Figures & data
Table 1. Properties of chitosan used in the study.
Figure 1. Gel retardation assay of the DNA/chitosan complexes. Complexes containing 2 μg of salmon testes DNA were analyzed by 1% agarose gel electrophoresis at various N/P ratios: (A) DNA/chi-87K complexes, (B) DNA/chi-18K complexes. Arrows indicate: (1) loading position, (2) unretarded DNA.
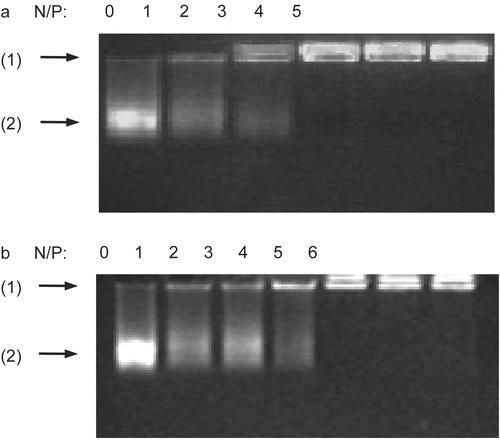
Figure 2. Zeta potential and particle size as a function of the N/P ratio of DNA/chitosan complexes prepared at the concentration of 0.2 mg/ml of DNA and chitosan. ▪ DNA/chi-87K complexes, o DNA/chi-18K complexes. The results are expressed as the mean ± SE (n = 5).
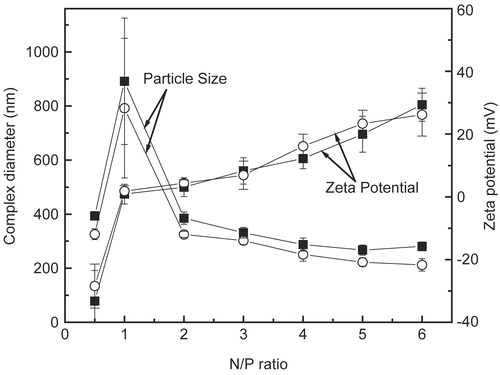
Figure 3. Transmission electron microscopy (TEM) photograph of DNA/chi-87K nanoparticles. Particles were observed after staining with 1% phosphotungstic acid. Bar = 100 nm.
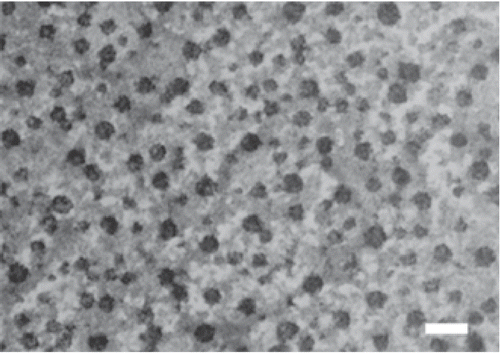
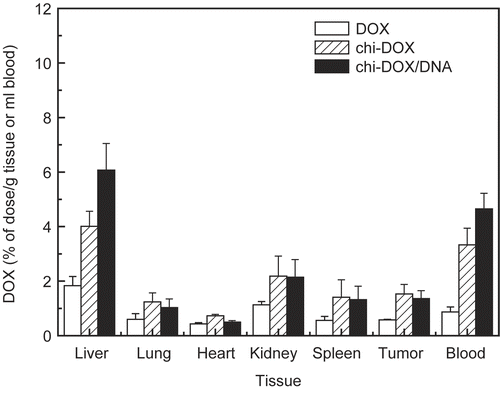
Figure 4. Tissue distribution of FITC-chitosan (A) and DNA/chitosan nanocomplexes (B) 4 hr after tail vain injection. ▪ FITC-chi-18K, □ FITC-chi-87K. Results are expressed as the mean ± SE (n = 5).
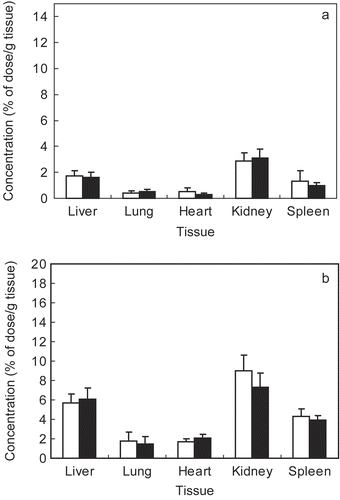
Figure 5. Time-course of FITC-chitosan concentrations in mouse liver (A) and blood (B) after tail vain injection. ▪ FITC-chi-87K, ▴ FITC-chi-18K, □ DNA/chi-87K nanocomplexes, △ DNA/chi-18K nanocomplexes. Results are expressed as the mean ± SE (n = 5).
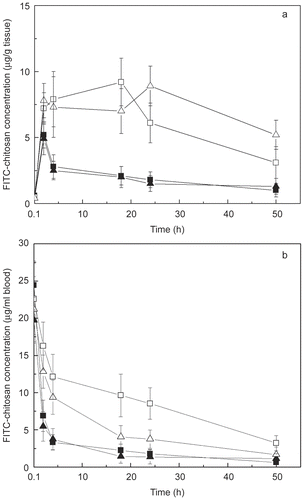
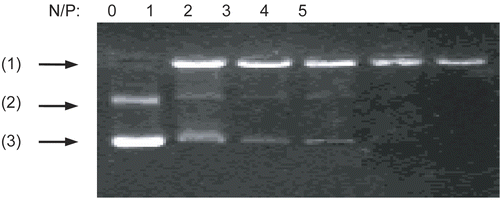
Figure 6. Gel retardation assay of DNA/chi-DOX complexes with various N/P ratios. Arrows indicate: (1) loading position, (2) open circle pDNA, and (3) supercoiled pDNA.