Figures & data
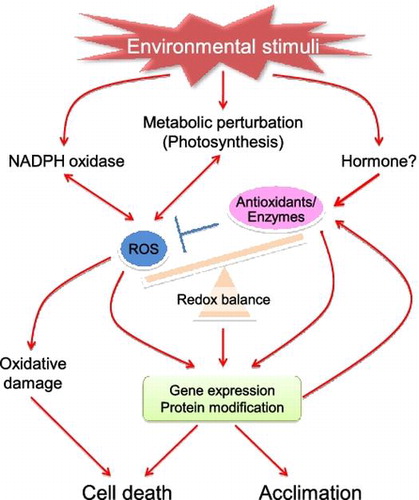
Fig. 2. The ascorbate-glutathione cycle.
Notes: APX, ascorbate peroxidase; AsA, L-Ascorbic acid; DHA, dehydroascorbate; DHAR, DHA reductase; GSH, reduced glutathione; GR, glutathione reductase; GSSG, oxidized glutathione; MDA, monodehydroascorbate; MDAR, MDA reductase.
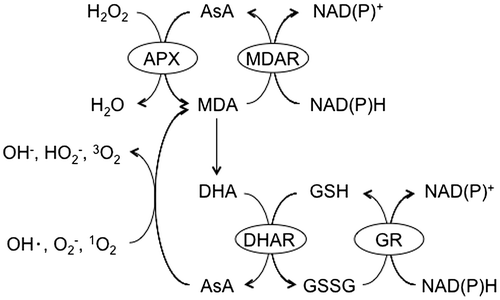
Fig. 3. Proposed ascorbate biosynthetic pathways in higher plants.
Notes: Thick arrows indicate the d-Man/l-Gal pathway. Parentheses indicate the names of ascorbate-deficient (VTC) Arabidopsis mutants. Enzymes: Enzymes: 1, phosphomannose isomerase; 2, phosphomannomutase; 3, GDP-d-Mannose pyrophosphorylase; 4, GDP-d-Mannose-3′,5′-epimerase; 5, GDP-l-Galactose phosphorylase; 6, l-Galactose-1P phosphatase; 7, l-Galactose dehydrogenase; 8, l-Galactono-1,4-lactone dehydrogenase; 9, l-Gulono-1,4-lactone dehydrogenase; 10, d-Galacturonate reductase; 11, aldonolactonase; 12, purple acid phosphatase; 13, myo-inositol oxygenase; 14, d-Glucuronate reductase.
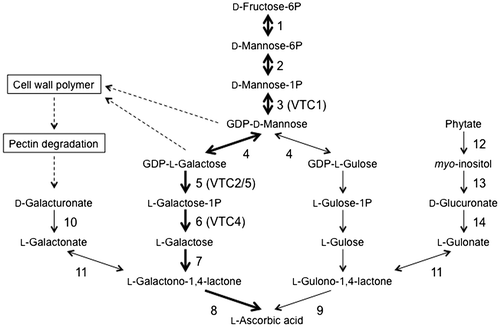
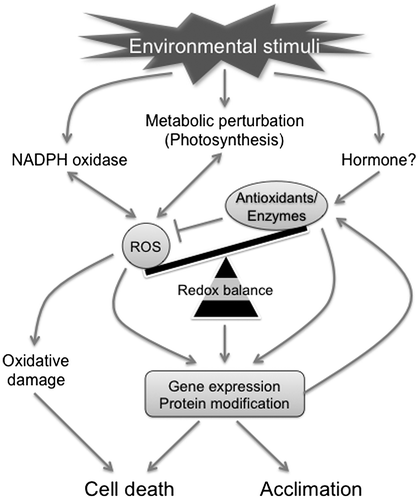
Fig. 4. Proposed model for chloroplastic H2O2-mediated signaling.
Notes: CAS, chloroplast calcium sensor; MVR, methylviologen-resistant; MVS, methylviologen-susceptible; PS, photosystem; SA, salicylic acid.
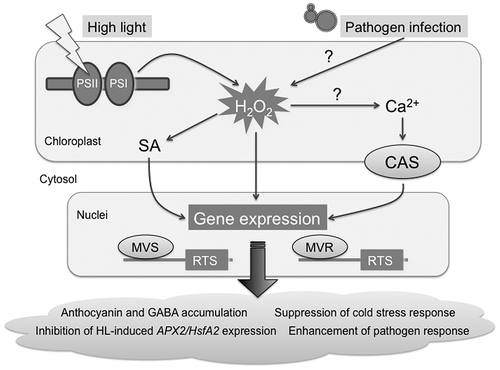
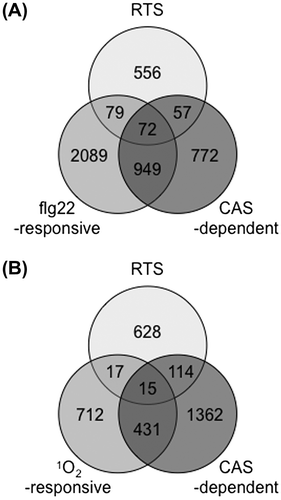
Fig. 5. The Relationship between chloroplastic ROS- and CAS-dependent responses.
Notes: (A) Overlap between chloroplastic H2O2-responsive,Citation102) flg22-responsive,Citation151) and CAS-dependent genes.Citation150) (B) Overlap between chloroplastic H2O2-responsive, 1O2-responsive,Citation137) and CAS-dependent genes.