Figures & data
Table 1. Species list and DNA sequence information employed for phylogenetic analysis.
Figure 2. Phylogenetic relationship among available species in the genus Aspicilia based on a maximum likelihood analysis of the dataset of ITS sequences. The tree was rooted with six Lobothallia and Teuvoa sequences. Maximum likelihood bootstrap values ≥ 70% and posterior probabilities ≥ 95% are shown above internal branches. Branches with bootstrap values ≥ 90% are shown in bold. The new species Aspicilia humida is presented in bold, and all species names are followed by the Genbank accession numbers. Reference provides the species related to the specific GenBank accession numbers and voucher information.
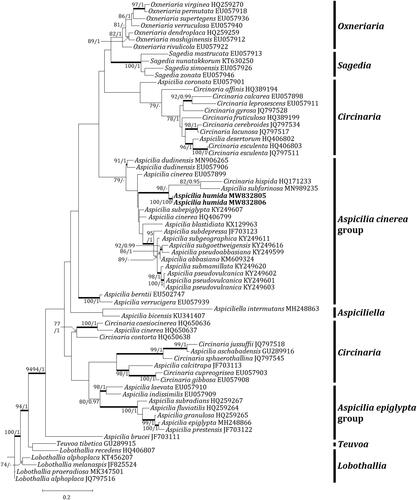
Figure 3. Phylogenetic relationships among available species in the genus Aspicilia based on a maximum likelihood analysis of the dataset of the mitochondrial small subunit (mtSSU) sequences. The tree was rooted with five Lobothallia sequences. Maximum-likelihood bootstrap values ≥ 70% and posterior probabilities ≥ 95% are shown above internal branches. Branches with bootstrap values ≥ 90% are shown in bold. The new species Aspicilia humida is presented in bold, and all species names are followed by the GenBank accession numbers. Reference provides the species related to the specific GenBank accession numbers and voucher information.
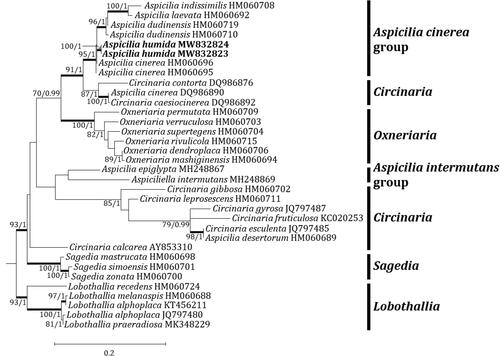
Figure 4. Aspicilia humida (BDNA-L-0000703, holotype for A–K; BDNA-L-0000711, paratype for L & M) in morphology. (A–D): Habitus and apothecia emerging single to several per an areole; (E): Adnate apothecia without constriction at the base in section; (F): Epihymenium in olive-brown pigment; (G–J): Clavate asci with eight spores; (K): Ellipsoid or globose ascospores with no septation; (L): Immersed pycnidia; (M): Thread-like pycnoconidia. Bars: A–D 1 mm; E 200 μm; F 50 μm; G–K 10 μm; L 100 μm; M 10 μm.
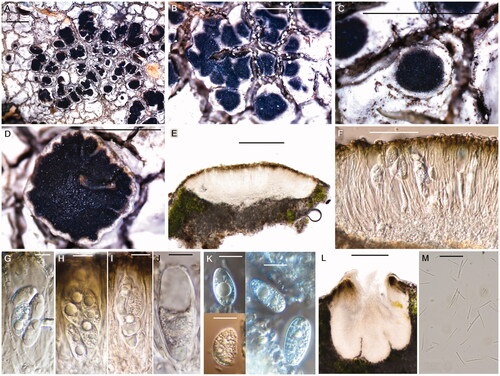
Table 2. Comparison of Aspicilia humida with closely-related species.

