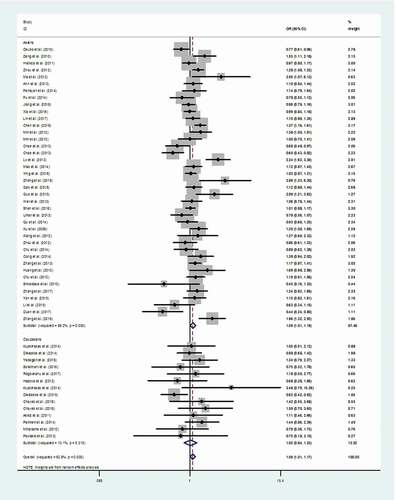
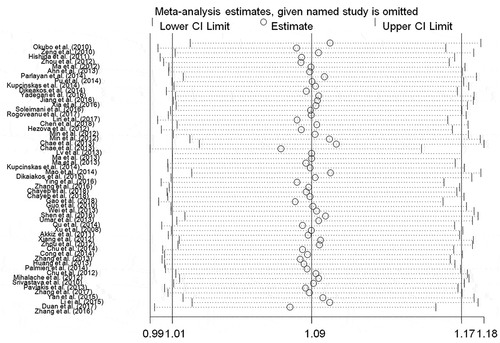
Table 4. Results of the meta-analysis from different comparative genetic models