Figures & data
Figure 1. Timeline showing advances in culture-dependent (a) and culture-independent (b) approaches to study microbial dark matter (MDM).
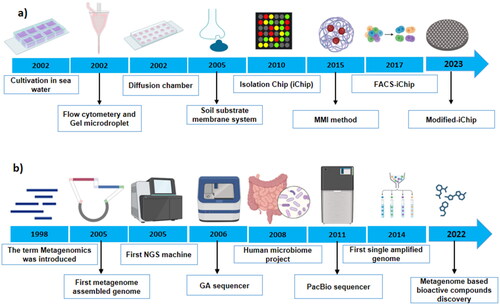
Figure 2. Schematic workflow used to generate SAGs and MAGs. Single-cell is physically separated from a complex environmental sample using microfluidics, flow cytometry, or a single-cell printer. Various genome binning tools are freely available, including MetaBAT (Kang et al. Citation2019), MaxBin (Wu et al. Citation2016), and CONCOCT (Alneberg et al. Citation2014). The quality of the draft genome is checked using CheckM or MDMcleaner and should follow the criteria for SAGs/MAGs suggested by the genome standard consortium (Bowers et al. Citation2017).
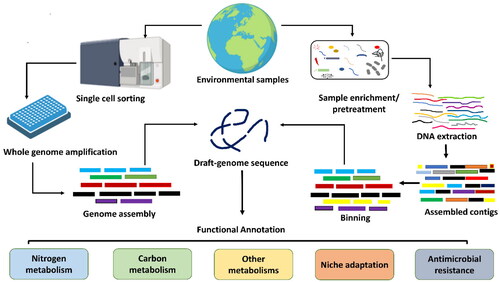
Table 1. Novel candidate prokaryotic phyla discovered through SAGs/MAGs.
Figure 3. Schematic representation of microbial dark matter (MDM) research progress and future opportunities.