Figures & data
Figure 1. (A) volcano plot, |Log2FC|≥0.107 and p < 0.05 were chosen as filtering conditions, scanned 264 upregulated genes and 202 downregulated genes. (B) Heatmap of top 50 different expressed genes.
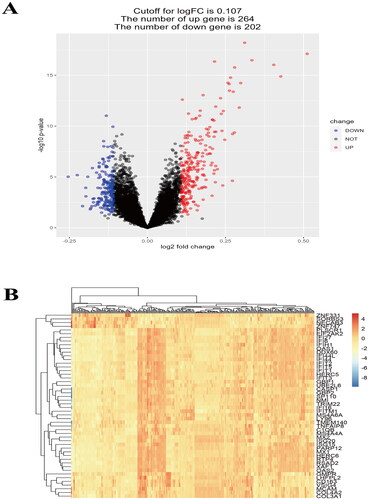
Figure 2. GO and DO enrichment analysis of DEGs. (A) BP enrichment analysis. (B) CC enrichment analysis. (C) MF analysis. (D) DO enrichment analysis.
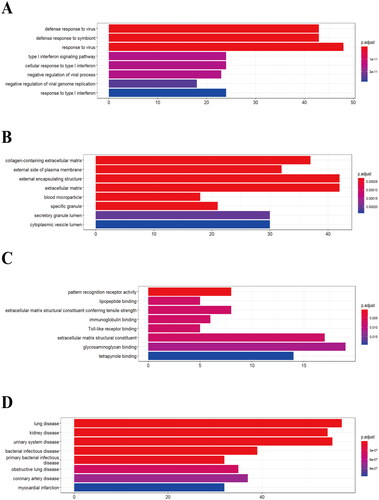
Figure 4. A. prediction performance of each machine learning model in training data and testing data, the models were arranged by average AUC in each dataset. B. PPI network of targeting genes.
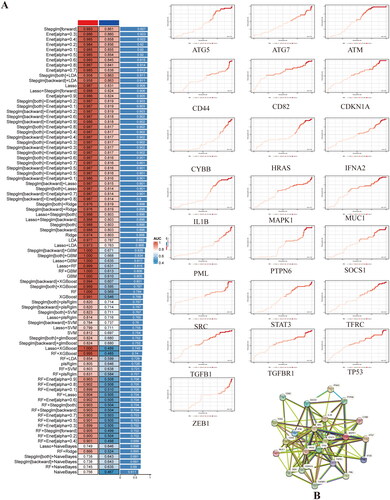
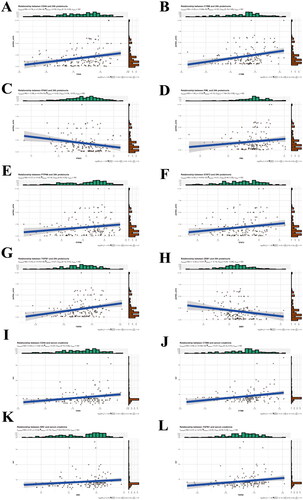
Figure 5. Correlation between targeting genes and clinical factors (proteinuria and serum creatinine (SCr).
Supplemental Material
Download Zip (1.3 MB)Data availability statement
We can acquire the data from GEO Database (GSE32591, GSE127797, GSE69438, GSE99967, GSE113342 and GSE200306) and FerrDb database.