Figures & data
Figure 2. PCR amplification products of NBS RGAs in mango. M: DNA Marker V; Lane 1: blank control; Lane 2: control using water as a template; Lane 3: PCR products of NBS genes amplified with forward and reverse primers for mango.
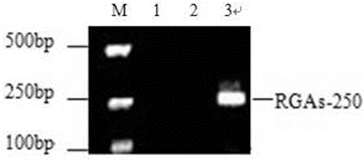
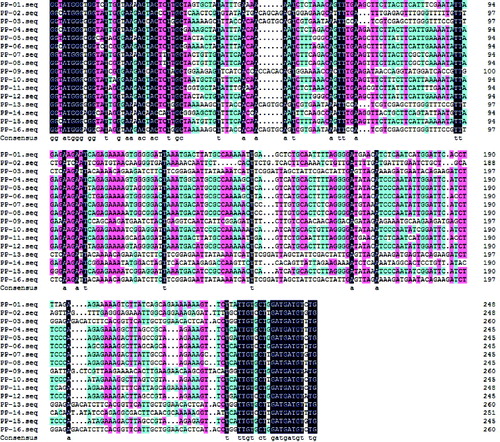
Table 1 Variation sites of 16 RGAs sequences in mango.
Figure 4. Polymorphism characteristic of NBS–LRR analogues isolated from mango (pp-01–16). The vertical axis represents the nucleotide diversity (Pi), and the horizontal axis represents the nucleotide position (bp), the curved line represents the change of Pi in different nucleotide positions.
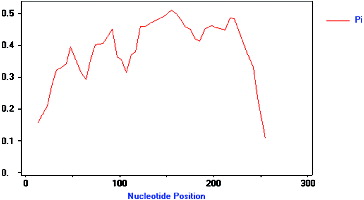
Table 2 Results from the BL2SEQ algorithm showing the extent of identity between the RGAs isolated in the present study.