Figures & data
Table 4. Morphological characters of C. olympicus compared with two similar chain-forming species: C. tenuissimus (this study) and C. wighamii (Bosak et al., 2015).
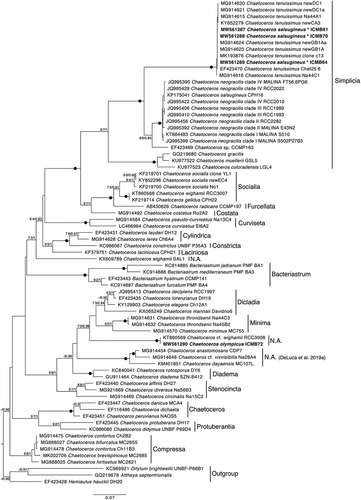
Fig. 45. LSU rDNA phylogenetic tree including sequences of some Chaetoceros representatives of interest and sequences obtained in this study (in bold). Sequences of Bacteriastrum species were used as outgroup. Statistical support shown in nodes corresponds to boostrap values (%) and Bayesian posterior probability. Only values >70% and 0.95 respectively are shown and black dots represent maximum statistical support. The different phylogenetic clusters were labelled according to morphological sections proposed in De Luca et al. (2019a). N.A. = species not assigned to any existing section. *C. tenuissimus after this work, see Discussion.