Figures & data
Table 1. Average colony forming units per milliliter for each GIT section.
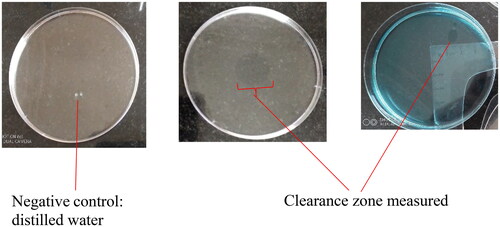
Table 2. Distribution of microbes showing clearance zones in sheep blood agar.
Table 3. Phenotypic characteristics of bacterial colonies subcultured from broiler birds’ GIT extracted samples.
Table 4. Oil spread test clearance zones and average emulsification index (E24) for biosurfactants from different sections of broiler GIT.
Figure 3. Dendogram of community relatedness of forty-nine bacterial strains based on the Jaccard’s coefficient.
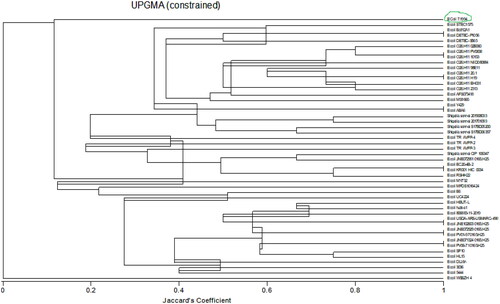
Table 5. Major phytochemical compounds identified in biosurfactant GCMS analysis.
Figure 4. A comparison of FTIR spectroscopy of Pseudomonas aeruginosa (bottom) and crude endogenous biosurfactant (top) extracted from E. coli strain. NB. Pseudomonas aeruginosa adapted from Khademolhosseini et al.Citation21
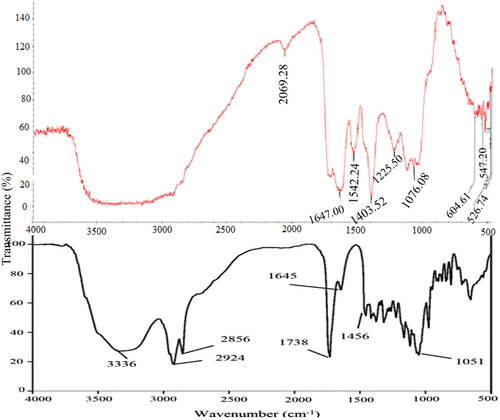
Table 6. FTIR peaks identified in Experimental biosurfactant.
Table 7. Small intestine extracted biosurfactant sample radio receptor assay output.
Table 8. Observed broiler GIT derived biosurfactants antimicrobial activity at 5% (m/v).
Data availability statement
Data are presented in-text, and a link has been provided for data in repository.