Figures & data
Figure 1. Deficiency in BRAP homologous protein expression caused mild anxiety-like behaviors in male mice.
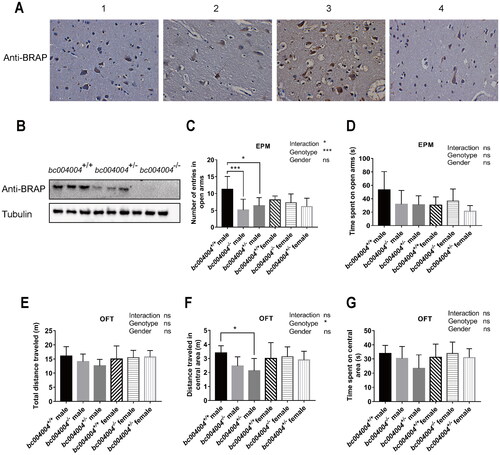
Figure 2. The analysis of behavioral tests after exposure to chronic unpredictable mild stress (CUMS). (A) Body weight changes throughout the CUMS procedure. (B) Immobility time of bc004004+/+ and bc004004−/− mice in the forced swimming test with treatment of CUMS. (C) Analysis of sucrose preference tests during the CUMS procedure. (D) Immobility time of bc004004+/+ and bc004004−/− mice in the tail suspension test after 21-day CUMS exposure. (E) Total distance traveled in the different regions in the open field test after 21-day CUMS exposure. Two-way ANOVA was performed followed by Tukey’s multiple comparisons. (F) The expression of BRAP homologous protein in mice detected by western blot with anti-BRAP antibody. The protein extracts of hippocampus tissues of the mice was used in western blot after 21-day of CUMS treatment. The expression of β-actin from the same sample was used as loading controls. Data are presented as mean ± SD. *p <.05, **p <.01, ***p <.001,****p <.0001, n = 8.
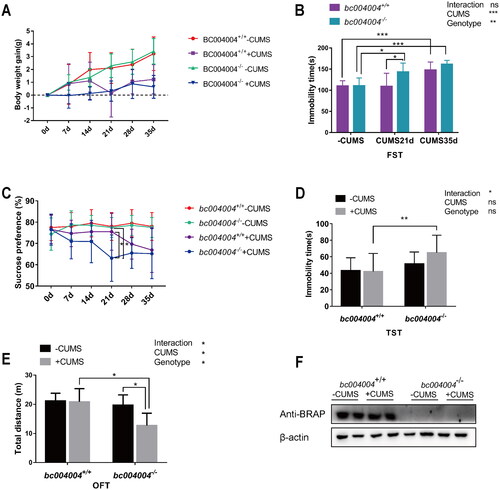
Figure 3. Novel object recognition (NOR) test in mice exposed to 21-day of CUMS treatment.
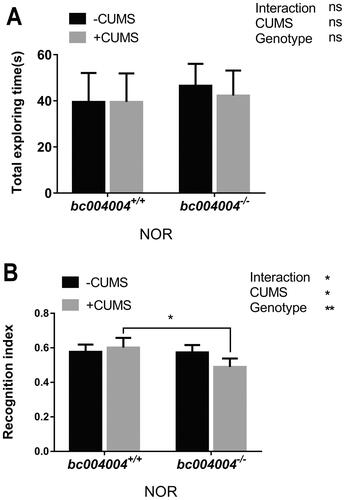
Figure 4. Analysis of dendritic morphology of the hippocampal neurons after 21-day of CUMS exposure.
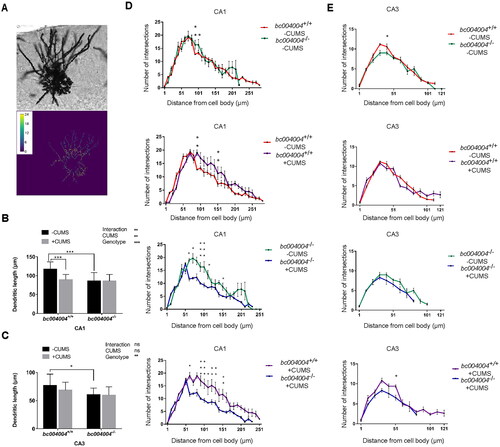
Figure 5. The amount and morphology of different dendritic spines on hippocampal CA1 and CA3 neurons.
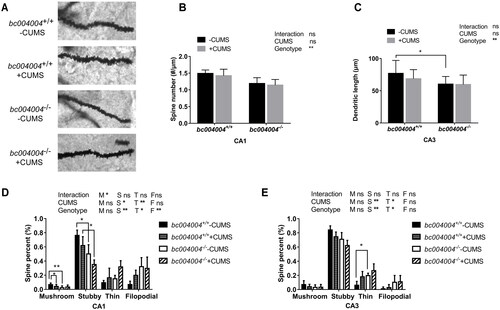
Figure 6. The expression of some proteins involved in synapse plasticity of the hippocampus.
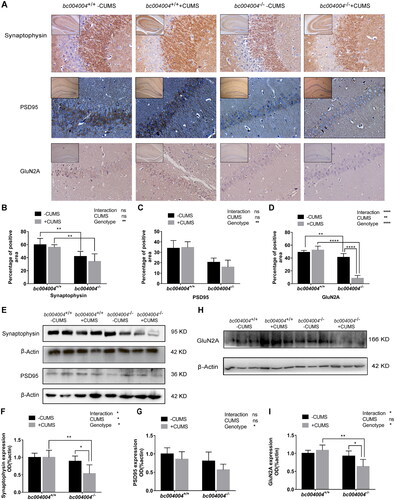
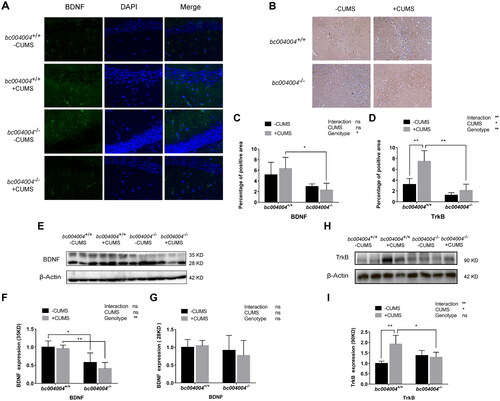
Figure 7. The impact of CUMS treatment on the expression of BDNF and TrkB in the hippocampal region of mice.