Figures & data
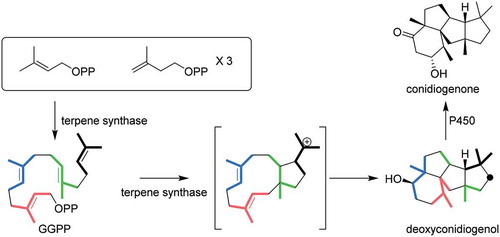
Figure 2. GC-MS and LC-MS profiles of the extracts from enzymatic reaction and heterologous expression.
(a). GC-MS profile of PchDS reaction. Upper panel: chromatogram showing total ion current (TIC). Lower panel: MS spectrum of 2. (b). GC-MS profile of metabolites extracted from AO-PrDS and A. oryzae NSAR1. Upper panel: chromatograms extracted at m/z 290. Lower panel: MS spectrum of 2. (c). LC-MS profile of AO-PrDS and AO-PrDS/PrP450. Chromatograms are extracted at m/z 305.
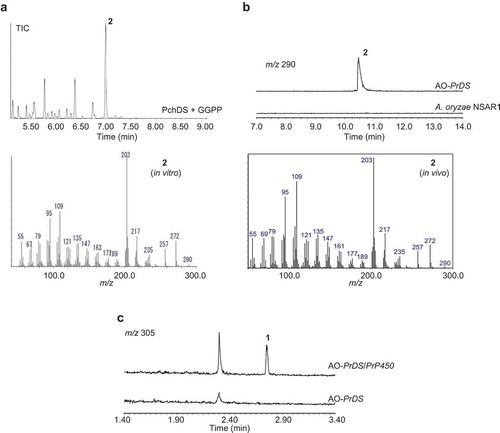
Table 1. 13C chemical shifts and multiplicity of [13C20]-2 (CDCl3, 150 MHz).
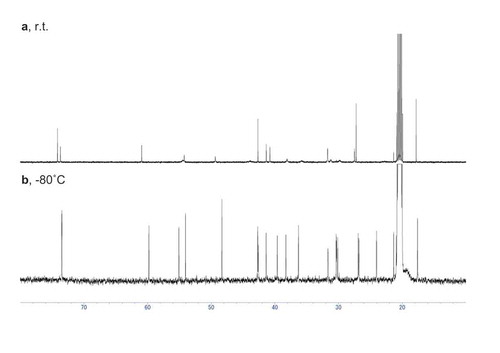
Figure 3. 13C NMR spectra of 13C-labeled 2 enzymatically synthesized by PchDS (CDCl3, 150 MHz).
The solvent signal at δC 77.0 was used as a reference.
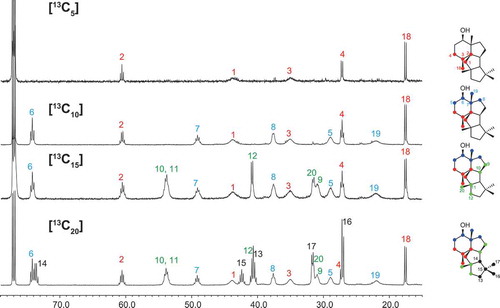
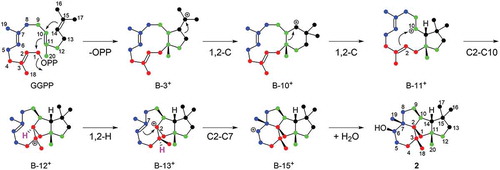
Scheme 1. Proposed cyclization mechanism catalyzed by PchDS/PrDS.
The numbers of intermediates are consistent with those in ref. 22.