Figures & data
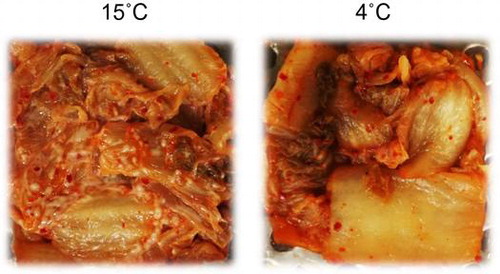
Figure 1. Changes in pH (A) and viable cell counts of LAB (B) and yeast (C) in kimchi A (circles), kimchi B (triangles), and kimchi C (squares). Kimchi was fermented at 4 °C (open symbols) or 15 °C (filled symbols) for 28 days. Viable cell counts of LAB and yeast were determined with GYP and YPD agar media, respectively.
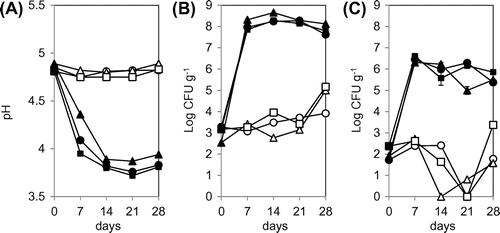
Figure 2. PCR-DGGE analysis of yeast communities in three types of kimchi fermented at 4 or 15 °C. Bands labeled with letters were excised and their DNA sequences were determined for identification of yeasts (see Table ). The numbers above each lane represent the fermentation period (days).
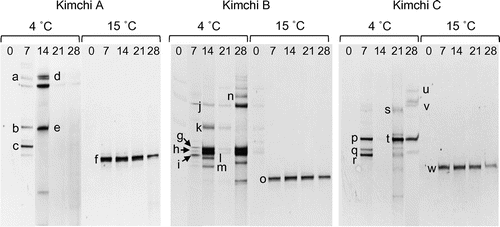
Table 1. Sequence analysis of 26S rRNA genes recovered from DGGE bands.
Figure 3. PCR-DGGE analysis of yeast communities in three types of kimchi after white-colony formation. Kimchi was fermented at 4, 10, 15, or 25 °C until white colonies appeared on the surface. Note that white-colony formation at 4 °C was detected only in kimchi C. Bands labeled with letters were excised, and their DNA sequences were determined for identification of yeasts (see text). Numbers above each lane represent fermentation temperature (°C).
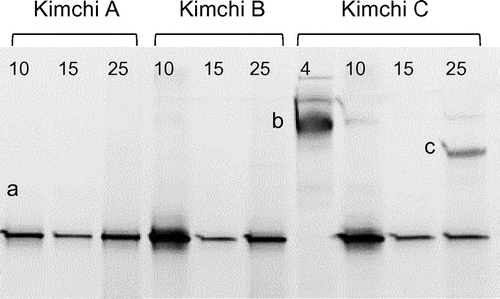
Table 2. Yeast strains isolated directly from white colonies formed on kimchi surface.
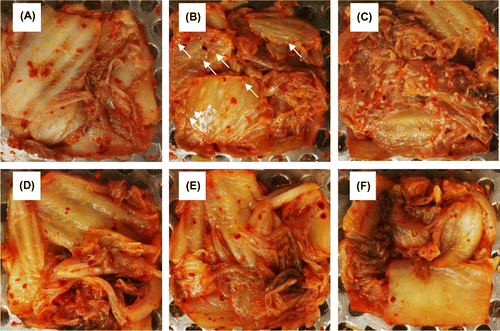
Figure 4. White-colony formation on the surface of kimchi A inoculated with yeast strains. Freshly prepared kimchi A was heat-treated and inoculated with 102 CFU g−1 of Kazachstania exigua (B and E) or K. pseudohumilis (C and F). Control kimchi without yeast inoculation is also shown (A and D). The kimchi was incubated at 15 °C for 21 days (panels A–C) or at 4 °C for 51 days (D–F). Two strains were inoculated individually for each yeast species, but essentially the same results were obtained. White arrows indicate the yeast colonies.