Figures & data
Table 1. Predicted amino-acid composition from the coding region.
Table 2. Contents of four amino acids in PFMG2, MSI60, MSI31, MSI7 and Nacrein.
Figure 2. Putative conserved sequence of the PFMG2 protein and phylogenetic analysis. (A) A putative conserved CH domain is found at the N-terminus. (B) Alignments of PFMG2 and Transgelin in Bos taurus [Bos] (NP_001039 614.1), Equus caballus [Equus] (NP_001103 604.1), Homo sapiens [Homo] (NP_001001 522.1), Rattus norvegicus [Rattus] (NP_113 737.1), Gallus gallus [Gallus] (NP_990 825.1), Xenopus laevis [Xenopus] (NP_001083 600.1), Sparus aurata [Sparus] (ADO67626.1), Mus musculus [Mus] (NP_035 656.1), Haliotis discus discus [Haliotis] (ABO26639.1). ‘*’ indicates residues that are identical in all sequences in the alignment. ‘:’ indicates the presence of conserved substitutions. ‘.’ indicates the presence of semi-conserved substitutions. (C) Phylogenetic analysis of PFMG2 protein and its closest putative homologues in different species. The phylogenetic tree shows the evolutionary relationship of the protein from various species mentioned in B.
![Figure 2. Putative conserved sequence of the PFMG2 protein and phylogenetic analysis. (A) A putative conserved CH domain is found at the N-terminus. (B) Alignments of PFMG2 and Transgelin in Bos taurus [Bos] (NP_001039 614.1), Equus caballus [Equus] (NP_001103 604.1), Homo sapiens [Homo] (NP_001001 522.1), Rattus norvegicus [Rattus] (NP_113 737.1), Gallus gallus [Gallus] (NP_990 825.1), Xenopus laevis [Xenopus] (NP_001083 600.1), Sparus aurata [Sparus] (ADO67626.1), Mus musculus [Mus] (NP_035 656.1), Haliotis discus discus [Haliotis] (ABO26639.1). ‘*’ indicates residues that are identical in all sequences in the alignment. ‘:’ indicates the presence of conserved substitutions. ‘.’ indicates the presence of semi-conserved substitutions. (C) Phylogenetic analysis of PFMG2 protein and its closest putative homologues in different species. The phylogenetic tree shows the evolutionary relationship of the protein from various species mentioned in B.](/cms/asset/5ab7ba2b-21f1-412d-8d6c-3a892eed7c1d/tbeq_a_1348254_f0002_oc.jpg)
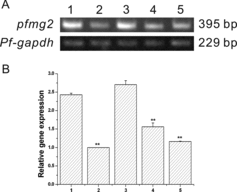
Figure 3. Pfmg2 gene expression pattern in P. fucata. (A) RT-PCR for analysis of pfmg2 gene expression in five tissues from P. fucata: 1, mantle; 2, foot; 3, adductor muscle; 4, gill; 5, viscera; gapdh was included as a positive control. The pfmg2 DNA PCR product is 395 bp in length, and Pf-gapdh is 229 bp. (B) 1, mantle; 2, foot; 3, adductor muscle; 4, gill; 5, viscera. Data are expressed as means ± SE from all experiments, as indicated **p < 0.01.