Figures & data
Table I. Chimeras GtrV→GtrX and GtrX→GtrV constructed by a PCR mediated approach and homologous recombination. P, positive. This table is reproduced in colour in Molecular Membrane Biology online.
Figure 1. Functional analysis of mutants. Electron micrographs (X50 000 magnification) of SFL1444 containing pNV1066 and 1067 tested with type V antibody. Secondary antibody used is conjugated to 10nm gold particles. Presence of gold particles is indicative of O-antigen modification. Description of each strain is shown below the image.
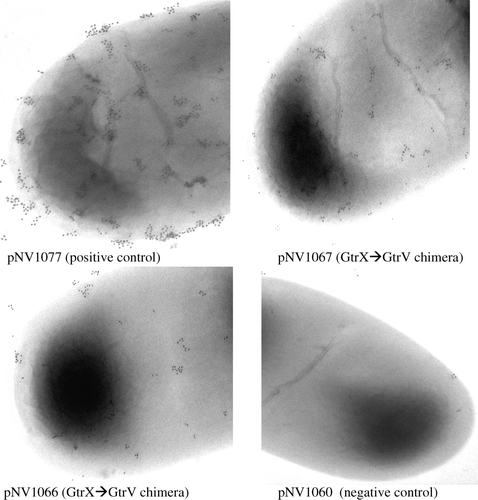
Figure 2. Western immunoblot of functional chimeras. Western immunoblot of chimeras pNV1066 and 1067 using mouse anti-E.coli PhoA antisera. pNV1105, positive control; pNV1060, negative control.

Figure 3. Best fit analysis of Gtrs. (A) Best fit analysis of GtrV and GtrX. Consensus is shown below sequence and in boxes. Note consensus regions boxed (red), indicative of motifs containing negatively charged residues. In vertical boxes shaded grey are the residues that were targeted by site-directed mutagenesis. (B) Best fit analysis of first periplasmic loop No 2 of N-terminus present in all Gtrs. The conserved aspartic acid (D) is shown in italics (grey box). Also in bold are either aspartic or glutamic acids part of the shown motif. This Figure is reproduced in colour in Molecular Membrane Biology online.
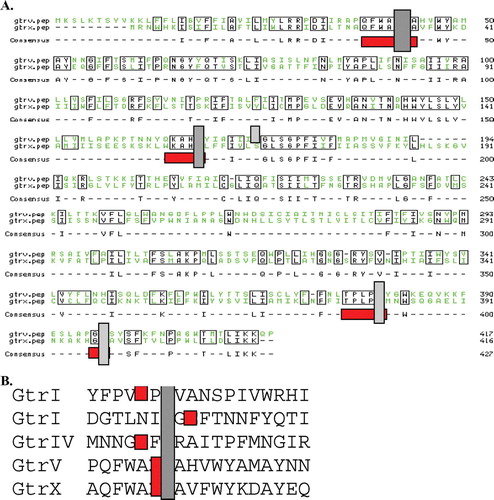
Figure 4. Position of critical GtrV residues and western immunoblot. Targeted residues in respect to the recently solved topology of GtrV. (A) Highlighted residues (bold-underlined) were mutated to residues shown next or above them. Note that important residues ED in the first periplasmic loop No 2 (N-terminus) were mutated to QN respectively, while residue D in loop No 10 (C-terminus) was mutated to N as well as A. (B) Western immunoblot of site-directed mutants pNV1173 and 1168 using mouse anti-E. coli PhoA antisera. A 98kDa band which includes both GtrV and dual reporter is observed in both constructs. This Figure is reproduced in colour in Molecular Membrane Biology online.
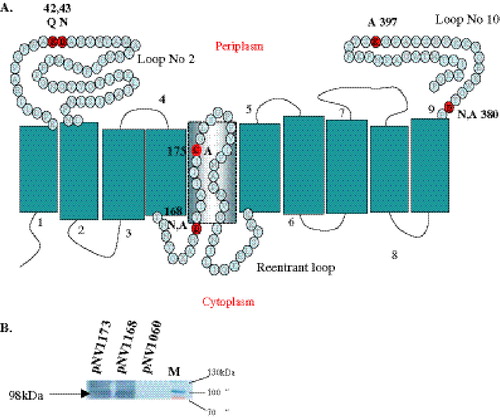
Table II. Site directed mutagenesis applied on selected GtrV residues. Mutations in bold are critical in function. P, Positive.
Supplementary Table I. Bacterial strains and plasmids used in this study.