Figures & data
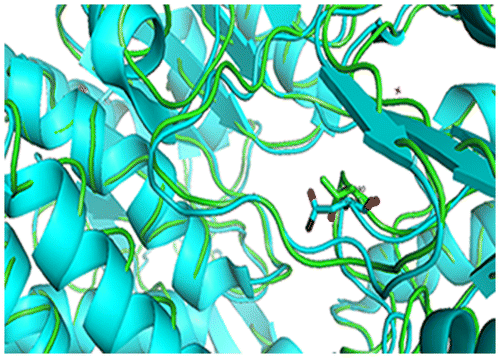
Fig. 1. Sequence and structure alignment of ALDA and BADH, ALDA shows in green, BADH shows in yellow.
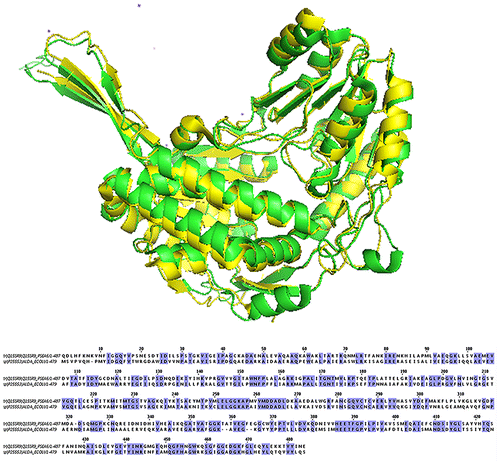
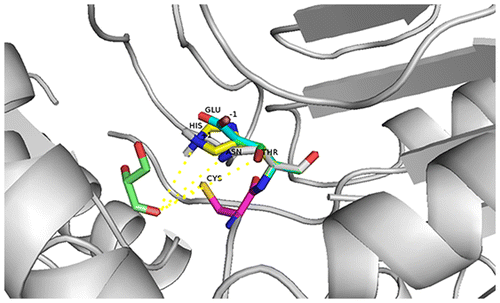
Table 1. Gibbs free energy of protein-substrate interaction calculated by Amber 12, with d-glyceraldehyde as substrate.
Table 2. Substrate specificity of ALDA and mutants.
Fig. 2. Sodium dodecyl sulfate polyacrylamide gel electrophoresis analysis of ALDA and mutants. M protein markers with indicated molecular mass (kDa) indicated alongside. ALDA wild-type ALDA, H449R mutants H449R, L158Y mutantsL158Y, N286H mutants N286H, N286E mutants N286E, N286T mutants N286T.
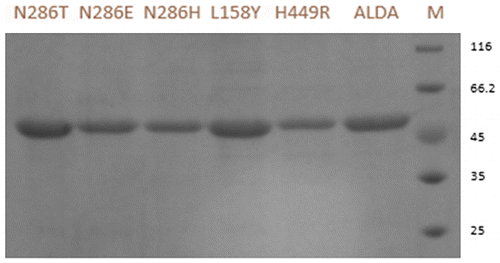
Table 3. Kinetic parameters of ALDA and mutants.
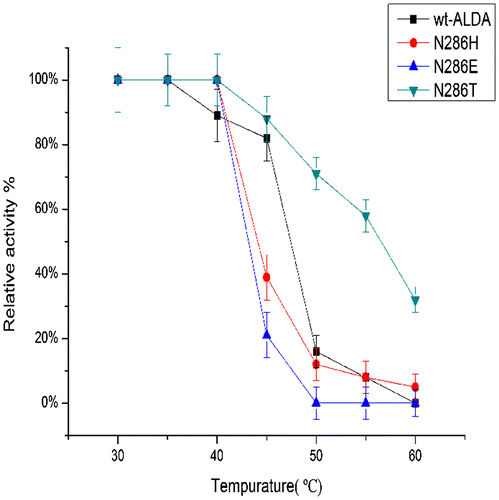
Fig. 5. Effects of temperature on the stability of ALDA and mutants, with l-lactaldehyde as substrate.