Figures & data
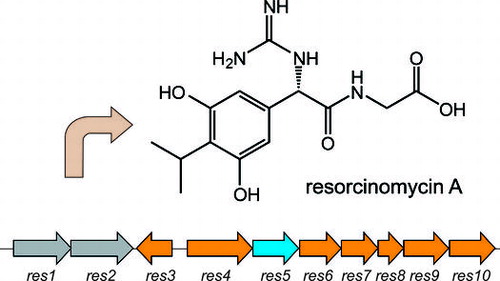
Fig. 1. Structures of resorcinomycin (1), pheganomycins (3), and their corresponding N-terminal nonproteinogenic amino acids (2 and 4).
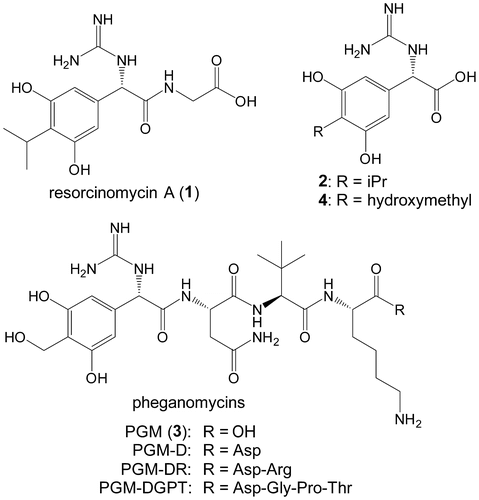
Fig. 2. The resorcinomycin biosynthetic gene cluster identified in Streptoverticillium roseoverticillatum.
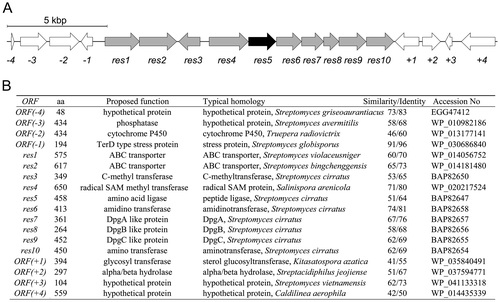
Fig. 3. Disruption of the res5 gene.
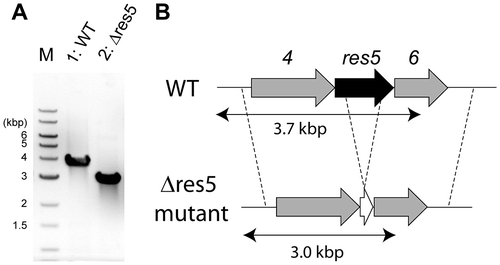
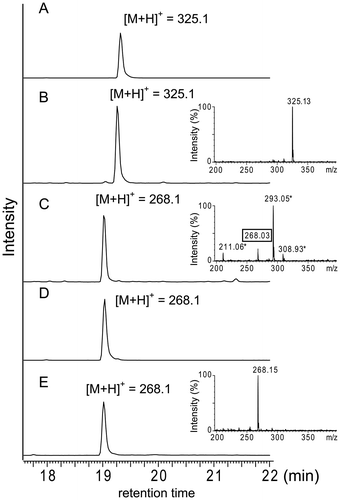
Fig. 4. LC-ESI-MS analysis of products accumulated in the culture broth of the res5 disruptant and S. lividans carrying the res gene cluster.