Figures & data
Table 1. Physico-chemical parameters of water samples from inshore and intertidal pools at Luz beach, Portugal.
Table 2. Target genes/sequences and oligonucleotide primers used in this study.
Figs 1–8. Sampling site and samples of the colonial cyanobacterium LEGE 06123. Figs 1, 2. The sampling site at Luz beach, showing location in southern Portugal (arrowhead). . Unicellular cyanobacterium (arrowhead) in mat collected from shallow puddle. . Isolate LEGE 06123 showing sub-spherical cells which divide irregularly in various planes with dispersion through disintegration of colonies after intensive cell division (arrow). . Firm sheath (arrow) and amorphous polysaccharide layer (arrowhead) surround cells stained with Alcian blue. . Ultrastructure (TEM) showing the laminated nature of the sheath and the concentric arrangement of thylakoids which can exhibit invaginations towards the centre of the cell (sh, sheath, th, thylakoids). Scale bars: 10 µm (light micrographs); 1 µm (TEM).

Table 3. Morphometric measurements of cyanobacterial strain LEGE 06123.a
Fig. 11. Growth curves of LEGE 06123 in ASNIII medium supplemented with various concentrations of iron (FeCl3 · 6H2O).
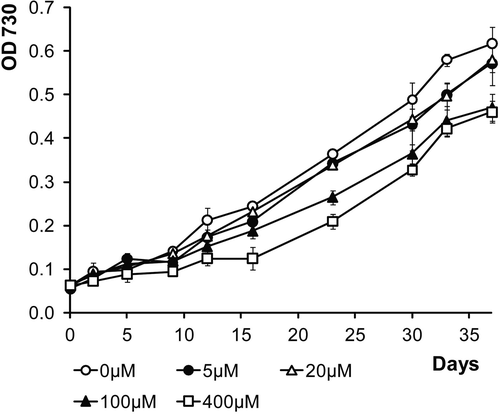
Fig. 12. Transmission electron micrograph of a 14-day-old culture of the cyanobacterium LEGE 06123 in liquid medium ASNIII supplemented with 400 µM of iron (FeCl3 · 6H2O). The arrow shows the metal accumulation outside the cell. Scale bar: 0.5 µm.
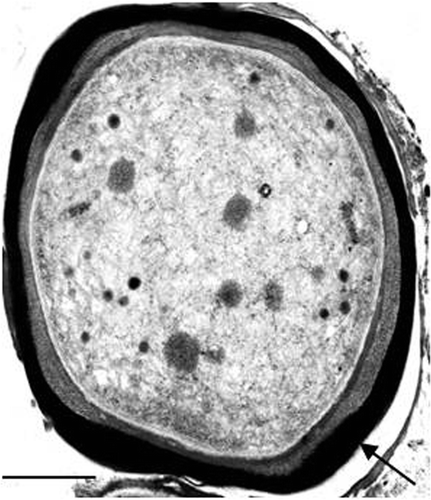
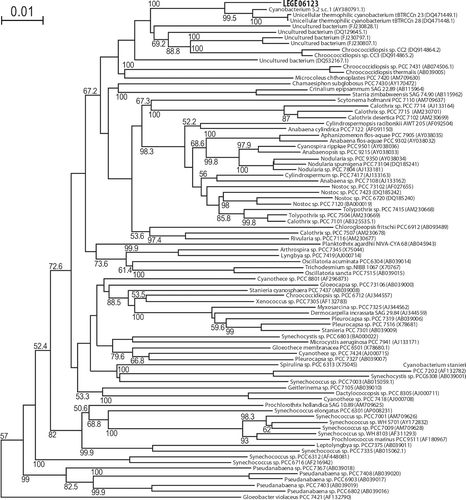
Fig. 13. Phylogenetic tree of 16S rRNA gene sequences from several cyanobacteria generated by NJ analysis. LEGE 06123 is shown in bold. The tree was rooted with the outgroup Chloroflexus aurantiacus J-10-fl (D38365.1), which was later pruned from the tree. Bootstrap values >50% are indicated beside the nodes. Each taxon is identified by the accession number (brackets) and whenever possible by the name.