Figures & data
Fig. 1. The phytoplankton community structure in surface seawater sampled at Huaniao Island in the coastal East China Sea in spring (17 April 2012 and 24 March–18 April 2013), summer (31 July 2012 and 18 July–8 August 2013), autumn (6 October 2012 and 23 October–12 November 2013) and winter (29 December 2012 and 1–12 January 2013) periods. ‘Others’ include Zetaproteobacteria, Euglenida, Klebsormidiophyceae, Trebouxiophyceae and Ulvophyceae (Form IAB), and Dictyochophyceae, Fragilariophyceae, Pelagophyceae, Eustigmatophyceae, Chrysophyceae and Raphidophyceae (Form ID).
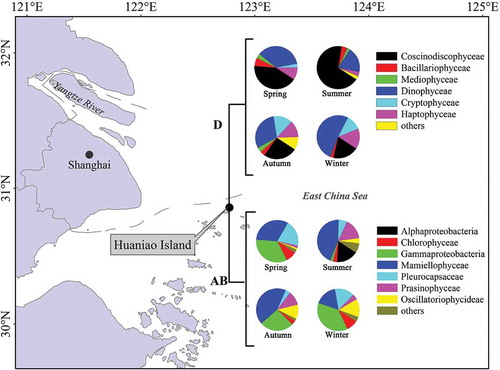
Table 1. Total number of clones, library coverage, number of OTUs, clones of dominant OTUs, diversity index and classification of dominant OTUs derived from Form 1AB and Form 1D rbcL for surface seawater sampled in the coastal East China Sea in spring (17 April 2012 and 24 March–18 April 2013), summer (31 July 2012 and 18 July–8 August 2013), autumn (6 October 2012 and 23 October–12 November 2013) and winter (29 December 2012 and 1–12 January 2013) periods.
Table 2. Closest relatives to clones amplified by Form 1AB and Form 1D rbcL for surface seawater sampled in the coastal East China Sea in spring (17 April 2012 and 24 March–18 April 2013), summer (31 July 2012 and 18 July–8 August 2013), autumn (6 October 2012 and 23 October–12 November 2013) and winter (29 December 2012 and 1–12 January 2013) periods. Relative OTUs (top three dominant OTUs) were obtained using the NCBI BLAST algorithm.
Fig. 2. Phylogenetic tree obtained from a Neighbour joining analysis for the most dominant OTUs (number of clones in parentheses) discriminated by Form IAB rbcL for surface seawater sampled in the coastal East China Sea in four seasons. Sequences existing in GenBank are included with accession numbers in parentheses. The numbers at the nodes represent full heuristic bootstrap values (with 1000 replicates) greater than 50%. The scale bar represents amino acid substitutions/site.
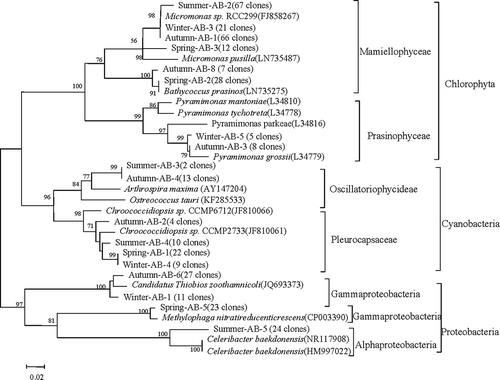
Fig. 3. Phylogenetic tree obtained from a Neighbour joining analysis of the most dominant OTUs (number of clones in parentheses) discriminated by Form 1D rbcL for surface seawater sampled in the coastal East China Sea in four seasons. Sequences in GenBank are included with accession numbers in parentheses. The numbers around the branches represent full heuristic bootstrap values (with 1000 replicates) greater than 50%. The scale bar represents substitutions/site.
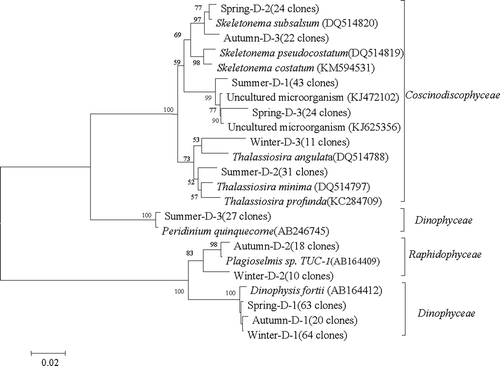
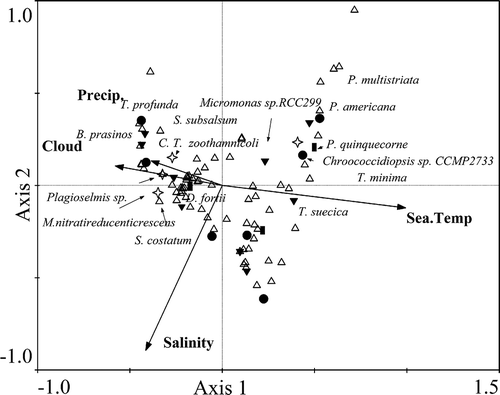
Fig. 4. The relationships between environmental factors and seawater samples collected in the coastal East China Sea in spring (samples 1–10, ▲), summer (samples 11–20, ◯), autumn (samples 21–30, □) and winter (samples 31–39, ²) analysed by Canonical Correspondence Analysis.
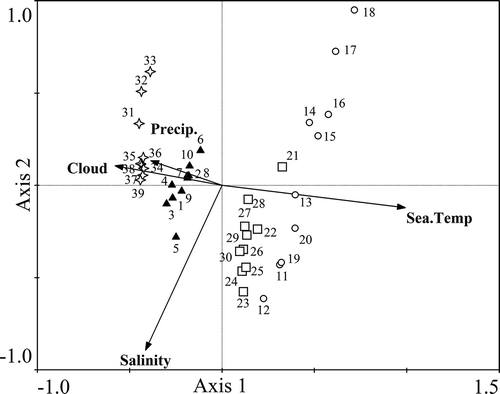
Fig. 5. Canonical correspondence analysis showing the relationships between environmental factors and dominant phytoplankton species including: Chroococcidiopsis sp. belonging to Cyanobacteria (●); Bathycoccus prasinos (B. prasinos), Micromonas sp. and Tetraselmis suecica (T. suecica) belonging to Chlorophyta (▼); Skeletonema costatum (S. costatum), Skeletonema subsalsum (S. subsalsum), Thalassiosira minima (T. minima), Pseudo-nitzschia multistriata (P. multistriata), Pseudo-nitzschia americana (P. americana) and Thalassiosira profunda (T. profunda) belonging to diatoms (△); Peridinium quinquecorne (P. quinquecorne) and Dinophysis fortii (D. fortii) belonging to dinoflagellates (■); Plagioselmis sp., Candidatus Thiobios zoothamnicoli (C. T. zoothamnicoli) and Methylophaga nitratireducenticrescens (M. nitratireducenticrescens) belonging to Proteobacteria (²).